E3 Answers to Exercise¶
E3.1 Exercise: listing¶
Using Python, produce a listing of the files in the subdirectory data of geogg122/Chapter3_Scientific_Numerical_Python that end with .nc and put this listing in a file called data/data.dat with each entry on a different line
A3.1 Answer: listing¶
Hopefully, you should already be in the directory geogg122/Chapter3_Scientific_Numerical_Python, if not, you may like to go there before starting this exercise.
If you were to do this from unix, you would get the listing with:
ls -l data/*.nc
-rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200901.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200902.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200903.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200904.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200905.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200906.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18758652 Sep 30 11:56 data/GlobAlbedo.200907.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200908.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200909.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200910.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200911.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.200912.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18758652 Sep 30 11:56 data/GlobAlbedo.201001.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201002.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201003.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201004.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201005.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201006.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201007.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18758652 Sep 30 11:56 data/GlobAlbedo.201008.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201009.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201010.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201011.mosaic.5.nc -rw-rw-r--. 1 plewis plewis 18669672 Sep 30 11:56 data/GlobAlbedo.201012.mosaic.5.nc
or similar.
To do this in Python, you should use glob, and the same 'pattern':
import glob
files = glob.glob('data/*.nc')
print files
['data/GlobAlbedo.201008.mosaic.5.nc', 'data/GlobAlbedo.200903.mosaic.5.nc', 'data/GlobAlbedo.200905.mosaic.5.nc', 'data/GlobAlbedo.201007.mosaic.5.nc', 'data/GlobAlbedo.201011.mosaic.5.nc', 'data/GlobAlbedo.200902.mosaic.5.nc', 'data/GlobAlbedo.201006.mosaic.5.nc', 'data/GlobAlbedo.201004.mosaic.5.nc', 'data/GlobAlbedo.200904.mosaic.5.nc', 'data/GlobAlbedo.200911.mosaic.5.nc', 'data/GlobAlbedo.201012.mosaic.5.nc', 'data/GlobAlbedo.201005.mosaic.5.nc', 'data/GlobAlbedo.200912.mosaic.5.nc', 'data/GlobAlbedo.201003.mosaic.5.nc', 'data/GlobAlbedo.200907.mosaic.5.nc', 'data/GlobAlbedo.200909.mosaic.5.nc', 'data/GlobAlbedo.201001.mosaic.5.nc', 'data/GlobAlbedo.201009.mosaic.5.nc', 'data/GlobAlbedo.201010.mosaic.5.nc', 'data/GlobAlbedo.200906.mosaic.5.nc', 'data/GlobAlbedo.201002.mosaic.5.nc', 'data/GlobAlbedo.200908.mosaic.5.nc', 'data/GlobAlbedo.200901.mosaic.5.nc', 'data/GlobAlbedo.200910.mosaic.5.nc']
This is a list. We want to write this to a file called data/data.dat.
First we open it, then simply use writelines to write the list of strings, the close the file.
filename = 'data/data.dat'
# open in write mode
fp = open(filename,'w')
fp.writelines(files)
fp.close()
Hopefully, that all worked well, but just to check from unix:
cat data/data.dat
data/GlobAlbedo.201008.mosaic.5.ncdata/GlobAlbedo.200903.mosaic.5.ncdata/GlobAlbedo.200905.mosaic.5.ncdata/GlobAlbedo.201007.mosaic.5.ncdata/GlobAlbedo.201011.mosaic.5.ncdata/GlobAlbedo.200902.mosaic.5.ncdata/GlobAlbedo.201006.mosaic.5.ncdata/GlobAlbedo.201004.mosaic.5.ncdata/GlobAlbedo.200904.mosaic.5.ncdata/GlobAlbedo.200911.mosaic.5.ncdata/GlobAlbedo.201012.mosaic.5.ncdata/GlobAlbedo.201005.mosaic.5.ncdata/GlobAlbedo.200912.mosaic.5.ncdata/GlobAlbedo.201003.mosaic.5.ncdata/GlobAlbedo.200907.mosaic.5.ncdata/GlobAlbedo.200909.mosaic.5.ncdata/GlobAlbedo.201001.mosaic.5.ncdata/GlobAlbedo.201009.mosaic.5.ncdata/GlobAlbedo.201010.mosaic.5.ncdata/GlobAlbedo.200906.mosaic.5.ncdata/GlobAlbedo.201002.mosaic.5.ncdata/GlobAlbedo.200908.mosaic.5.ncdata/GlobAlbedo.200901.mosaic.5.ncdata/GlobAlbedo.200910.mosaic.5.nc
which isn't quite what we wanted: we need to insert a newline character at the end of each string before writing.
