Table of Contents¶
- 1 3.1 In this session
- 2 E3.1 Exercise: listing
- 3 3.2 Getting and Plotting Some Data: netCDF format
- 3.1 3.2.1 GlobAlbedo data
- 3.2 3.2.2 Loading from within Python
- 3.3 Exercise E3.2 Read Image Data into 3D List
- 3.4 Exercise E3.3 Read More Image Data into a 4D List
- 3.5 3.2.3 Plotting an image
- 3.6 E3.2 Exercise: Making Movies
- 3.7 E3.2.1 Software
- 3.8 E3.2.2 Looping over a set of images
- 3.9 E3.2.3 Make the movie
- 4 Methods
- 5 3.3 Numpy
- 6 E3.3 Exercise: 3D Masked Array
- 7 3.4 Average Albedo
- 8 3.5 Solar Radiation
- 9 E3.5 Exercise: Solar Radiation Code: A chance for you to get feedback
- 10 3.6 Saving and loading numpy data
- 11 E3.6 Exercise: awkward reading
- 12 Advanced and Answers
3. Plotting and Numerical Python¶
3.1 In this session¶
In this session, we will introduce and use some packages that you will commonly use in scientific programming.
These are:
- numpy: NumPy is the fundamental package for scientific computing with Python
- matplotlib: Python 2D plotting library
We will also introduce some additional programming concepts, and set an exercise that you can do and get feedback on.
3.2.1 Getting Started¶
To get started for this session, you should go to your Data area and download or update your material for this course.
If you haven't previously downloaded the course, type the following at the unix prompt (illustrated by berlin% here):
berlin% cd DATA
berlin% git clone https://github.com/profLewis/geogg122.git
berlin% cd geogg122/Chapter3_Scientific_Numerical_Python
If you already have a clone of the course repository (which you should have from last week):
berlin% cd DATA/geogg122
berlin% git reset --hard HEAD
berlin% git pull
berlin% cd Chapter3_Scientific_Numerical_Python
This will update your local copy with any new notes or files. Any additional files that you yourself have created should be still where they were.
E3.1 Exercise: listing¶
Using Python, produce a listing of the files in the subdirectory data of geogg122/Chapter3_Scientific_Numerical_Python that end with .nc and put this listing in a file called data/data.dat with each entry on a different line
3.2 Getting and Plotting Some Data: netCDF format¶
3.2.1 GlobAlbedo data¶
Before we start to use these new packages, we will start by getting some data for you to use and show you how to read it in and do some basic plotting.
Today, we will be using the ESA GlobAlbedo data.
These data come in different spatial resolutions, but here we will use the global product at 0.5 degrees.
The files are acessible via http in the directory:
http://gws-access.cems.rl.ac.uk/public/qa4ecv/albedo/netcdf_cf/05deg/monthly
The filenames are of the pattern:
GlobAlbedo.merge.albedo.05.YYYYMMM.nc
where YYYY is the year (e.g. 2009) and MM is the month (01 is January, 12 is December).
The data are in netCDF format which is a common binary data format.
ga_root_url = 'http://gws-access.cems.rl.ac.uk/public/qa4ecv/albedo/netcdf_cf/05deg/monthly/'
# example filename : use formatting string:
# %d%02d
year = 2009
month = 1
url = ga_root_url + '/GlobAlbedo.merge.albedo.05.%d%02d.nc' % (
year, month )
print url
# Or alternatively
url = ga_root_url + \
'/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc'.format( year, month )
print url
http://gws-access.cems.rl.ac.uk/public/qa4ecv/albedo/netcdf_cf/05deg/monthly//GlobAlbedo.merge.albedo.05.200901.nc http://gws-access.cems.rl.ac.uk/public/qa4ecv/albedo/netcdf_cf/05deg/monthly//GlobAlbedo.merge.albedo.05.200901.nc
3.2.2 Loading from within Python¶
We have found how to write the URL for each of this netcdf files. We could then just download them. For this we'd have to:
- Decide what year and month we want,
- Combine the root part of the URL with the filename, substituting the month and year in the latter,
- Download the file
- Save the file into a file.
We have already provided data in the data folder, but here's how you could do it yourself with the requests method (see also advanced section):
import requests
# Import requests module
while False:
# We want to iterate over two years, 2009 and 2010
for year in [2009, 2010]:
# Each year has 12 months, from 1 to 12, so we produce a list between 1 and 13-1
for month in range(1,13):
# the URL is composed as above
url = ga_root_url + '/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc'.format( year, month )
# This obtains the file from the remote server
r = requests.get(url)
# We first check whether the connection was OK
if r.ok:
# It worked!!! Yay!!! We extract the filename as the lat bit of the URL
# split by "/" and select the last element
fname = url.split("/")[-1]
# We open the destination file in the data folder
with open("data/" + fname, 'wb') as fp:
# Dump content of remote file into local file
fp.write(r.content)
# finished writing!
print "Wrote {}".format(fname)
else:
# something went wrong!
print "Couldn't download {}".format(url)
!ls data/GlobAlbedo*.nc
data/GlobAlbedo.merge.albedo.05.200901.nc data/GlobAlbedo.merge.albedo.05.200902.nc data/GlobAlbedo.merge.albedo.05.200903.nc data/GlobAlbedo.merge.albedo.05.200904.nc data/GlobAlbedo.merge.albedo.05.200905.nc data/GlobAlbedo.merge.albedo.05.200906.nc data/GlobAlbedo.merge.albedo.05.200907.nc data/GlobAlbedo.merge.albedo.05.200908.nc data/GlobAlbedo.merge.albedo.05.200909.nc data/GlobAlbedo.merge.albedo.05.200910.nc data/GlobAlbedo.merge.albedo.05.200911.nc data/GlobAlbedo.merge.albedo.05.200912.nc data/GlobAlbedo.merge.albedo.05.201001.nc data/GlobAlbedo.merge.albedo.05.201002.nc data/GlobAlbedo.merge.albedo.05.201003.nc data/GlobAlbedo.merge.albedo.05.201004.nc data/GlobAlbedo.merge.albedo.05.201005.nc data/GlobAlbedo.merge.albedo.05.201006.nc data/GlobAlbedo.merge.albedo.05.201007.nc data/GlobAlbedo.merge.albedo.05.201008.nc data/GlobAlbedo.merge.albedo.05.201009.nc data/GlobAlbedo.merge.albedo.05.201010.nc data/GlobAlbedo.merge.albedo.05.201011.nc data/GlobAlbedo.merge.albedo.05.201012.nc
We use the gdal module to read netCDF data:
import gdal
# example filename : use formatting string:
# %d%02d
year = 2009
month = 1
folder = "data"
filename = folder + \
'/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc'.format( year, month )
print filename
g = gdal.Open(filename)
# g should now be a GDAL dataset, but if the file isn't found
# g will be none. Let's test this:
if g is None:
print "Problem opening file %s!" % filename
else:
print "File %s opened fine" % filename
subdatasets = g.GetSubDatasets()
for name, fname in subdatasets:
print name
print "\t", fname
data/GlobAlbedo.merge.albedo.05.200901.nc File data/GlobAlbedo.merge.albedo.05.200901.nc opened fine NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_VIS [1x360x720] DHR_VIS (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_NIR [1x360x720] DHR_NIR (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_SW [1x360x720] DHR_SW (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_VIS [1x360x720] BHR_VIS (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_NIR [1x360x720] BHR_NIR (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_SW [1x360x720] BHR_SW (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_sigmaVIS [1x360x720] DHR_sigmaVIS (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_sigmaNIR [1x360x720] DHR_sigmaNIR (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_sigmaSW [1x360x720] DHR_sigmaSW (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_sigmaVIS [1x360x720] BHR_sigmaVIS (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_sigmaNIR [1x360x720] BHR_sigmaNIR (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_sigmaSW [1x360x720] BHR_sigmaSW (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_alpha_VIS_NIR [1x360x720] DHR_alpha_VIS_NIR (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_alpha_VIS_SW [1x360x720] DHR_alpha_VIS_SW (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_alpha_NIR_SW [1x360x720] DHR_alpha_NIR_SW (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_alpha_VIS_NIR [1x360x720] BHR_alpha_VIS_NIR (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_alpha_VIS_SW [1x360x720] BHR_alpha_VIS_SW (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":BHR_alpha_NIR_SW [1x360x720] BHR_alpha_NIR_SW (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":Weighted_Number_of_Samples [1x360x720] Weighted_Number_of_Samples (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":Goodness_of_Fit [1x360x720] Goodness_of_Fit (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":Time_to_the_Closest_Sample [1x360x720] Time_to_the_Closest_Sample (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":Data_Mask [1x360x720] Data_Mask (8-bit integer) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":Snow_Fraction [1x360x720] Snow_Fraction (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":Solar_Zenith_Angle [1x360x720] Solar_Zenith_Angle (32-bit floating-point) NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":Relative_Entropy [1x360x720] Relative_Entropy (32-bit floating-point)
Here, we will want to access 'DHR_VIS', 'DHR_NIR' and 'DHR_SW', which are the bihemispherical reflectance (DHR: 'black sky albedo' -- the albedo under directional illumination conditions) for visible, near infrared and total shortwave wavebands.
