Pyura (Piura) chilensis Transcriptome¶
Objectives¶
- Gain Experience using IPython Notebook (or alternative) to document research so that it is useful to future you.
- Use iPlant Collaborative Discovery Environment
- Install and software locally (BLAST)
- Explore structured data tables with commandline
- Use SQLShare to join and query table.
What is Piura?¶
In [1]:
from IPython.display import HTML
HTML('<iframe src=http://en.m.wikipedia.org/wiki/Pyura_chilensis width=80% height=350></iframe>')
Out[1]:
Map¶
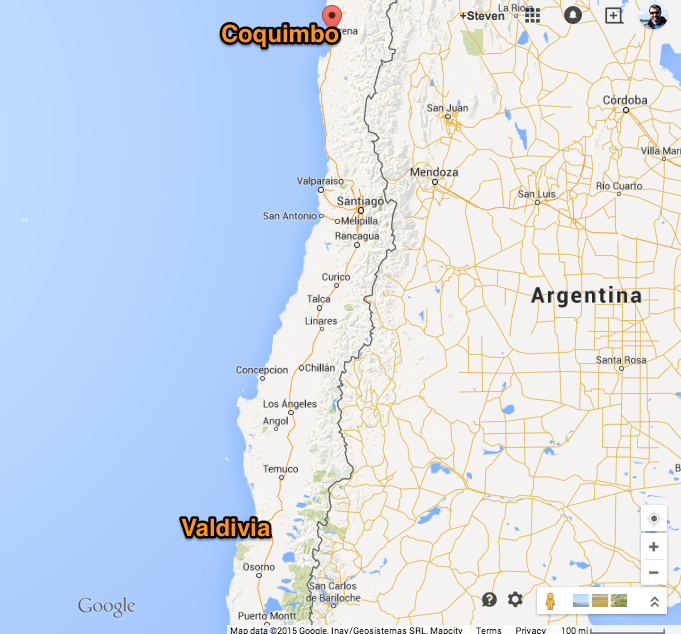
Scenario¶
Sequencing data from 454 Platform was just sequenced by a core facility. You need to take it, make sure it is of decent quality, your data, and determine the functional category of genes expressed.
Location of Raw Data¶
Step 1 Upload to iPlant Discovery Environment¶
Step 2 Convert SFF to Fastq¶
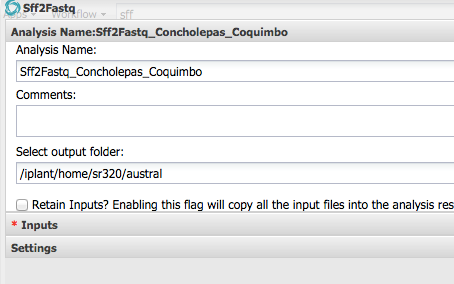
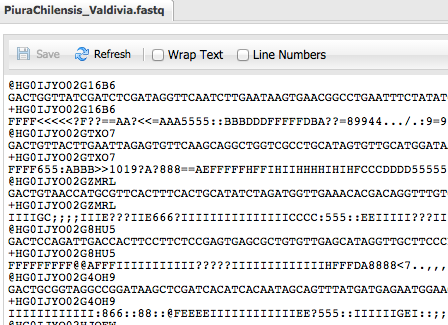
Step 3 Check Sequence Quality - FASTQC¶
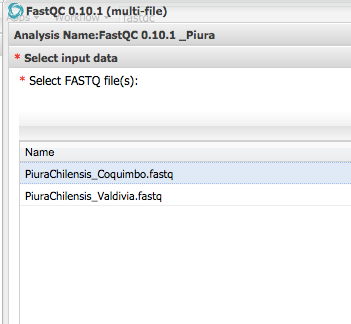
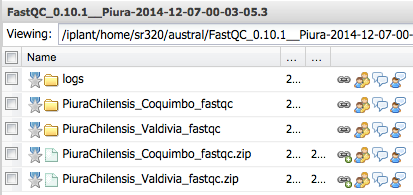
Step 3b Download zip file and view graphs¶
In [2]:
from IPython.display import HTML
HTML('<iframe src=../fastqc/PiuraChilensis_Coquimbo_fastqc/fastqc_report.html width=100% height=650></iframe>')
Out[2]:
In [3]:
from IPython.display import HTML
HTML('<iframe src=../fastqc/PiuraChilensis_Valdivia_fastqc/fastqc_report.html width=100% height=650></iframe>')
Out[3]:
Step 4 Trim Sequences¶
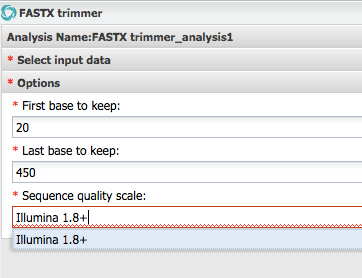
Step 5 Recheck Sequence Quality¶
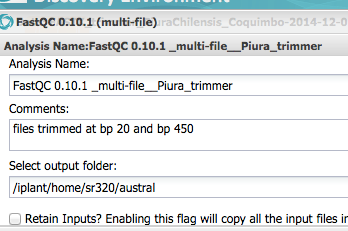
In [12]:
from IPython.display import HTML
HTML('<iframe src=../fastqc/trimmer/PiuraChilensis_Coquimbo_trimmer_out_fastqc/fastqc_report.html width=100% height=650></iframe>')