There are several ways to do this, e.g.:
files = glob.glob('data/*.nc')
for i,file in enumerate(files):
files[i] = file + '\n'
print files
['data/GlobAlbedo.201008.mosaic.5.nc\n', 'data/GlobAlbedo.200903.mosaic.5.nc\n', 'data/GlobAlbedo.200905.mosaic.5.nc\n', 'data/GlobAlbedo.201007.mosaic.5.nc\n', 'data/GlobAlbedo.201011.mosaic.5.nc\n', 'data/GlobAlbedo.200902.mosaic.5.nc\n', 'data/GlobAlbedo.201006.mosaic.5.nc\n', 'data/GlobAlbedo.201004.mosaic.5.nc\n', 'data/GlobAlbedo.200904.mosaic.5.nc\n', 'data/GlobAlbedo.200911.mosaic.5.nc\n', 'data/GlobAlbedo.201012.mosaic.5.nc\n', 'data/GlobAlbedo.201005.mosaic.5.nc\n', 'data/GlobAlbedo.200912.mosaic.5.nc\n', 'data/GlobAlbedo.201003.mosaic.5.nc\n', 'data/GlobAlbedo.200907.mosaic.5.nc\n', 'data/GlobAlbedo.200909.mosaic.5.nc\n', 'data/GlobAlbedo.201001.mosaic.5.nc\n', 'data/GlobAlbedo.201009.mosaic.5.nc\n', 'data/GlobAlbedo.201010.mosaic.5.nc\n', 'data/GlobAlbedo.200906.mosaic.5.nc\n', 'data/GlobAlbedo.201002.mosaic.5.nc\n', 'data/GlobAlbedo.200908.mosaic.5.nc\n', 'data/GlobAlbedo.200901.mosaic.5.nc\n', 'data/GlobAlbedo.200910.mosaic.5.nc\n']
files = glob.glob('data/*.nc')
# or:
files = [file + '\n' for file in files]
print files
['data/GlobAlbedo.201008.mosaic.5.nc\n', 'data/GlobAlbedo.200903.mosaic.5.nc\n', 'data/GlobAlbedo.200905.mosaic.5.nc\n', 'data/GlobAlbedo.201007.mosaic.5.nc\n', 'data/GlobAlbedo.201011.mosaic.5.nc\n', 'data/GlobAlbedo.200902.mosaic.5.nc\n', 'data/GlobAlbedo.201006.mosaic.5.nc\n', 'data/GlobAlbedo.201004.mosaic.5.nc\n', 'data/GlobAlbedo.200904.mosaic.5.nc\n', 'data/GlobAlbedo.200911.mosaic.5.nc\n', 'data/GlobAlbedo.201012.mosaic.5.nc\n', 'data/GlobAlbedo.201005.mosaic.5.nc\n', 'data/GlobAlbedo.200912.mosaic.5.nc\n', 'data/GlobAlbedo.201003.mosaic.5.nc\n', 'data/GlobAlbedo.200907.mosaic.5.nc\n', 'data/GlobAlbedo.200909.mosaic.5.nc\n', 'data/GlobAlbedo.201001.mosaic.5.nc\n', 'data/GlobAlbedo.201009.mosaic.5.nc\n', 'data/GlobAlbedo.201010.mosaic.5.nc\n', 'data/GlobAlbedo.200906.mosaic.5.nc\n', 'data/GlobAlbedo.201002.mosaic.5.nc\n', 'data/GlobAlbedo.200908.mosaic.5.nc\n', 'data/GlobAlbedo.200901.mosaic.5.nc\n', 'data/GlobAlbedo.200910.mosaic.5.nc\n']
# or all at once if you like:
files = [file + '\n' for file in glob.glob('data/*.nc')]
print files