From the information printed in the loop above, we note that the field of information we need to access is of the form:
NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS
The way we do this in gdal is to use gdal.Open(f) where f is the full data layer specification (in this case e.g. NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS) to open a particular dataset within the file.
We then read the (raster) data into an array with g.Open(f).ReadAsArray()
for example then:
f = 'NETCDF:"data/GlobAlbedo.merge.albedo.05.200901.nc":DHR_VIS'
data = gdal.Open(f).ReadAsArray()
print data
[[ nan nan nan ..., nan nan
nan]
[ nan nan nan ..., nan nan
nan]
[ nan nan nan ..., nan nan
nan]
...,
[ 0.71308476 0.71428406 0.74763721 ..., 0.71503514 0.71508819
0.71491981]
[ nan 0.71366459 0.71380746 ..., 0.71256566 0.71183181
0.71170306]
[ 0.94759613 0.94759613 0.94759613 ..., 0.94759613 0.94759613
0.94759613]]
We can make this more flexible by building the string f from the specific pieces of information we require.
In this case, it helps to use a form of template for the string we need:
# form a generic string of the form
# NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS
file_template = 'NETCDF:"{:s}":{:s}'
# now make a list of the datset names we want
# so we can loop over this
selected_layers = ['DHR_VIS','DHR_NIR','DHR_SW']
# ----------------------------------
# try it out:
root = 'data/'
# example filename : use formatting string:
# %d%02d
year = 2009
month = 1
filename = root + '/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc'.format( year, month )
print filename
# set up an empty dictionary to load the data into
data = {}
# use enumerate here to loop over
# the list selected_layers and also have
# access to an index i
for i, layer in enumerate ( selected_layers ):
this_file = file_template.format( filename, layer )
print "Opening Layer %d: %s" % (i, this_file )
g = gdal.Open ( this_file )
# test that the opening worked
# and raise an error otherwise
if g is None:
raise IOError
data[layer] = g.ReadAsArray()
print "\t>>> Read %s!" % layer
data//GlobAlbedo.merge.albedo.05.200901.nc Opening Layer 0: NETCDF:"data//GlobAlbedo.merge.albedo.05.200901.nc":DHR_VIS >>> Read DHR_VIS! Opening Layer 1: NETCDF:"data//GlobAlbedo.merge.albedo.05.200901.nc":DHR_NIR >>> Read DHR_NIR! Opening Layer 2: NETCDF:"data//GlobAlbedo.merge.albedo.05.200901.nc":DHR_SW >>> Read DHR_SW!
albedo = []
for layer in selected_layers:
albedo.append ( data[layer])
print len(albedo)
albedo = [ data[layer] for layer in selected_layers]
3
If we really want the data in a single (3D) array, we could then create this from the dictionary entries:
# here, we just make a *list*
# as we will see below, a list might be a bit limited for these purposes and
# we would normally use a numpy array
albedo = [data[f] for f in selected_layers]
# print some information about the list we have made
print len(albedo),len(albedo[0]),len(albedo[0][0])
print albedo[0].shape
3 360 720 (360, 720)
Exercise E3.2 Read Image Data into 3D List¶
Write some python code that directly reads the data layers 'DHR_VIS','DHR_NIR','DHR_SW' into a list for a given month and year.
Exercise E3.3 Read More Image Data into a 4D List¶
You should now have some code that reads the 3 albedo datasets into a 3D list (3 x 360 x 720) for a given month and year.
Put a loop around this code to make a 4D list dataset (12 x 3 x 360 x 720) for all months in a given year.
3.2.3 Plotting an image¶
We can plot image data using the package pylab (from matplotlib).
If you are running this as a notebook, make sure you use:
ipython notebook --pylab=inline if you want the images to appear in the notebook.
First we have to import pylab, which we will refer to a plt (se import pylab as plt).
The basic function for plotting an image is plt.imshow:
import matplotlib.pyplot as plt
%matplotlib inline
# declare a new figure and its size
#plt.figure(figsize=(10,4))
plt.imshow(albedo[0])
<matplotlib.image.AxesImage at 0x7f414bcf3690>
Thats fine, but we might apply a few modifications to make a better plot:
# change the figure size
plt.figure(figsize=(10,4))
# use nearest neighbour interpolation
plt.imshow(albedo[0],interpolation='nearest')
# show a colour bar
plt.colorbar()
print albedo[0]
[[ nan nan nan ..., nan nan
nan]
[ nan nan nan ..., nan nan
nan]
[ nan nan nan ..., nan nan
nan]
...,
[ 0.71308476 0.71428406 0.74763721 ..., 0.71503514 0.71508819
0.71491981]
[ nan 0.71366459 0.71380746 ..., 0.71256566 0.71183181
0.71170306]
[ 0.94759613 0.94759613 0.94759613 ..., 0.94759613 0.94759613
0.94759613]]
Albedo should lie between 0 and 1, so there are clearly a few 'funnies' that we might flag later.
For now, we might like to plot only values between 0 and 1 (thresholding at those values).
Also if we don't like this colour map, we can try another:
# change the figure size
plt.figure(figsize=(10,4))
# no interpolation
plt.imshow(albedo[0], interpolation='nearest',
cmap=plt.get_cmap('gist_stern'), vmin=0.0, vmax=1.0)
# show a colour bar
plt.colorbar()
<matplotlib.colorbar.Colorbar at 0x7f414bc69e90>
If we want to make use of strings to label each month. We can do this if we for example load up all the albedo data into a 12-element monthly list, where each element is an the monthly albedo array, and then looping over. The following snippet does this. Note what we do:
- We define template strings for the netcdf complete name, as well as for the actual filename
- We then loop over the month number, first generating the actual filename string, and then introducing it in the NETCDF complete name, also selecting the layer (in this case,
BHR_VIS) - We append the data to an empty list called
albedo
Here's how the code looks like:
file_template = 'NETCDF:"{:s}":{:s}' # Template for the Netcdf path
layer = "BHR_VIS" # the layer we'll use
fname_template = 'data/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc' # the actual filename
year = 2009 # Year
albedo = [] # Empty list
for month in range(12):
# Looping over all months
fname = fname_template.format(year, month+1)
# fname now holds the actual filename
fich = file_template.format(fname, layer)
# We put together the full netcdf indicator
g = gdal.Open(fich)