Out[12]:
In [13]:
from IPython.display import HTML
HTML('<iframe src=../fastqc/trimmer/PiuraChilensis_Valdivia_trimmer_out_fastqc/fastqc_report.html width=100% height=650></iframe>')
Out[13]:
Step 6 Assemble Reads into Transcriptome (Trinity)¶
Lets do each sample separately
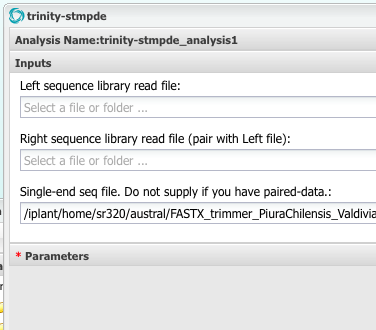
Step 6b Assemble Reads into Transcriptome (CLC)¶
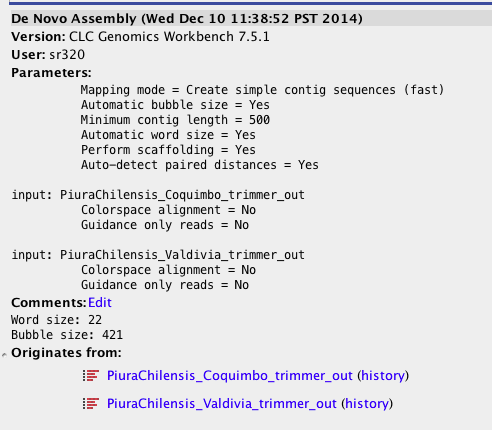
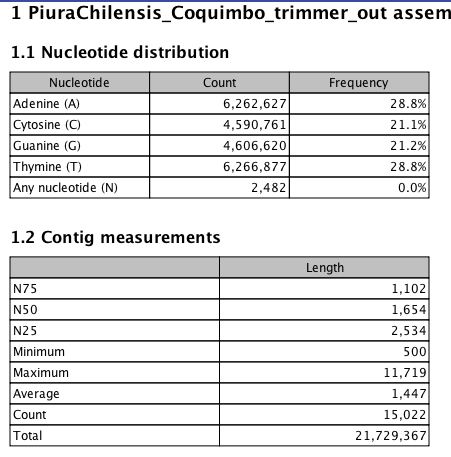
In [5]:
#in repo
!head ../data/Piura_v1_contigs.fa
>PiuraChilensis_v1_contig_1 ATTTACAATACGAAGTAAAATAGATAACGTGAAAATAATCTTGGTGCTGGATGATCGATC AAGTTCACCAATATTTTATTGTAAAAAATCATTCTAAACAGCATGAAATCGTGTACAATG TATAAACAAGCAAATATATAACACTAAAGCAAGAGGGCGTAAGTGGGGGGGTGGGTGAGA GTAAAAAATTCAAACATGTCAAATACCCCGGCGTTAGCCTTAAAAGCACCATGGACTTCT GCCTTCAATAAGCATAAAATTAAAACACCTAATACACAATGAATATACAGATAAAACAGA TTTATGAATAGTTGGTGTTACATCTTTTACAGCCATAAGCCTTCATTTTGCTTCCAAACG TATAAAATCTGACTTGGAACAATATACAGCCATGAGATATGACACAGCGAGCACTACAAT ATATATTTATCTTGTACTATACAGCCTGTACAAGAAAATTCTGGAATTGTCTTCACAAGA GACAGAAAAATAGTTGCAATGTGAATGCTAGTCTACTATTTGATCACAATTGGATAGAAA
Step 7 Annotate Contigs (Blast)¶
In [ ]:
!blastx \
-query ../data/Piura_v1_contigs.fa \
-db /Volumes/Bay3/Software/ncbi-blast-2.2.29\+/db/uniprot_sprot_r2013_12 \
-out ../data/Piura_v1_uniprot_sprot.tab \
-evalue 1E-05 \
-max_target_seqs 1 \
-max_hsps 1 \
-outfmt 6 \
-num_threads 2
In [1]:
#in repo
!tail -2 ../data/Piura_v1_uniprot_sprot.tab
PiuraChilensis_v1_contig_15018 sp|O18973|RABX5_BOVIN 79.07 43 9 0 888 1016 321 363 3e-15 80.1 PiuraChilensis_v1_contig_15021 sp|Q9Z1Z1|E2AK3_RAT 51.61 93 45 0 100 378 971 1063 8e-22 97.8
In [2]:
#still needs to be run