['data/GlobAlbedo.201008.mosaic.5.nc\n', 'data/GlobAlbedo.200903.mosaic.5.nc\n', 'data/GlobAlbedo.200905.mosaic.5.nc\n', 'data/GlobAlbedo.201007.mosaic.5.nc\n', 'data/GlobAlbedo.201011.mosaic.5.nc\n', 'data/GlobAlbedo.200902.mosaic.5.nc\n', 'data/GlobAlbedo.201006.mosaic.5.nc\n', 'data/GlobAlbedo.201004.mosaic.5.nc\n', 'data/GlobAlbedo.200904.mosaic.5.nc\n', 'data/GlobAlbedo.200911.mosaic.5.nc\n', 'data/GlobAlbedo.201012.mosaic.5.nc\n', 'data/GlobAlbedo.201005.mosaic.5.nc\n', 'data/GlobAlbedo.200912.mosaic.5.nc\n', 'data/GlobAlbedo.201003.mosaic.5.nc\n', 'data/GlobAlbedo.200907.mosaic.5.nc\n', 'data/GlobAlbedo.200909.mosaic.5.nc\n', 'data/GlobAlbedo.201001.mosaic.5.nc\n', 'data/GlobAlbedo.201009.mosaic.5.nc\n', 'data/GlobAlbedo.201010.mosaic.5.nc\n', 'data/GlobAlbedo.200906.mosaic.5.nc\n', 'data/GlobAlbedo.201002.mosaic.5.nc\n', 'data/GlobAlbedo.200908.mosaic.5.nc\n', 'data/GlobAlbedo.200901.mosaic.5.nc\n', 'data/GlobAlbedo.200910.mosaic.5.nc\n']
or several other ways ...
Putting this together:
import glob
files = [file + '\n' for file in glob.glob('data/*.nc')]
filename = 'data/data.dat'
# open in write mode
fp = open(filename,'w')
fp.writelines(files)
fp.close()
then checking:
cat data/data.dat
data/GlobAlbedo.201008.mosaic.5.nc data/GlobAlbedo.200903.mosaic.5.nc data/GlobAlbedo.200905.mosaic.5.nc data/GlobAlbedo.201007.mosaic.5.nc data/GlobAlbedo.201011.mosaic.5.nc data/GlobAlbedo.200902.mosaic.5.nc data/GlobAlbedo.201006.mosaic.5.nc data/GlobAlbedo.201004.mosaic.5.nc data/GlobAlbedo.200904.mosaic.5.nc data/GlobAlbedo.200911.mosaic.5.nc data/GlobAlbedo.201012.mosaic.5.nc data/GlobAlbedo.201005.mosaic.5.nc data/GlobAlbedo.200912.mosaic.5.nc data/GlobAlbedo.201003.mosaic.5.nc data/GlobAlbedo.200907.mosaic.5.nc data/GlobAlbedo.200909.mosaic.5.nc data/GlobAlbedo.201001.mosaic.5.nc data/GlobAlbedo.201009.mosaic.5.nc data/GlobAlbedo.201010.mosaic.5.nc data/GlobAlbedo.200906.mosaic.5.nc data/GlobAlbedo.201002.mosaic.5.nc data/GlobAlbedo.200908.mosaic.5.nc data/GlobAlbedo.200901.mosaic.5.nc data/GlobAlbedo.200910.mosaic.5.nc
which is what we wanted.
E3.2 Read Image Data into 3D List¶
Write some python code that directly reads the data layers 'DHR_VIS','DHR_NIR','DHR_SW' into a list for a given month and year.