# Opened with gdal, should check it works, but...
data = g.ReadAsArray()
# Read array
albedo.append(data)
#Append to empty list
print "The new list has {} elements".format(len(albedo))
The new list has 12 elements
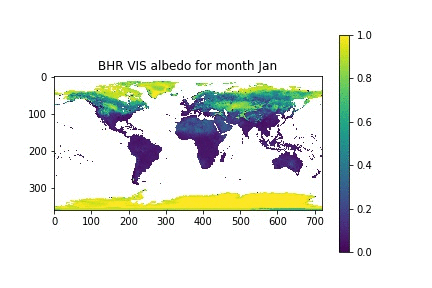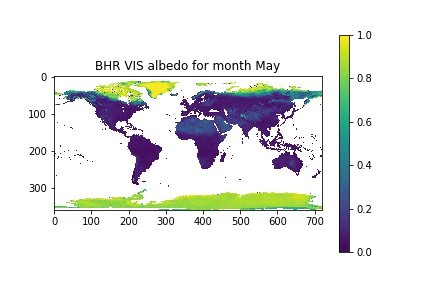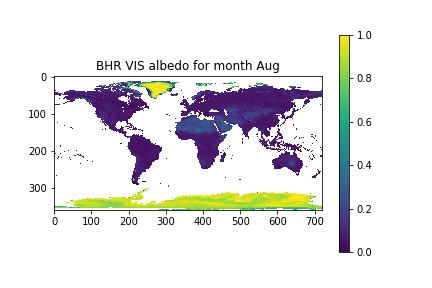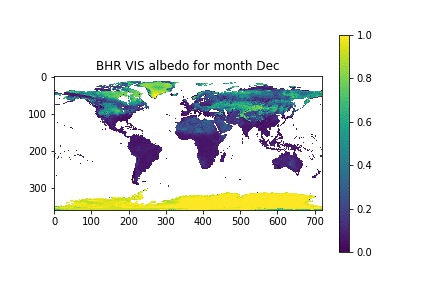
Now, let's assume that we have a list with the month names, similar to what we had in the previous session. We can then loop over the month numbers (0 to 11), open a new figure, plot the data for that month, and set the title for that month by looking up the relevant position on the month_names list that we have put together.
month_names = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec']
for i in xrange(12):
plt.figure()
plt.imshow(albedo[i], vmin=0, vmax=1, interpolation="nearest")
plt.title("BHR VIS albedo for month {}".format(month_names[i]))
plt.colorbar()
You can click on the plots above and save them to individual files, but that's not practical in many situations. We can use the savefig command. This command takes up a filename with a known extension (usually .png, .jpg or .pdf, as well some other extra options that define the plot resolution.
year = 2009
month = 1
# change the figure size
plt.figure(figsize=(10,4))
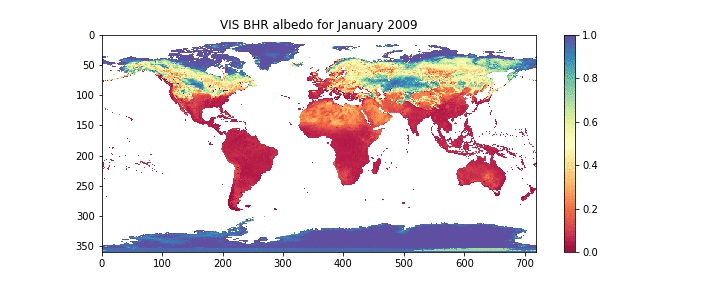
plt.title('VIS BHR albedo for %s %d'%(month_names[month],year))
# use nearest neighbour interpolation
plt.imshow(albedo[0],interpolation='nearest',
cmap=plt.get_cmap('Spectral'),vmin=0.0,vmax=1.0)
# show a colour bar
plt.colorbar()
# save the plot to a jpeg image file
plt.savefig("data/albedo.pdf")
E3.2 Exercise: Making Movies¶
E3.2.1 Software¶
You can sort of make movies in pylab, but you generally have to make a system call to unix at some point, so it's probably easier to do this all in unix with the utility convert.
At the unix prompt, chack that you have access to convert:
berlin% which convert
/usr/bin/convert
If this doesn't come up with anything useful, there is probably a version in /usr/bin/convert or /usr/local/bin/convert (If you don't have it on your local machine, install ImageMagick which contains the command line tool convert).
To use this, e.g.:
from the unix command line:
berlin% cd ~/DATA/geogg122/Chapter3_Scientific_Numerical_Python
berlin% convert data/albedo.jpg data/albedo.gif
or from within a notebook:
!convert data/albedo.jpg data/albedo.gif
Or, more practically here, you can run a unix command directly from Python:
import subprocess
subprocess.call(["convert", "data/albedo.jpg", "data/albedo.gif"])
0
This will convert the file files/data/albedo.jpg (in jpeg format) to files/data/albedo.gif (in gif format).
We can also use convert to make animated gifs, which is one way of making a movie.
E3.2.2 Looping over a set of images¶
You have all of the code you need above to be able to read a GlobAlbedo file for a given month and waveband in Python and save a picture in jpeg format, but to recap for BHR_VIS:
for i in xrange(12):
plt.figure()
plt.imshow(albedo[i], vmin=0, vmax=1, interpolation="nearest")
plt.title("BHR VIS albedo for month {}".format(month_names[i]))
plt.colorbar()
output_filename = "data/albedo_{:02d}.jpg".format(i+1)
plt.savefig(output_filename)
subprocess.call(["convert", output_filename,
output_filename.replace(".jpg", ".gif")])
plt.close() # Close figure
Modify the code above to loop over each month, so that it generates a set of gif format files for the TOTAL SHORTWAVE ALBEDO
You should confirm that these exist, and that the file modification time is when you ran it (not when I generated the files for these notes, which is Oct 10 2013).
ls -l data/albedo_??.gif
-rw-rw-r-- 1 ucfajlg ucfajlg 26054 Oct 24 20:11 data/albedo_01.gif -rw-rw-r-- 1 ucfajlg ucfajlg 26015 Oct 24 20:11 data/albedo_02.gif -rw-rw-r-- 1 ucfajlg ucfajlg 26320 Oct 24 20:11 data/albedo_03.gif -rw-rw-r-- 1 ucfajlg ucfajlg 26169 Oct 24 20:11 data/albedo_04.gif -rw-rw-r-- 1 ucfajlg ucfajlg 26237 Oct 24 20:11 data/albedo_05.gif -rw-rw-r-- 1 ucfajlg ucfajlg 25913 Oct 24 20:11 data/albedo_06.gif -rw-rw-r-- 1 ucfajlg ucfajlg 25505 Oct 24 20:11 data/albedo_07.gif -rw-rw-r-- 1 ucfajlg ucfajlg 25356 Oct 24 20:11 data/albedo_08.gif -rw-rw-r-- 1 ucfajlg ucfajlg 26418 Oct 24 20:11 data/albedo_09.gif -rw-rw-r-- 1 ucfajlg ucfajlg 26295 Oct 24 20:11 data/albedo_10.gif -rw-rw-r-- 1 ucfajlg ucfajlg 25749 Oct 24 20:11 data/albedo_11.gif -rw-rw-r-- 1 ucfajlg ucfajlg 26026 Oct 24 20:11 data/albedo_12.gif
E3.2.3 Make the movie¶
The unix command to convert these files to an animated gif is:
convert -delay 100 -loop 0 data/albedo_??.gif data/albedo_2009.gif
Run this (ideally, from within Python) to create the animated gif albedo_2009.gif
Again, confirm that you created this file (and it is not just a version you downloaded):
ls -l data/albedo_2009.gif
-rw-rw-r-- 1 ucfajlg ucfajlg 311922 Oct 24 18:58 data/albedo_2009.gif
Methods¶
We've seen how to load data, do plots and so on. We can see that this is something we might want to repeat many times, and one way to minimise the errors from "copying and pasting" things around is to pack everything into a method or a function (we'll use the term interchangeably).