!/Applications/bioinfo/ncbi-blast-2.2.30/bin/blastx \
-query ../data/PiuraC_Val_Trinity_2ndhalf.fasta \
-db /Users/sr320/data-genomic/blast/db/uniprot_sprot_r2015_01 \
-out ../data/PiuraC_Val_Trinity_uniprot_sprot_2ndhalf.tab \
-evalue 1E-05 \
-max_target_seqs 1 \
-max_hsps 1 \
-outfmt 6 \
-num_threads 6
In [4]:
!tail ../data/PiuraC_Val_Trinity_uniprot_sprot_2ndhalf.tab
comp31014_c0_seq1 sp|Q63159|COQ3_RAT 50.93 108 50 1 3 317 155 262 5e-29 111 comp31018_c0_seq1 sp|P43695|GAT5A_XENLA 68.97 58 16 1 409 242 226 283 2e-18 84.7 comp31019_c0_seq1 sp|P90747|YE56_CAEEL 48.53 68 33 1 54 251 567 634 5e-14 71.2 comp31028_c0_seq1 sp|Q5RAG7|XCT_PONAB 44.00 100 56 0 302 3 298 397 4e-13 69.7 comp31030_c0_seq1 sp|Q9CQJ2|PIHD1_MOUSE 50.57 87 38 2 403 143 57 138 3e-23 96.7 comp31033_c0_seq1 sp|Q6BEA2|PRS27_RAT 71.43 28 8 0 132 215 53 80 6e-06 46.6 comp31037_c0_seq1 sp|O95425|SVIL_HUMAN 31.94 72 48 1 10 225 1793 1863 1e-06 49.3 comp31054_c0_seq1 sp|A2AGA4|RHBL2_MOUSE 43.93 107 57 2 10 324 195 300 5e-16 76.6 comp31056_c0_seq1 sp|Q9NVH0|EXD2_HUMAN 40.30 134 66 3 6 389 170 295 1e-20 91.3 comp31058_c0_seq1 sp|Q9WUA2|SYFB_MOUSE 69.70 66 20 0 53 250 353 418 3e-27 108
Step 8 Explore files in commandline¶
In [2]:
!wc -l ../data/Piura_v1_uniprot_sprot.tab
9498 ../data/Piura_v1_uniprot_sprot.tab
In [3]:
!head -2 ../data/Piura_v1_uniprot_sprot.tab
PiuraChilensis_v1_contig_3 sp|Q6P9A1|ZN530_HUMAN 33.33 105 61 3 825 1118 414 516 1e-07 57.4 PiuraChilensis_v1_contig_4 sp|Q8TGM6|TAR1_YEAST 70.91 55 16 0 3829 3665 22 76 3e-15 80.1
In [4]:
!tr '|' "\t" <../data/Piura_v1_uniprot_sprot.tab> ../data/Piura_v1_uniprot_sprot_sql.tab
!echo SQLShare ready version has Pipes converted to Tabs ....