A3.2 Read Image Data into 3D List¶
We start off with the code from the class and attempt to modify this for our purposes:
import gdal
# form a generic string of the form
# NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS
file_template = 'NETCDF:"%s":%s'
# now make a list of the datset names we want
# so we can loop over this
selected_layers = ['DHR_VIS','DHR_NIR','DHR_SW']
# ----------------------------------
# try it out:
root = 'data/'
# example filename : use formatting string:
# %d%02d
year = 2009
month = 1
filename = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
print filename
# set up an empty dictionary to load the data into
data = {}
# use enumerate here to loop over
# the list selected_layers and also have
# access to an index i
for i, layer in enumerate ( selected_layers ):
this_file = file_template % ( filename, layer )
print "Opening Layer %d: %s" % (i, this_file )
g = gdal.Open ( this_file )
# test that the opening worked
# and raise an error otherwise
if g is None:
raise IOError
data[layer] = g.ReadAsArray()
print "\t>>> Read %s!" % layer
# post-hoc load into array called albedo
albedo = [data[f] for f in selected_layers]
data/GlobAlbedo.200901.mosaic.5.nc Opening Layer 0: NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS >>> Read DHR_VIS! Opening Layer 1: NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_NIR >>> Read DHR_NIR! Opening Layer 2: NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_SW >>> Read DHR_SW!
Now, instead of loading the data into a dictionary {}, we simply append onto a list:
import gdal
# NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS
file_template = 'NETCDF:"%s":%s'
selected_layers = ['DHR_VIS','DHR_NIR','DHR_SW']
root = 'data/'
# example filename : use formatting string:
# %d%02d
year = 2009
month = 1
filename = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
# set up an empty list to load the data into
data = []
# loop over
# the list selected_layers
for layer in ( selected_layers ):
this_file = file_template % ( filename, layer )
g = gdal.Open ( this_file )
# test that the opening worked
# and raise an error otherwise
if g is None:
raise IOError
# it is a list, so we will just append the entry here
data.append(g.ReadAsArray())
# check it workwed
print 'previously, we had:',albedo
print '...... we now have:',data
previously, we had: [array([[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[ 0.69980866, 0.69980866, 0.69980866, ..., 0.70370543,
0.70370543, 0.70370543],
[ 0.69980866, 0.69980866, 0.69980866, ..., 0.70370543,
0.70370543, 0.70370543],
[ 0.69980866, 0.69980866, 0.69980866, ..., 0.70370543,
0.70370543, 0.70370543]], dtype=float32), array([[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[ 0.45321581, 0.45321581, 0.45321581, ..., 0.46333486,
0.46333486, 0.46333486],
[ 0.45321581, 0.45321581, 0.45321581, ..., 0.46333486,
0.46333486, 0.46333486],
[ 0.45321581, 0.45321581, 0.45321581, ..., 0.46333486,
0.46333486, 0.46333486]], dtype=float32), array([[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[ 0.57224423, 0.57224423, 0.57224423, ..., 0.57901061,
0.57901061, 0.57901061],
[ 0.57224423, 0.57224423, 0.57224423, ..., 0.57901061,
0.57901061, 0.57901061],
[ 0.57224423, 0.57224423, 0.57224423, ..., 0.57901061,
0.57901061, 0.57901061]], dtype=float32)]
...... we now have: [array([[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[ 0.69980866, 0.69980866, 0.69980866, ..., 0.70370543,
0.70370543, 0.70370543],
[ 0.69980866, 0.69980866, 0.69980866, ..., 0.70370543,
0.70370543, 0.70370543],
[ 0.69980866, 0.69980866, 0.69980866, ..., 0.70370543,
0.70370543, 0.70370543]], dtype=float32), array([[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[ 0.45321581, 0.45321581, 0.45321581, ..., 0.46333486,
0.46333486, 0.46333486],
[ 0.45321581, 0.45321581, 0.45321581, ..., 0.46333486,
0.46333486, 0.46333486],
[ 0.45321581, 0.45321581, 0.45321581, ..., 0.46333486,
0.46333486, 0.46333486]], dtype=float32), array([[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[ 0.57224423, 0.57224423, 0.57224423, ..., 0.57901061,
0.57901061, 0.57901061],
[ 0.57224423, 0.57224423, 0.57224423, ..., 0.57901061,
0.57901061, 0.57901061],
[ 0.57224423, 0.57224423, 0.57224423, ..., 0.57901061,
0.57901061, 0.57901061]], dtype=float32)]
E3.3 Read More Image Data into a 4D List¶
You should now have some code that reads the 3 albedo datasets into a 3D list (3 x 360 x 720) for a given month and year.