A function takes a number of parameters (some of them can have some default values), does something with the data and returns something back. For example, we can think that to read some GA data for any given month and year, we need:
- The folder where the data are
- The month
- The year
- The layer
We would get back a 2D array with the data loaded up. The function is called read_globalbedo (you can call it whatever you want, but remember that you'll want to remember it!)
def read_globalbedo(month, year, folder="data/", layer="BHR_VIS"):
"""Read globalbedo data when given month and year.
Parameters
------------
month: int
The month from 1 to 12.
year: int
The year, currently only 2009 and 2010 available, but can download more.
folder: str
The folder where the GA data are stored.
layer: str
The data layer
Returns
---------
A 2D array"""
file_template = 'NETCDF:"{:s}":{:s}' # Template for the Netcdf path
fname_template = '{:s}/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc' # the actual filename
fname = fname_template.format(folder, year, month)
netcdf_fname = file_template.format(fname, layer)
g = gdal.Open(netcdf_fname)
if g is None:
raise IOError("Problem with reading file {}".format(fname))
data = g.ReadAsArray()
return data
In the above code the function takes two mandatory parameters: month and year, and two optional parameters, that if they are not specified will the ones in the function header (e.g. if you don't specify folder, it'll be automatically set to data/, and if you don't specify layer, you'll get BHR_VIS automatically). The function will return a 2D albedo array as before:
data = read_globalbedo(1, 2009)
3.3 Numpy¶
3.3.1 Numpy Arrays¶
At the heart of the numpy module is a a powerful N-dimensional array object.
Among many other things, this allows us to do arithmetic (and other operations) directly on arrays, rather than having to loop over each element (as we did with lists for example).
Two main benefits of this are:
- the code is much easier to read
- running the code is much more efficient (because, 'under the hood' as it were of the programming language, we can do fast operations such as vector processing).
As our first example, let's read the visible albedo data we examined above into a numpy array.
We will start with just one month of data:
import numpy as np
data = read_globalbedo(2, 2009)
# some interesting things about numpy arrays
print "the array type is now",type(data)
print "the shape of the array is",data.shape
print "the size of the array is",data.size
print "the number of dimensions is",data.ndim
print "the data type in the array is",data.dtype
the array type is now <type 'numpy.ndarray'> the shape of the array is (360, 720) the size of the array is 259200 the number of dimensions is 2 the data type in the array is float32
A few things to notice here.
GDAL reads the data and returns a numpy array directly. However, if we want to have access to all of Numpy's functionality, we first need to import it using
import numpy as np
which means that locally, we refer to this module as np (saves us writint numpy).
We can convert lists that have all elements of the same data type into an array by using the array method of the numpy library:
np.array([1,2,3])
Then we saw at the end of this code a number of commonly used methods for the numpy class that provide us with information on the shape of the array (N.B. this is (rows,cols) as you might notice from the plots we generated), the total size (number of elements), number of dimansions.
We also saw that we could access the array data type (dtype). This is very different to a list or tuple then, because a numpy array can only contain data of a single data type (but lists or tuples can have different data types in each element).
astype¶
We can convert between data types using astype:
a = np.array([1,2,3])
print a.dtype
int64
b = a.astype(float)
print b
print b.dtype
[ 1. 2. 3.] float64
slice¶
As you would expect, we can slice in a numpy array. All that is different is that we set up a slice of each dimension, e.g.:
# see if you can work out why this is
# the size, shape and ndim it is
print data[-3:-1,10:30:5]
print data[-3:-1,10:30:5].shape
print data[-3:-1,10:30:5].ndim
[[ 0.87582344 0.87606007 0.87689805 0.87557447] [ 0.79205 0.65575886 0.87453514 0.87763792]] (2, 4) 2
If you want to specify all elements in a given dimension you need to use at least one :, e.g.
# what does this mean?
print data[10:20,0]
print data[10:20,0].shape
[ nan nan nan nan nan nan nan nan nan nan] (10,)
nan, inf¶
Some of the data in the array band appear as nan ('not a number'), which is how, in this dataset, non-valid pixels are specified.
nan is also what you would get for undefined results from arithmetic:
a = np.array([0., 1.])
b = np.array([0., 0.])
c = a/b
print c
print 'so 0./0. =',c[0]
print 'so 1./0. =',c[1]
[ nan inf] so 0./0. = nan so 1./0. = inf
/opt/anaconda/lib/python2.7/site-packages/ipykernel_launcher.py:3: RuntimeWarning: divide by zero encountered in divide This is separate from the ipykernel package so we can avoid doing imports until /opt/anaconda/lib/python2.7/site-packages/ipykernel_launcher.py:3: RuntimeWarning: invalid value encountered in divide This is separate from the ipykernel package so we can avoid doing imports until
inf here means 'infinity', which is what anything other than zero divided by zero is.
We can check to see if some value is nan or inf e.g:
print c
print 'is c nan?',np.isnan(c)
print 'is c inf?',np.isinf(c)
[ nan inf] is c nan? [ True False] is c inf? [False True]
Generally we would want to avoid processing array values that contained nan or inf as any arithmetic involving these is unlikely tell us more than we already know:
print 'inf + 1\t\t=',np.inf + 1
print 'inf - np.inf\t=',np.inf - np.inf
print 'nan / 2 \t\t=',np.nan / 2
inf + 1 = inf inf - np.inf = nan nan / 2 = nan
Or, trying to e.g. calculate the sum of values in the array band:
print data.sum()
nan
which is not really what we wanted.
We have seen above that np.isnan(c) and np.isinf(c) result in a bool array of the same shape as the input array.
We can use these to mask out things we don't want, e.g.
no_data = np.isnan(data)
print no_data
[[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [False False False ..., False False False] [ True False False ..., False False False] [False False False ..., False False False]]
This will be True where the data in band were nan ... but what we might often want is the opposite of this, i.e. True where the data are valid.
We might guess that we could simply type:
valid_data = ~np.isnan(data)
print valid_data
[[False False False ..., False False False] [False False False ..., False False False] [False False False ..., False False False] ..., [ True True True ..., True True True] [False True True ..., True True True] [ True True True ..., True True True]]
where ~ means a logical NOT.
masked array¶
There is a special type of object in numpy called a masked array. With this array representation, operations are only performed where the data mask is False. We need to import the masked array module ma from numpy:
import numpy.ma as ma
masked_band = ma.array(data)
print masked_band.mask
False
When we convert the representation from a normal numpy array to a masked array using ma.array(), the default mask is False (which means no mask).
In this case, we want to set the mask to True where the data are nan. We would normally do this when setting up the masked array:
masked_band = ma.array(data,mask=np.isnan(data))
print "so the array is now\n",masked_band
print "\nwith a mask\n",masked_band.mask
so the array is now [[-- -- -- ..., -- -- --] [-- -- -- ..., -- -- --] [-- -- -- ..., -- -- --] ..., [0.6507693529129028 0.6519480347633362 0.7674967646598816 ..., 0.6532170176506042 0.6534419059753418 0.6529830694198608] [-- 0.6506087779998779 0.6500169038772583 ..., 0.6474758386611938 0.6462251543998718 0.6453946232795715] [0.8676198124885559 0.8676198124885559 0.8676198124885559 ..., 0.8676198124885559 0.8676198124885559 0.8676198124885559]] with a mask [[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [False False False ..., False False False] [ True False False ..., False False False] [False False False ..., False False False]]
3.3.2 Reading data from multiple files¶
zeros, ones, empty¶
Suppose we want to read in all 12 months of SW albedo data following on from the example above.
def read_globalbedo(month, year, folder="data/", layer="BHR_VIS"):
"""Read globalbedo data when given month and year.