!head -1 ../data/Piura_v1_uniprot_sprot_sql.tab
SQLShare ready version has Pipes converted to Tabs .... PiuraChilensis_v1_contig_3 sp Q6P9A1 ZN530_HUMAN 33.33 105 61 3 825 1118 414 516 1e-07 57.4
Step 9 Joining Blast results with other information (SQLShare)¶
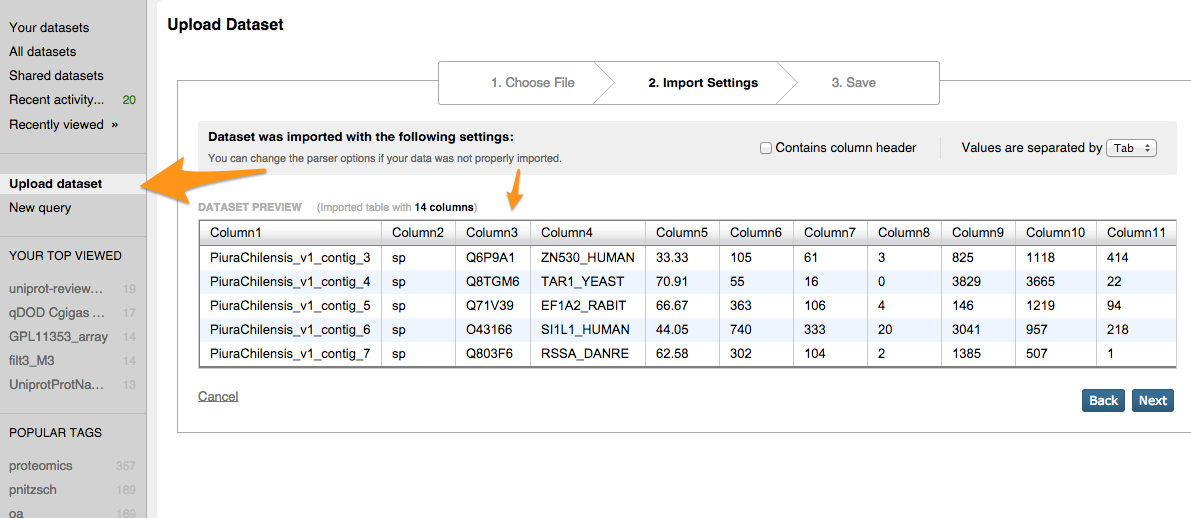
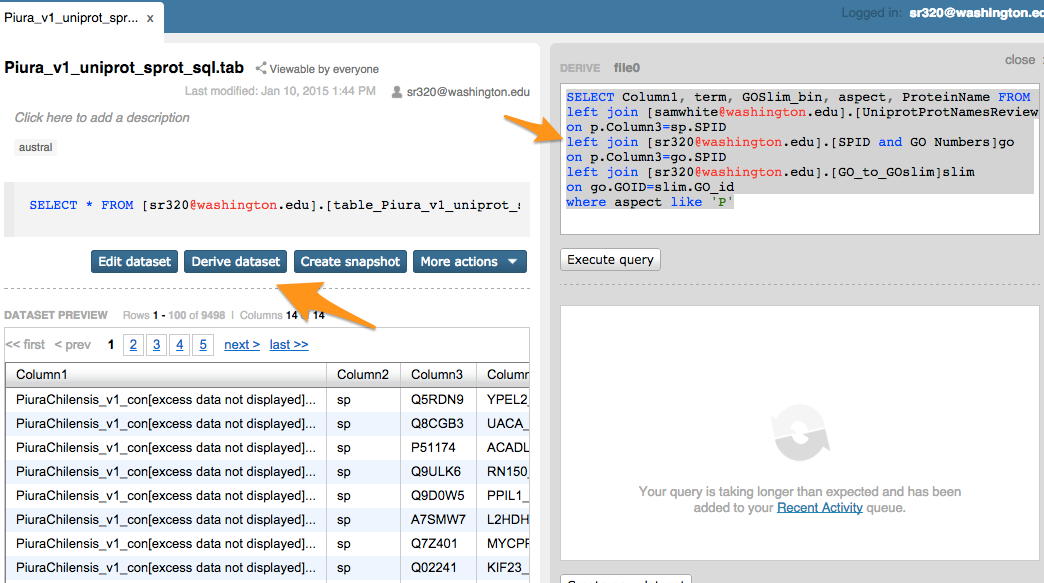
SELECT Column1, term, GOSlim_bin, aspect, ProteinName FROM [sr320@washington.edu].[Piura_v1_uniprot_sprot_sql.tab]p
left join [samwhite@washington.edu].[UniprotProtNamesReviewed_yes20130610]sp