Put a loop around this code to make a 4D list dataset (12 x 3 x 360 x 720) for all months in a given year.
A3.3 Read More Image Data into a 4D List¶
Let's start from the code we have above:
import gdal
# NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS
file_template = 'NETCDF:"%s":%s'
selected_layers = ['DHR_VIS','DHR_NIR','DHR_SW']
root = 'data/'
# example filename : use formatting string:
# %d%02d
year = 2009
month = 1
filename = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
# set up an empty list to load the data into
data = []
# loop over
# the list selected_layers
for layer in ( selected_layers ):
this_file = file_template % ( filename, layer )
g = gdal.Open ( this_file )
# test that the opening worked
# and raise an error otherwise
if g is None:
raise IOError
# it is a list, so we will just append the entry here
data.append(g.ReadAsArray())
All we really need to do is to replace where it sets month = 1 by a (for) loop, remembering to indent the code appropriately, and put an 'outer' list (called alldata here) which we put each of the data lists into.
import gdal
# NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS
file_template = 'NETCDF:"%s":%s'
selected_layers = ['DHR_VIS','DHR_NIR','DHR_SW']
root = 'data/'
# example filename : use formatting string:
# %d%02d
year = 2009
# important to put the alldata empty list
# setup outside of the loop, ie before
# we start looping over month
alldata = []
for month in range(1,13):
print "I'm reading month",month,"of year",year
filename = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
# just the data for this month in the list data
data = []
for layer in ( selected_layers ):
this_file = file_template % ( filename, layer )
g = gdal.Open ( this_file )
# test that the opening worked
# and raise an error otherwise
if g is None:
raise IOError
# it is a list, so we will just append the entry here
data.append(g.ReadAsArray())
# now (note indentation!!)
# append to the alldata list
alldata.append(data)
# now check how big ...
print 'dataset dimensions',len(alldata),'x',len(alldata[0]),'x',\
len(alldata[0][0]),'x',len(alldata[0][0][0])
I'm reading month 1 of year 2009 I'm reading month 2 of year 2009 I'm reading month 3 of year 2009 I'm reading month 4 of year 2009 I'm reading month 5 of year 2009 I'm reading month 6 of year 2009 I'm reading month 7 of year 2009 I'm reading month 8 of year 2009 I'm reading month 9 of year 2009 I'm reading month 10 of year 2009 I'm reading month 11 of year 2009 I'm reading month 12 of year 2009 dataset dimensions 12 x 3 x 360 x 720
E3.2 Exercise: Making Movies¶
E3.2.1 Software¶
You can sort of make movies in pylab, but you generally have to make a system call to unix at some point, so it's probably easier to do this all in unix with the utility convert.
At the unix prompt, chack that you have access to convert:
berlin% which convert
/usr/bin/convert
If this doesn't come up with anything useful, there is probably a version in /usr/bin/convert or /usr/local/bin/convert (If you don't have it on your local machine, install ImageMagick which contains the command line tool convert).
To use this, e.g.:
from the unix command line:
berlin% cd ~/Data/geogg122/Chapter3_Scientific_Numerical_Python
berlin% convert data/albedo.jpg files/data/albedo.gif
or from within a notebook:
!convert data/albedo.jpg data/albedo.gif
Or, more practically here, you can run a unix command directly from Python:
import os
cmd = 'convert data/albedo.jpg data/albedo.gif'
os.system(cmd)
0
This will convert the file data/albedo.jpg (in jpeg format) to data/albedo.gif (in gif format).
We can also use convert to make animated gifs, which is one way of making a movie.
E3.2.2 Looping over a set of images¶
You have all of the code you need above to be able to read a GlobAlbedo file for a given month and waveband in Python and save a picture in jpeg format, but to recap for BHR_VIS:
import gdal
import pylab as plt
import os
import calendar
layer = 'BHR_VIS'
# form a generic string of the form
# NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":DHR_VIS
file_template = 'NETCDF:"%s":%s'
root = 'data/'
# example filename : use formatting string:
# %d%02d
year = 2009
# set the month (1-based, i.e. 1 == January)
month = 1
filename = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
g = gdal.Open ( file_template % ( filename, layer ) )
if g is None:
raise IOError
data = g.ReadAsArray()
''' Plot the data and save as picture jpeg format '''
# make a string with the output file name
out_file = root + 'GlobAlbedo.%d%02d.jpg'%(year,month)
# plot
plt.figure(figsize=(10,4))
plt.clf()
# %9s forces the string to be 8 characters long
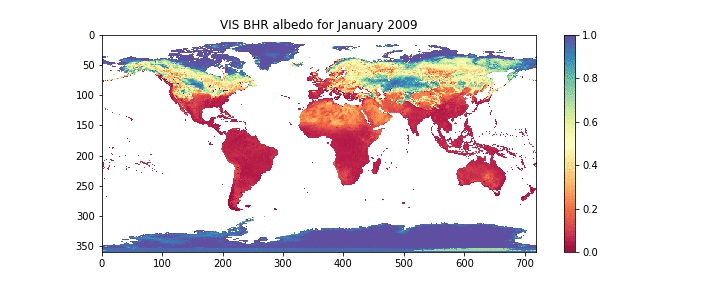
plt.title('VIS BHR albedo for %8s %d'%(calendar.month_name[month],year))
# use nearest neighbour interpolation
# load the array data
plt.imshow(data,interpolation='nearest',cmap=plt.get_cmap('Spectral'),vmin=0.0,vmax=1.0)
# show a colour bar
plt.colorbar()
plt.savefig(out_file)
# convert to gif
# set up the unix command which is of the form
# convert input output
# Here input will be out_file
# and output we can get with out_file.replace('.jpg','.gif')
# i.e. replacing where it says .jpg with .gif
cmd = 'convert %s %s'%(out_file,out_file.replace('.jpg','.gif'))
os.system(cmd)
0
Modify the code above to loop over each month, so that it generates a set of gif format files for the TOTAL SHORTWAVE ALBEDO
You should confirm that these exist, and that the file modification time is when you ran it (not when I generated the files for these notes, which is Oct 10 2013).
ls -l data/GlobAlbedo.??????.gif
-rw-rw-r--. 1 plewis plewis 50845 Oct 3 12:13 data/GlobAlbedo.200901.gif -rw-rw-r--. 1 plewis plewis 28139 Sep 30 11:56 data/GlobAlbedo.200902.gif -rw-rw-r--. 1 plewis plewis 28259 Sep 30 11:56 data/GlobAlbedo.200903.gif -rw-rw-r--. 1 plewis plewis 28249 Sep 30 11:56 data/GlobAlbedo.200904.gif -rw-rw-r--. 1 plewis plewis 28468 Sep 30 11:56 data/GlobAlbedo.200905.gif -rw-rw-r--. 1 plewis plewis 28672 Sep 30 11:56 data/GlobAlbedo.200906.gif -rw-rw-r--. 1 plewis plewis 28656 Sep 30 11:56 data/GlobAlbedo.200907.gif -rw-rw-r--. 1 plewis plewis 28275 Sep 30 11:56 data/GlobAlbedo.200908.gif -rw-rw-r--. 1 plewis plewis 28952 Sep 30 11:56 data/GlobAlbedo.200909.gif -rw-rw-r--. 1 plewis plewis 28450 Sep 30 11:56 data/GlobAlbedo.200910.gif -rw-rw-r--. 1 plewis plewis 28570 Sep 30 11:56 data/GlobAlbedo.200911.gif -rw-rw-r--. 1 plewis plewis 28438 Sep 30 11:56 data/GlobAlbedo.200912.gif
A3.2.2 Answer: Looping over a set of images¶
Really all you need to do here is to make month appear in a loop, e.g. using:
for month in range(1,13):
and then make sure that all of the code below is in that loop (i.e. indented) as below.
and finally, make sure you change the title
You should also however, go through the code above line by line, making sure you appreciate what is going on at each stage and why we have done these things (in this order).
# lets try something out to see how we can loop easily in this case
# we know that the mionth names are contained in calendar.month_name
# so we might try to just loop over that
import calendar
for month_name in calendar.month_name:
print month_name
January February March April May June July August September October November December
# but we might also want access to the month index, so we might use
# enumerate
import calendar
for month,month_name in enumerate(calendar.month_name):
print month,month_name
0 1 January 2 February 3 March 4 April 5 May 6 June 7 July 8 August 9 September 10 October 11 November 12 December
# but thats not quite right ... we don't want
# the blank zero entry so we only
# loop over the slice [1:] in month_name
import calendar
for month,month_name in enumerate(calendar.month_name[1:]):
print month,month_name
0 January 1 February 2 March 3 April 4 May 5 June 6 July 7 August 8 September 9 October 10 November 11 December
# but now we have a zero index for January
# which isnt what we want
# we could just add one to this when we use it
# but that is a bit ugly ...
# so instead, use a feature of enumerate()
import calendar
for month,month_name in enumerate(calendar.month_name[1:],start=1):
print month,month_name
1 January 2 February 3 March 4 April 5 May 6 June 7 July 8 August 9 September 10 October 11 November 12 December
So, lets put that together with the other code:
import gdal
import pylab as plt
import os
import calendar
# set the layer we want
layer = 'BHR_SW'
# form a generic string of the form
# NETCDF:"data/GlobAlbedo.200901.mosaic.5.nc":BHR_SW
file_template = 'NETCDF:"%s":%s'
root = 'data/'
# example filename : use formatting string:
# %d%02d
year = 2009
# set the month (1-based, i.e. 1 == January)
# its a good idea to use enumerate here
# but we want the numbers to start at 1
# so we set start=1 in the enumerate call
for month,month_name in enumerate(calendar.month_name[1:],start=1):
'''
Loop over each month
setting month = 1,2,..12
month_name= 'January', ... 'December'
(unless we change the locale!)
'''
filename = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
g = gdal.Open ( file_template % ( filename, layer ) )
if g is None:
raise IOError
# note that we are re-using the varioable data
# each time in the loop as we haver no need to store it as yet
data = g.ReadAsArray()
''' Plot the data and save as picture jpeg format '''
# make a string with the output file name
out_file = root + 'GlobAlbedo.%d%02d.jpg'%(year,month)
# plot
plt.figure(figsize=(10,4))
plt.clf()
# %9s forces the string to be 8 characters long
plt.title('SW BHR albedo for %8s %d'%(month_name,year))
# use nearest neighbour interpolation
# load the array data
plt.imshow(data,interpolation='nearest',cmap=plt.get_cmap('Spectral'),vmin=0.0,vmax=1.0)
# show a colour bar
plt.colorbar()
plt.savefig(out_file)
# convert to gif
# set up the unix command which is of the form
# convert input output
# Here input will be out_file
# and output we can get with out_file.replace('.jpg','.gif')
# i.e. replacing where it says .jpg with .gif
cmd = 'convert %s %s'%(out_file,out_file.replace('.jpg','.gif'))
os.system(cmd)
E3.2.3 Make the movie¶
The unix command to convert these files to an animated gif is:
convert -delay 100 -loop 0 data/GlobAlbedo.2009??.gif data/GlobAlbedo.2009.SW.gif
Run this (ideally, from within Python) to create the animated gif GlobAlbedo.2009.SW.gif
Again, confirm that you created this file (and it is not just a version you downloaded):
ls -l data/GlobAlbedo.2009.SW.gif
-rw-rw-r--. 1 plewis plewis 340658 Sep 30 11:56 data/GlobAlbedo.2009.SW.gif
A3.2.3 Answer: Make the movie¶
You could just type the command at the unix prompt ... but to do it using a system call from within Python, e.g.:
import os
# this is quite a neat way of generating the string for the input files
out_file = 'data/GlobAlbedo.%d.SW.gif'%year
in_files = out_file.replace('.SW.gif','??.gif')
cmd = 'convert -delay 100 -loop 0 %s %s'%(in_files,out_file)
# check the cmd is ok
print cmd
convert -delay 100 -loop 0 data/GlobAlbedo.2009??.gif data/GlobAlbedo.2009.SW.gif
# good ... so now run it
os.system(cmd)
0

To view the animated gif you have generated, open it in a browser (or view in the notebook):
E3.3 Exercise: 3D Masked Array¶
from netCDF4 import Dataset
import numpy as np
root = 'files/data/'
year = 2009
# which months?
months = xrange(1,13)
# empty list
data = []
# loop over month
# use enumerate so we have an index counter
for i,month in enumerate(months):
# this then is the file we want
local_file = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
# load the netCDF data from the file local_file
nc = Dataset(local_file,'r')
# append what we read to the list called data
data.append(np.array(nc.variables['DHR_SW']))
# convert data to a numpy array (its a list of arrays at the moment)
data = np.array(data)
N.B. Do this exercise before proceeding to the next section.
Taking the code above as a starting point, generate a masked array of the GlobAlbedo dataset for the year 2009.
A3.3 Answer: 3D Masked Array¶
To recap, to make a masked array, we use some code such as:
import numpy.ma as ma
band = np.array(nc.variables['DHR_SW'])
masked_band = ma.array(band,mask=np.isnan(band))
print masked_band.mask
[[ True True True ..., True True True] [ True True True ..., True True True] [ True True True ..., True True True] ..., [False False False ..., False False False] [False False False ..., False False False] [False False False ..., False False False]]
So we can do this for each band as we loop over each month:
from netCDF4 import Dataset
import numpy as np
import numpy.ma as ma
root = 'files/data/'
year = 2009
# which months?
months = xrange(1,13)
# empty list
data = []
# loop over month
# use enumerate so we have an index counter
for i,month in enumerate(months):
# this then is the file we want
local_file = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
# load the netCDF data from the file local_file
nc = Dataset(local_file,'r')
# load into the variable 'band'
band = np.array(nc.variables['DHR_SW'])
# convert to a masked array
masked_band = ma.array(band,mask=np.isnan(band))
# append what we read to the list called data
data.append(masked_band)
# convert data to a numpy array (its a list of arrays at the moment)
data = np.array(data)
That's a good start, but we used:
data = np.array(data)
at the end, which means that we have a 3D numpy array ... rather than a masked array:
print type(data)
print data.shape
print data.ndim
<type 'numpy.ndarray'> (12, 360, 720) 3
so we need to replace this with a function to make it into a masked array:
from netCDF4 import Dataset
import numpy as np
import numpy.ma as ma
root = 'files/data/'
year = 2009
# which months?
months = xrange(1,13)
# empty list
data = []
# loop over month
# use enumerate so we have an index counter
for i,month in enumerate(months):
# this then is the file we want
local_file = root + 'GlobAlbedo.%d%02d.mosaic.5.nc'%(year,month)
# load the netCDF data from the file local_file
nc = Dataset(local_file,'r')
# load into the variable 'band'
band = np.array(nc.variables['DHR_SW'])
# convert to a masked array
masked_band = ma.array(band,mask=np.isnan(band))
# append what we read to the list called data
data.append(masked_band)
# convert data to a numpy array (its a list of arrays at the moment)
data = ma.array(data)
print type(data)
print data.shape
print data.ndim
<class 'numpy.ma.core.MaskedArray'> (12, 360, 720) 3
we might notice that now, the data mask (data.mask) is a 3D mask:
print data.mask.shape
print data.mask.ndim
(12, 360, 720) 3
and we might just check that the mask is the same for all months:
First, lets sum the masks over axis 0 (i.e. over all months):
sum = (data.mask).sum(axis=0)
print sum.shape
(360, 720)
print sum
[[12 12 12 ..., 12 12 12] [12 12 12 ..., 12 12 12] [12 12 12 ..., 12 12 12] ..., [ 0 0 0 ..., 0 0 0] [ 0 0 0 ..., 0 0 0] [ 0 0 0 ..., 0 0 0]]
This should be 12 (where mask is True) or 0 (where mask is False). How can we chack that quickly?
Fortunately, there is a convenient numpy function np.unique that will give you the unique values in an array:
np.unique(sum)
array([ 0, 12])
The only values in here are 12 and 0, so the masks must be consistent!