Parameters
------------
month: int
The month from 1 to 12.
year: int
The year, currently only 2009 and 2010 available, but can download more.
folder: str
The folder where the GA data are stored.
layer: str
The data layer
Returns
---------
A 2D array"""
file_template = 'NETCDF:"{:s}":{:s}' # Template for the Netcdf path
fname_template = '{:s}/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc' # the actual filename
fname = fname_template.format(folder, year, month)
netcdf_fname = file_template.format(fname, layer)
g = gdal.Open(netcdf_fname)
if g is None:
raise IOError("Problem with reading file {}".format(fname))
data = g.ReadAsArray()
return data
# which months?
months = range(1,13)
# loop over month
# but right now we'll just pretend that we're
# doing that and consider what happens the first time
# we are in our loop
#
# set i as index counter
# set the variable month to be
# months[0]
i = 0
band = read_globalbedo(i+1,2009, layer='DHR_SW')
Looking at what would happen when we read the first file, we can see that we don't actually know how large the whole dataset will be until we have read our first file.
We know that we will want dimensions of (12,nrows,ncols) (or better still, (len(months),nrows,ncols) in case we decide to change months), but until we have read the data into the variable band with:
band = readGA(month=months[i],layer='DHR_SW')
we don't know how many rows or columns the dataset has (well, we might do ... but we want to try to design flexible code, where if the size of the datasets changed, our code would still operate correctly!).
But, when we have first read some data, then we would be in a position to say what the shape of the whole dataset should be. At that point, we could use the method:
np.empty(shape)
to (in essence) set aside ('allocate') some memory to store these data. There are three main options for this:
np.zeros(shape): set up an array with shape defined byshapeand initialise with 0np.ones(shape): set up an array with shape defined byshapeand initialise with 1np.empty(shape): set up an array with shape defined byshapebut don't initialise (so, in theory, a little faster, but can be filled with randon junk)
we can control the dtype of the array with a dtype option. e.g.:
print '-'*30;print '1-D arrays';print '-'*30;
print 'np.zeros(1):',np.zeros(1) # 1 : 1D array
print 'np.zeros(2):',np.zeros(2) # 2 : 1D array
print '\n' + '-'*30;print '2- and 3-D arrays';print '-'*30;
print 'np.zeros((1,2)):\n',np.zeros((1,2)) # default dtype is float
print 'np.zeros((2,2)):\n',np.zeros((2,2)) # 2 X 2
print 'np.zeros((2,2,3)):\n',np.zeros((2,2,3)) # 2 X 2 X 3
print '\n' + '-'*30;print 'empty and ones';print '-'*30;
print 'np.empty((1,2)):\n',np.empty((1,2)) # default dtype is float
print 'np.ones((1,2)):\n',np.ones((1,2)) # default dtype is float
print '\n' + '-'*30;print 'Changing the data type';print '-'*30;
print 'np.ones((1,2),dtype=int):\n',np.ones((1,2),dtype=int) # dtype int
print 'np.empty((1,2),dtype=int):\n',np.empty((1,2),dtype=int) # dtype int
print 'np.ones((1,2),dtype=bool):\n',np.ones((1,2),dtype=bool) # dtype bool
print 'np.zeros((1,2),dtype=bool):\n',np.zeros((1,2),dtype=bool) # dtype bool
print 'np.empty((1,2),dtype=bool):\n',np.empty((1,2),dtype=bool) # dtype bool
print '\n' + '-'*30;print 'Note that we can have a string array';print '-'*30;
print 'np.ones((1,2),dtype=str):\n',np.ones((1,2),dtype=str) # dtype str
print '\n' + '-'*30;print 'Or even e.g. a list array';print '-'*30;
print 'np.ones((2,2),dtype=list):\n',np.ones((2,2),dtype=list) # dtype list
print 'np.empty((2,2),dtype=list):\n',np.empty((2,2),dtype=list) # dtype list
------------------------------ 1-D arrays ------------------------------ np.zeros(1): [ 0.] np.zeros(2): [ 0. 0.] ------------------------------ 2- and 3-D arrays ------------------------------ np.zeros((1,2)): [[ 0. 0.]] np.zeros((2,2)): [[ 0. 0.] [ 0. 0.]] np.zeros((2,2,3)): [[[ 0. 0. 0.] [ 0. 0. 0.]] [[ 0. 0. 0.] [ 0. 0. 0.]]] ------------------------------ empty and ones ------------------------------ np.empty((1,2)): [[ 0. 0.]] np.ones((1,2)): [[ 1. 1.]] ------------------------------ Changing the data type ------------------------------ np.ones((1,2),dtype=int): [[1 1]] np.empty((1,2),dtype=int): [[1 1]] np.ones((1,2),dtype=bool): [[ True True]] np.zeros((1,2),dtype=bool): [[False False]] np.empty((1,2),dtype=bool): [[False False]] ------------------------------ Note that we can have a string array ------------------------------ np.ones((1,2),dtype=str): [['1' '1']] ------------------------------ Or even e.g. a list array ------------------------------ np.ones((2,2),dtype=list): [[1 1] [1 1]] np.empty((2,2),dtype=list): [[None None] [None None]]
So, once we have read one band, we could write:
def read_globalbedo(month, year, folder="data/", layer="BHR_VIS"):
"""Read globalbedo data when given month and year.
Parameters
------------
month: int
The month from 1 to 12.
year: int
The year, currently only 2009 and 2010 available, but can download more.
folder: str
The folder where the GA data are stored.
layer: str
The data layer
Returns
---------
A 2D array"""
file_template = 'NETCDF:"{:s}":{:s}' # Template for the Netcdf path
# the actual filename
fname_template = '{:s}/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc'
fname = fname_template.format(folder, year, month)
netcdf_fname = file_template.format(fname, layer)
g = gdal.Open(netcdf_fname)
if g is None:
raise IOError("Problem with reading file {}".format(fname))
data = g.ReadAsArray()
return data
# Loop over all months
for month in range(12):
# Read data for current month. NOte that read_globalbedo
# takes months from 1 to 12, not 0 to 11!
this_data = read_globalbedo(month + 1, 2009, layer="DHR_SW")
# If we're on the first month...
if month == 0:
# Get the array shape (rows, cols)
ny, nx = this_data.shape
# Create an empty 3D array, with 12 "layers" of 2D arrays
data = np.empty((12, ny, nx))
# data is created on the first iteration, so it's always available
# Stash the new data into the output
data[month, :, :] = this_data
print data.shape
(12, 360, 720)
We also use slicing when assigning ('loading', if you like) data into a larger array:
data[i,:,:] = band
But this will only work if what you are trying to load and where you are tryng to load it are the same shape:
data[i,:,:].shape == band.shape
which they are in this case.
An alternative way of achieving this same effect is to use a list, creating an empty list before going into the loop and appending band to the list each time we read a new band.
# Start with an empty list
data = []
# Loop over all months
for month in range(12):
# Read data for current month. NOte that read_globalbedo
# takes months from 1 to 12, not 0 to 11!
this_data = read_globalbedo(month + 1, 2009, layer="DHR_SW")
data.append(this_data)
data = np.array(data)
print 'shape', data.shape
shape (12, 360, 720)
In many ways, this is a simpler and 'cleaner' way of setting up a 3-D array from reading multiple 2-D arrays. If the array is very large, it can be less efficient (you have to convert a huge list to an array at the end), but that is not normally something to worry about and this approach is often preferable.
In fact, if we know how many entries (in time) we are expecting, we can simply set up the storage array before going into the loop:
# Start with an empty array of the same size
data = np.empty((12, 360, 720))
# Loop over all months
for month in range(12):
# Read data for current month. NOte that read_globalbedo
# takes months from 1 to 12, not 0 to 11!
this_data = read_globalbedo(month + 1, 2009, layer="DHR_SW")
data[month, :, :] = this_data
print 'shape', data.shape
shape (12, 360, 720)
1. Writing a completely new method
2. Writing a new `read_globalbedo_annual` method that builds up from `read_globalbedo`
sum¶
To sum data in a numpy (or masked) array, we use the function sum.
e.g.:
print data.sum(),data[0,0,0]
nan nan
Thats not very useful in this case, because we have nan in the dataset ... though if we use a masked array it will work as we expect.
np.sum here adds all of the values in the array.
We could select only values that are >= 0 with the logical statement:
not_valid = np.isnan(data)
print not_valid
[[[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [False False False ..., False False False] [ True False False ..., False False False] [False False False ..., False False False]] [[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [False False False ..., False False False] [ True False False ..., False False False] [False False False ..., False False False]] [[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [ True True True ..., True True True] [ True True True ..., True True True] [False False False ..., False False False]] ..., [[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [False False False ..., False False False] [ True False False ..., False False False] [False False False ..., False False False]] [[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [False False False ..., False False False] [ True False False ..., False False False] [False False False ..., False False False]] [[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [False False False ..., False False False] [ True False False ..., False False False] [False False False ..., False False False]]]
so not_valid here is an array of the same size and shape as data, with True where the value is not valid (nan) and False elsewhere.
We can use this to select only data elements in data which are False in not_valid (not not_valid is valid ...), or we could change this into a valid array:
valid = not_valid == False
ndata = data[valid]
print ndata
[ 0.79814297 0.74479127 0.82455897 ..., 0.75297731 0.75297731 0.75297731]
Now, the array is a 1-D array. It has fewer elements that the original data array:
print 'original data size\t:',data.size
print 'new data size\t\t:',ndata.size
print 'data shape\t\t:',data.shape
print 'new data shape\t\t:',ndata.shape
original data size : 3110400 new data size : 1121705 data shape : (12, 360, 720) new data shape : (1121705,)
This is however, one way we can use np.sum to add up the values in data:
print data[valid].sum()
463425.186595
mean¶
Similarly, if we wanted the mean or standard deviation and we tried to do this on the array that has nan in it:
print data.mean(),data.std()
nan nan
but selecting only the values that are valid:
print data[valid].mean(),data[valid].std()
0.413143550751 0.308557692958
we get what we might expect.
Remember that this is for normal (i.e. not masked) numpy arrays. Hassles of this nature with nan in the dataset is one good reason to use masked arrays.
axis selction¶
As used above, functions such as sum, mean and std will give you the sum, mean and standard deviation over the whole array.
Often though, we only want to apply these functions over a particular dimension.
Recall that the shape of data is:
data.shape
(12, 360, 720)
The first dimension here is 'month' then, the second is 'latitude' and the third 'longitude'.
If we wanted to know the mean albedo for each month then then, we would want to apply mean over axis 1 and 2:
In numpy, this is done by specifying axis in the function:
month_mean = data.mean(axis=(1,2))
print month_mean.shape
(12,)
Thats the correct shape, but if we look at the values:
print month_mean
[ nan nan nan nan nan nan nan nan nan nan nan nan]
we see that we get nan ...
That's hardly surprising, as this is the same issue we had earlier with just calling sum or mean over the whole array.
Rather than end this section with a negative result, lets consider an array without nan.
We have previously generated the array valid, which is a logical array, but we could e.g. convert this to float:
gdata = (valid).astype(float)
print gdata.sum(axis=(1,2))
print gdata.std(axis=(1,2))
print gdata.mean(axis=(1,2))
[ 94781. 94642. 92534. 92549. 92584. 92587. 92576. 92540. 92546. 94785. 94790. 94791.] [ 0.48161682 0.48146692 0.47911435 0.47913162 0.47917188 0.47917533 0.47916268 0.47912126 0.47912816 0.48162113 0.4816265 0.48162758] [ 0.36566744 0.36513117 0.35699846 0.35705633 0.35719136 0.35720293 0.35716049 0.3570216 0.35704475 0.36568287 0.36570216 0.36570602]
There are many useful functions in numpy for manipulating array data, from basic arithmetic through to these function here for some basic stats.
They are all very easy to use and make for clear code, but if you have masks in your dataset (as we often do with geospatial data), you need to think carefully about how you want to process in the presence of masks.
It is often a good idea when you have masks to use a masked array.
E3.3 Exercise: 3D Masked Array¶
Taking the code above as a starting point, generate a masked array of the GlobAlbedo dataset for the year 2009.
3.4 Average Albedo¶
If you run the exercise above, data should now be a masked array.
If you haven't (e.g. when going through this in the class), you can run a function in the file python/masked.py to do this now:
import sys
# look in the local python (code) directory
sys.path.insert(0,'python')
from masked import masked
import numpy as np
import numpy.ma as ma
import gdal
def masked(months=range(1, 13), years=[2009], folder="data/", layer="BHR_VIS"):
"""Read globalbedo data when given month and year.
Parameters
------------
month: int
The month from 1 to 12.
year: int
The year, currently only 2009 and 2010 available, but can download more.
folder: str
The folder where the GA data are stored.
layer: str
The data layer
Returns
---------
A 2D array"""
data = []
file_template = 'NETCDF:"{:s}":{:s}' # Template for the Netcdf path
# the actual filename
fname_template = '{:s}/GlobAlbedo.merge.albedo.05.{:d}{:02d}.nc'
for year in years:
for month in months:
fname = fname_template.format(folder, year, month)
netcdf_fname = file_template.format(fname, layer)
g = gdal.Open(netcdf_fname)
if g is None:
raise IOError("Problem with reading file {}".format(fname))
the_data = g.ReadAsArray()
masked_data = np.ma.array(the_data,mask=np.isnan(the_data))
data.append(masked_data)
output_data = np.ma.array(data)
return output_data
data = masked(layer='BHR_SW')
print type(data)
print data.shape
print data.ndim
<class 'numpy.ma.core.MaskedArray'> (12, 360, 720) 3
Now we have the albedo dataset for one year stacked in a 3-D masked array array (assuming you sucessfully completed exercise E3.3 or ran the code above), we can do some interesting things with it with numpy (without the nan hassles above):
3.4.1 Annual Mean and Standard Deviation¶
# calculate the mean albedo over the year
# The axis=0 option tells numpy which
# dimension to apply the function mean() over
mean_albedo = data.mean(axis=0)
# just confirm this is the ndim and shape we expect ...
print mean_albedo.ndim
print mean_albedo.shape
2 (360, 720)
# do a plot
# change the figure size
plt.figure(figsize=(10,4))
plt.title('SW BHR mean albedo for %d'%(year))
# use nearest neighbour interpolation
plt.imshow(mean_albedo,interpolation='none',\
cmap=plt.get_cmap('Spectral'),vmin=0.0,vmax=1.0)
# show a colour bar
plt.colorbar()
<matplotlib.colorbar.Colorbar at 0x7f41492e87d0>
#std
std_albedo = data.std(axis=0)
plt.figure(figsize=(10,4))
plt.title('SW BHR std albedo for %d'%(year))
# use nearest neighbour interpolation
plt.imshow(std_albedo,interpolation='none',\
cmap=plt.get_cmap('Spectral'),vmin=0.0, vmax=0.5)
# show a colour bar
plt.colorbar()
<matplotlib.colorbar.Colorbar at 0x7f4149758610>
We can see form this that most of the intra-annual variation in total shortwave albedo is at high latitudes (but not Greenland or Antarctica, for instance).
Why do you think that is?
3.4.2 NDVI¶
It might be interesting to look at NDVI data in this same way and to examine a few more stats using numpy. Even though this is a broadband albedo dataset, we would expect NDVI to give some indication of vegetation activity.
min, max¶
#import sys
#sys.path.insert(0,'files/python')
#from masked import masked
vis = masked(layer='BHR_VIS')
nir = masked(layer='BHR_NIR')
ndvi = (nir-vis)/(nir+vis)
# set negative and ndvi > 1 (error) to 0
#ndvi[ndvi>1] = 0
#ndvi[ndvi<0] = 0
mean_ndvi = ndvi.mean(axis=0)
max_ndvi = ndvi.max(axis=0)
min_ndvi = ndvi.min(axis=0)
range_ndvi = max_ndvi - min_ndvi
std_ndvi = ndvi.std(axis=0)
# set up a dictionary
datasets = {"mean NDVI 2009":mean_ndvi,\
"max NDVI 2009":max_ndvi,\
"min NDVI 2009":min_ndvi,\
"NDVI range 2009":range_ndvi,\
"std NDVI 2009":std_ndvi}
vmax = {"mean NDVI 2009":1.0,\
"max NDVI 2009":1.,\
"min NDVI 2009":1.,\
"NDVI range 2009":1.,\
"std NDVI 2009":0.25}
# plotting loop
for k in datasets.keys():
# note the order is not maintained
# from when we set up the dictionary
plt.figure(figsize=(10,4))
plt.title(k)
plt.imshow(datasets[k],interpolation='none',\
cmap=plt.get_cmap('YlGn'),vmin=0.0,vmax=vmax[k])
plt.colorbar()
The mean and max NDVI is pretty much what you would expect (MODIS NDVI, Nov-Dec, 2007 shown here):

but again we see the large variations in NDVI in the Northern Hemisphere.
3.5 Solar Radiation¶
One common use for albedo data is to calculate the amount of absorbed solar radiation at the surface.
In this section, we will build a model and dataset of downwelling solar radiation.
We can build a simple model of solar radiation:
Incoming solar radiation, $A$, ignoring the effect of the atmosphere, clouds, etc is given by
$$ A = E_{0}\sin\theta $$where $E_{0}$ is the solar constant (given as $1360Wm^{-2}$), and $\theta$ is the solar elevation angle. The solar elevation angle is approximately given by $$ \sin \theta = \cos h \cos \delta \cos \varphi + \sin \delta \sin \varphi $$
where $h$ is the hour angle, $\delta$ is the solar declination angle and $\varphi$ is the latitude. The solar declination can be approximated by
$$ \delta = - \arcsin \left [ 0.39779 \cos \left ( 0.98565 \left (N + 10 \right ) + 1.914 \sin \left ( 0.98565 \left ( N - 2 \right ) \right ) \right ) \right ] $$where $N$ is the day of year beginning with N=0 at 00:00:00 UTC on January 1
We have the latitude associated with the dataset from knowing that latitudes stretch from -90 to +90 degrees at 0.5 degree intervals, and longitudes go from -180 to 180 degrees at 0.5 degree intervals. We can use the meshgrid method to create the two 2D arrays from vectors of the dimensions.
We note that one of the big advantages of using arrays is that we can do operations on many grid cells simultaneously just by virtue of using arrays of the right shape(s). In this case, we will apply a series of calculations to all the grid cells in the albedo dataset in just one go. We could do this with loops, but netsting loops in Python for large datasets as is the case here is very inefficient. Numpy arrays allow us to do away with all this.
lon, lat= np.meshgrid(np.arange(-180 + 0.25, 180+0.25, 0.5),
np.arange(90 - 0.25, -90, -0.5))
print lat.shape
print lon.shape
(360, 720) (360, 720)
Quick sanity check plotting: longitude should increase from left to right, and latitutde should increase from bottom to top:
plt.imshow(lon, interpolation="nearest")
plt.figure()
plt.imshow(lat, interpolation="nearest")
<matplotlib.image.AxesImage at 0x7f414bc7e610>
# The boolean mask is stored as vis.mask
print 'shape of vis.mask',vis.mask.shape
# we want a single time slice so mask[0]
print vis.mask[0].shape,lat.shape
lat = ma.array(lat,mask=vis.mask[0])
print lat.shape
plt.figure(figsize=(10,4))
plt.imshow(lat)
plt.title('latitude')
plt.colorbar()
shape of vis.mask (12, 360, 720) (360, 720) (360, 720) (360, 720)
<matplotlib.colorbar.Colorbar at 0x7f41486f39d0>
for day of year N then, we have a formula for solar declination.
Approximating N for each month by an average:
# mean days per month
av_days = 365.25 / 12.
half = av_days/2.
N = np.arange(half,365.25,av_days)
print N
[ 15.21875 45.65625 76.09375 106.53125 136.96875 167.40625 197.84375 228.28125 258.71875 289.15625 319.59375 350.03125]
rad2deg, deg2rad, cos, sin, arcsin, arccos, pi¶
we can approximate the solar declination as:
t0 = np.deg2rad (0.98565*(N-2))
t1 = 0.39779*np.cos( np.deg2rad ( 0.98565*(N+10) + 1.914*np.sin ( t0 ) ) )
delta = -np.arcsin ( t1 )
plt.plot(N,np.rad2deg(delta))
plt.xlabel('day of year')
plt.ylabel('solar declination angle (degrees)')
plt.xlim(0,365.25)
(0, 365.25)
Although we might think the formulae a little complicated, the translation from the formula to numpy code is really very clear and readable. In particular, by having N as an array object, we are able to apply the formulae to the array and not have to obfuscate the code with loops.
Note that the formulae above are given in degrees, but trignometric functions in most programming languages work in radians.
In numpy, we can use the functions deg2rad and rad2deg to convert between degrees and radians and vice-versa (alternatively, multiply by np.pi/180. and 180./np.pi respectively).
The trig functions cosine and sine are cos and sin respectively, and their inverses arccos and arcsin.
For our solar energy calculations, we will set the 'hour angle' to zero (this is solar noon), so:
h = 0.0
To estimate the solar elevation angle then, we need to include the latitude. At present, we have an array lat for latitude and N for day of year:
print 'lat:',lat.shape
print 'N:',N.shape
lat: (360, 720) N: (12,)
what we need is lat and N in the full 3-D array shape (12,360,720).
There are various ways to do this, but a simple one is to just use np.mlist with an extra (time) dimension:
dd = 0.5
month,lat,lon = np.mgrid[0:12,-90+dd/2:90+dd/2:dd,-180+dd/2:180+dd/2:dd]
print '------ latitude ------'
print lat.shape
print '------ longitude ------'
print lon.shape
------ latitude ------ (12, 360, 720) ------ longitude ------ (12, 360, 720)
def: functions¶
At this point, we might consider it useful to write a function. We have used an example of this above, but go into it in more detail now.
These are of the form:
def fn_name(args):
...
return value
def declination(N):
'''Calculate solar declination (in degrees)'''
t0 = np.deg2rad (0.98565*(N-2))
t1 = 0.39779*np.cos( np.deg2rad ( 0.98565*(N+10) + 1.914*np.sin ( t0 ) ) )
delta = -np.arcsin ( t1 )
return np.rad2deg(delta)
def solar_elevation(delta,h,lat):
'''solar elevation in degrees'''
lat = np.deg2rad(lat)
delta = np.deg2rad(delta)
h = np.deg2rad(h)
sin_theta = np.cos (h)*np.cos (delta)*np.cos(lat) + np.sin ( delta)*np.sin(lat)
return np.rad2deg(np.arcsin(sin_theta))
# create numpy arrays of the right shape
# using meshgrid for simplicity
N, lat, lon = np.meshgrid(np.arange(half, 365.25, av_days),
np.arange(90 - 0.25, -90, -0.5),
np.arange(-180 + 0.25, 180+0.25, 0.5), indexing="ij")
# zeros_like creates array of the same shape as N here (or, lat or lon)
# filled with zeros. Similarly ones_like
h2 = np.zeros_like(N) + h
# now put these things together
# copy the arrays, so we don't overwrite the data
delta = declination(N)
e0 = 1360.
sea = solar_elevation(delta,h2,lat)
sin_theta = np.sin(np.deg2rad(sea))
rad = e0*sin_theta
# threshold at zero
rad[rad < 0] = 0.0
incoming_rad = rad
# now mask this with data mask
rad = ma.array(incoming_rad,mask=data.mask)
# plot to visualise the data and see it looks reasonable
plt.figure(figsize=(10,4))
plt.title('Jan')
plt.imshow(rad[0],interpolation='none',cmap=plt.get_cmap('jet'))
plt.colorbar()
plt.figure(figsize=(10,4))
plt.title('Jun')
plt.imshow(rad[5],interpolation='none',cmap=plt.get_cmap('jet'))
plt.colorbar()
<matplotlib.colorbar.Colorbar at 0x7f41499f4110>
This sort of application really shows the power of manipulating arrays.
E3.5 Exercise: Solar Radiation Code: A chance for you to get feedback¶
In the section above, we have developed some code to estimate solar radiation and we have packaged some of it into functions.
In a file called python/solar.py, develop some Python code that will
- Read GlobAlbedo data
DHR_SW(shortwave directional-hemispherical reflectance) into a masked arraydata - Generate a spatial Solar Radiation dataset
radof the same shape asdata - Calculate the amount of solar radiation absorbed at the surface (W m^-2)
- Calculate the latitudinal total absorbed radiation for each month (W m^-2)
- Calculate the global total absorbed radiation power for each month (W)
Where appropriate, produce images, graphs and/or movies of your results.
Comment on the patterns of absorbed solar radiation that you see.
All of your code should be commented as well as possible.
It should indicate which parts of the code you have simply taken from the material above and which parts you have developed yourself.
If possible, use an ipython notebook to illustrate your code and its operation. As a miniumum, submit your code and examples of its output.
If possible, wrap your codes into functions to group things that belong together together.
You will probably want to save the arrays you generated (np.savez) so that you can show your results later.
Hints:
- All of the code you need for parts 1. and 2. can be found in the material above ... you just need to put it together
- Recall that albedo is the proportion of radiation reflected.
- Recall that in a lat/long projection the area represented by each pixel varies as a function of latitude.
We will not release model answers for this exercise at present.
Instead, we would like you to complete this exercise over the coming week and submit it by the following Monday morning. If you do this, there will be sessions run in (extended) office hours (11-12 Thursday, 4-5 Thursday, 9-10 Wednesday) when you will bve given feedback on your submission. To get the most out of these feedback sessions, it would be adviseable to request a time slot (You should be getting a doodle poll email to that effect)
We strongly recommend that you attempt this exercise on your own. Given the amount of material you are given here, this should be achievable for all of you.
This work does not form part of your formal assessment. Instead it is intended to provide you with an opportunity for feedback on how you are getting on in the course at an early stage. So, you are not required to do this. If you do not though, we cannot give yuou feedback.
Come along to the feedback session prepared to explain how your code operates and how to run it (one reason why a notebook would be useful if you can manage that).
3.6 Saving and loading numpy data¶
savez, load¶
When we have gone to lots of effort (well, lots of processing time) to do some calculation, we probably want to save the result.
In numpy, we can save one or more numpy arrays (and similar data types) with np.savez().
So, e.g. to save the arrays data and rad:
import numpy as np
# there is a problem saving masked arrays at the moment,
# so convert and save the mask
np.savez('data/solar_rad_data.npz',\
rad=np.array(rad),data=np.array(data),mask=data.mask)
To load this again use np.load, which returns a dictionary:
f = np.load('data/solar_rad_data.npz')
data = ma.array(f['data'],mask=f['mask'])
rad = ma.array(f['rad'],mask=f['mask'])
loadtxt, savetxt¶
As an aside here, we note that numpy also has convenient functions for loading ASCII text (which is much simplerto use than the methods we learned last session, but you need to know how to do it from scratch if the data format is very awkward).
That said, we can deal with some awkward issues in a dataset:
!head -15 < data/heathrowdata.txt
Heathrow (London Airport)
Location 5078E 1767N 25m amsl
Estimated data is marked with a * after the value.
Missing data (more than 2 days missing in month) is marked by ---.
Sunshine data taken from an automatic Kipp & Zonen sensor marked with a #, otherwise sunshine data taken from a Campbell Stokes recorder.
yyyy mm tmax tmin af rain sun
degC degC days mm hours
1948 1 8.9 3.3 --- 85.0 ---
1948 2 7.9 2.2 --- 26.0 ---
1948 3 14.2 3.8 --- 14.0 ---
1948 4 15.4 5.1 --- 35.0 ---
1948 5 18.1 6.9 --- 57.0 ---
1948 6 19.1 10.3 --- 67.0 ---
1948 7 21.7 12.0 --- 21.0 ---
1948 8 20.8 11.7 --- 67.0 ---
# read sun hours from the heathrow data file
filename = 'data/heathrowdata.txt'
# In this dataset, we need to know what
# to do when we get '---' in the dataset
# so we set up a
# translation dictionary
def awkward(x):
if x == '---':
return np.nan
return x
trans = {6: awkward}
# skip the first 7 rows
# use the awkward translation for column 6
# unpack gets the shape the right way around
sun_hrs = np.loadtxt(filename,skiprows=7,usecols=[0,1,6],converters=trans,unpack=True)
print sun_hrs.shape
# plot decimal year and sun hours
plt.plot(sun_hrs[0]+sun_hrs[1]/12.,sun_hrs[2])
# zoom in to 1970 to 1990
plt.xlim(1970,1990)
(3, 785)
(1970, 1990)
We can save this to a text file with:
header = 'year month sun_hours'
# note, we transpose it if we want the data as columns
np.savetxt('data/sunshine.txt',sun_hrs.T,header=header)
!head -10 < data/sunshine.txt
# year month sun_hours 1.948000000000000000e+03 1.000000000000000000e+00 nan 1.948000000000000000e+03 2.000000000000000000e+00 nan 1.948000000000000000e+03 3.000000000000000000e+00 nan 1.948000000000000000e+03 4.000000000000000000e+00 nan 1.948000000000000000e+03 5.000000000000000000e+00 nan 1.948000000000000000e+03 6.000000000000000000e+00 nan 1.948000000000000000e+03 7.000000000000000000e+00 nan 1.948000000000000000e+03 8.000000000000000000e+00 nan 1.948000000000000000e+03 9.000000000000000000e+00 nan
E3.6 Exercise: awkward reading¶
The file data/elevation.dat look like:
!head -10 < data/elevation.dat
2013/10/8 00:00:00 -44.2719952943 2013/10/8 00:30:00 -43.5276412785 2013/10/8 00:59:59 -41.9842746582 2013/10/8 01:30:00 -39.7226999863 2013/10/8 02:00:00 -36.8452198361 2013/10/8 02:30:00 -33.459799008 2013/10/8 03:00:00 -29.6691191507 2013/10/8 03:30:00 -25.5652187709 2013/10/8 03:59:59 -21.2281801291 2013/10/8 04:30:00 -16.7272357302
Write some code to read these data in using np.loadtxt
Hint: You will need to write a function that decodes column 0 and 1, for example:
def col0(x):
# interpret column 1 as a sort of
# decimal year
# NB this is very rough ...
# not all months have 30 days ...
y,m,d = x.split('/')
dec_year = y + (m-1)/12. + (d-1)/30.
return dec_year
# translation ... just for column 1 here
# you'd need to define your own for column 2
trans = {0 : col0}
Advanced and Answers¶
If you feel comfortable with the contents of this session, or simply would like something a little more challenging, examine the notes in the advanced section.
Answers for the exercises in this session are available, as are the answers for the advanced section.