on p.Column3=sp.SPID
left join [sr320@washington.edu].[SPID and GO Numbers]go
on p.Column3=go.SPID
left join [sr320@washington.edu].[GO_to_GOslim]slim
on go.GOID=slim.GO_id
where aspect like 'P'
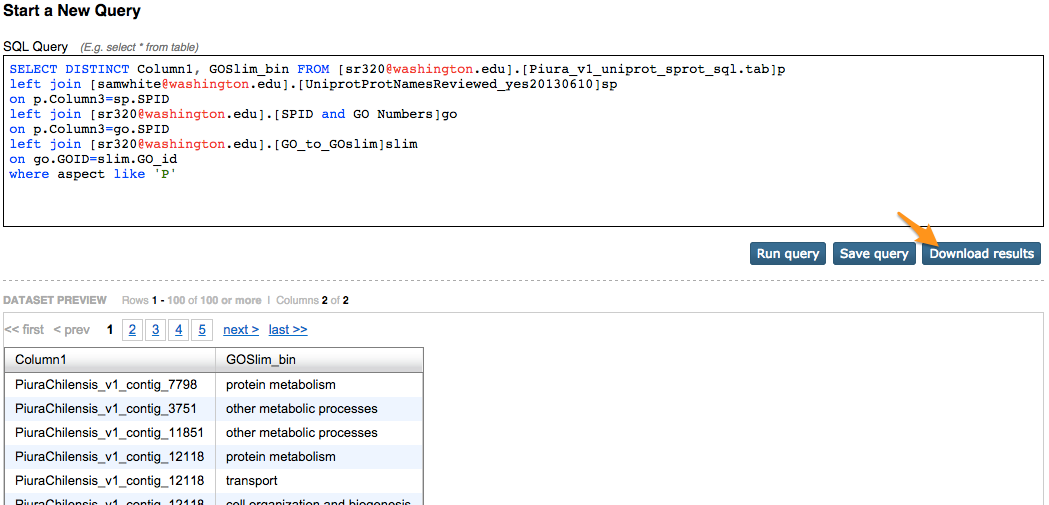
SELECT DISTINCT Column1, GOSlim_bin FROM [sr320@washington.edu].[Piura_v1_uniprot_sprot_sql.tab]p
left join [sr320@washington.edu].[SPID and GO Numbers]go
on p.Column3=go.SPID
left join [sr320@washington.edu].[GO_to_GOslim]slim
on go.GOID=slim.GO_id
where aspect like 'P'
Bonus Step! Automate - fasta2slim¶
In [ ]: