BLAST¶
Screenshot of Blast page at NCBI.
Download Stand-alone BLAST¶
ftp://ftp.ncbi.nlm.nih.gov/blast/executables/blast+/LATEST/
In [28]:
#unzipping [-]x --extract --get; -v, --verbose; -z, --gzip; -f, --file F
!tar -xzvf ncbi-blast-2.2.29+-universal-macosx.tar.gz
x ncbi-blast-2.2.29+/ x ncbi-blast-2.2.29+/bin/ x ncbi-blast-2.2.29+/bin/makembindex x ncbi-blast-2.2.29+/bin/tblastn x ncbi-blast-2.2.29+/bin/psiblast x ncbi-blast-2.2.29+/bin/rpsblast x ncbi-blast-2.2.29+/bin/legacy_blast.pl x ncbi-blast-2.2.29+/bin/blastdbcmd x ncbi-blast-2.2.29+/bin/makeblastdb x ncbi-blast-2.2.29+/bin/tblastx x ncbi-blast-2.2.29+/bin/blastn x ncbi-blast-2.2.29+/bin/blastp x ncbi-blast-2.2.29+/bin/segmasker x ncbi-blast-2.2.29+/bin/dustmasker x ncbi-blast-2.2.29+/bin/blastx x ncbi-blast-2.2.29+/bin/blast_formatter x ncbi-blast-2.2.29+/bin/windowmasker x ncbi-blast-2.2.29+/bin/blastdb_aliastool x ncbi-blast-2.2.29+/bin/convert2blastmask x ncbi-blast-2.2.29+/bin/update_blastdb.pl x ncbi-blast-2.2.29+/bin/deltablast x ncbi-blast-2.2.29+/bin/blastdbcheck x ncbi-blast-2.2.29+/bin/rpstblastn x ncbi-blast-2.2.29+/bin/makeprofiledb x ncbi-blast-2.2.29+/doc/ x ncbi-blast-2.2.29+/doc/README.txt x ncbi-blast-2.2.29+/README x ncbi-blast-2.2.29+/ncbi_package_info x ncbi-blast-2.2.29+/LICENSE x ncbi-blast-2.2.29+/ChangeLog
In [31]:
cd ncbi-blast-2.2.29+/bin
/Volumes/Bay3/Software/ncbi-blast-2.2.29+/bin
In [ ]:
In [33]:
ls -1
blast_formatter* blastdb_aliastool* blastdbcheck* blastdbcmd* blastn* blastp* blastx* convert2blastmask* deltablast* dustmasker* legacy_blast.pl* makeblastdb* makembindex* makeprofiledb* psiblast* rpsblast* rpstblastn* segmasker* tblastn* tblastx* update_blastdb.pl* windowmasker*
In [35]:
#check to see if "works"
!blastx -h
USAGE
blastx [-h] [-help] [-import_search_strategy filename]
[-export_search_strategy filename] [-db database_name]
[-dbsize num_letters] [-gilist filename] [-seqidlist filename]
[-negative_gilist filename] [-entrez_query entrez_query]
[-db_soft_mask filtering_algorithm] [-db_hard_mask filtering_algorithm]
[-subject subject_input_file] [-subject_loc range] [-query input_file]
[-out output_file] [-evalue evalue] [-word_size int_value]
[-gapopen open_penalty] [-gapextend extend_penalty]
[-xdrop_ungap float_value] [-xdrop_gap float_value]
[-xdrop_gap_final float_value] [-searchsp int_value]
[-max_hsps_per_subject int_value] [-max_intron_length length]
[-seg SEG_options] [-soft_masking soft_masking] [-matrix matrix_name]
[-threshold float_value] [-culling_limit int_value]
[-best_hit_overhang float_value] [-best_hit_score_edge float_value]
[-window_size int_value] [-ungapped] [-lcase_masking] [-query_loc range]
[-strand strand] [-parse_deflines] [-query_gencode int_value]
[-outfmt format] [-show_gis] [-num_descriptions int_value]
[-num_alignments int_value] [-html] [-max_target_seqs num_sequences]
[-num_threads int_value] [-remote] [-comp_based_stats compo]
[-use_sw_tback] [-version]
DESCRIPTION
Translated Query-Protein Subject BLAST 2.2.28+
Use '-help' to print detailed descriptions of command line arguments
Create a Blast Database¶
I would like to make a database of UniProt/Swiss-prot.
Screenshot:
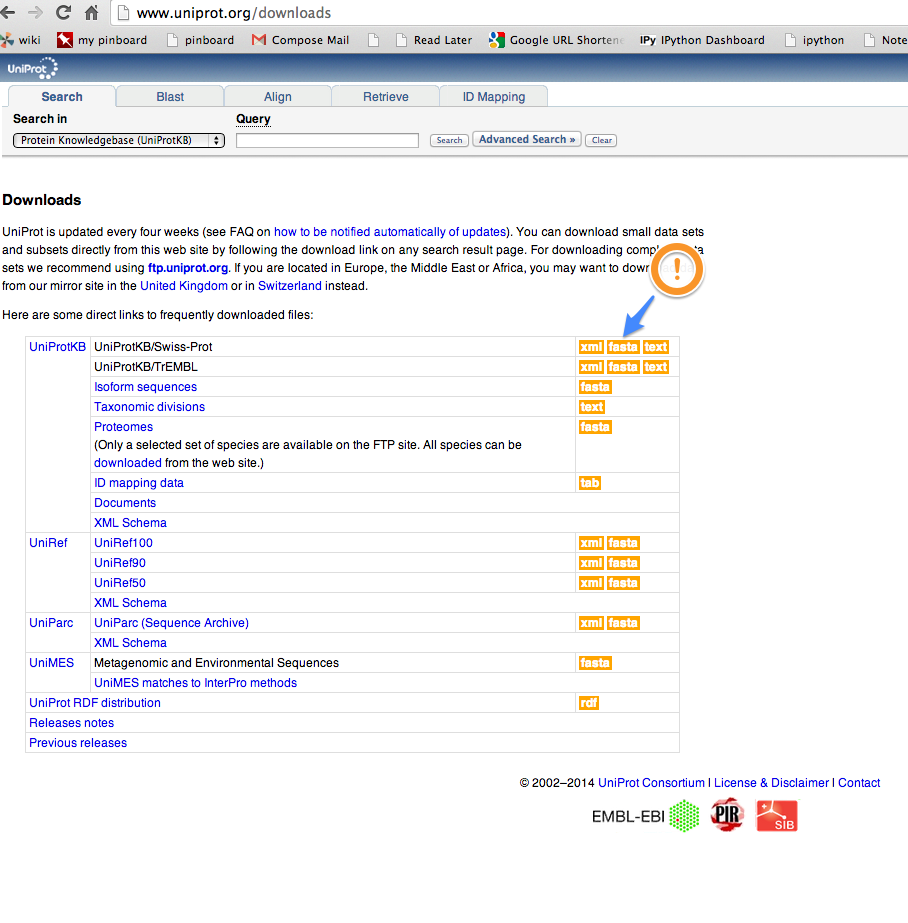
In [53]:
cd ncbi-blast-2.2.29+/
[Errno 2] No such file or directory: 'ncbi-blast-2.2.29+/' /Volumes/Bay3/Software/ncbi-blast-2.2.29+/db
In [54]:
cd db
[Errno 2] No such file or directory: 'db' /Volumes/Bay3/Software/ncbi-blast-2.2.29+/db
In [56]:
ls
uniprot_sprot.fasta.gz
In [62]:
!gzip -d uniprot_sprot.fasta.gz
In [63]:
ls
uniprot_sprot.fasta
In [65]:
pwd
Out[65]:
u'/Volumes/Bay3/Software/ncbi-blast-2.2.29+/db'
In [66]:
#note I am working in dir db, thus can just use file names. Most times you might use the complete path.
!makeblastdb -in uniprot_sprot.fasta -dbtype prot -out uniprot_sprot_r2013_12
Building a new DB, current time: 01/08/2014 11:34:36 New DB name: uniprot_sprot_r2013_12 New DB title: uniprot_sprot.fasta Sequence type: Protein Keep Linkouts: T Keep MBits: T Maximum file size: 1000000000B Adding sequences from FASTA; added 541954 sequences in 53.9535 seconds.
Get a Query Sequence¶
In [79]:
#creating new directory;
!pwd
/Volumes/Bay3/Software/ncbi-blast-2.2.29+/db
In [80]:
cd ..
/Volumes/Bay3/Software/ncbi-blast-2.2.29+
In [83]:
!mkdir query
In [85]:
cd query/
/Volumes/Bay3/Software/ncbi-blast-2.2.29+/query
In [86]:
#getting file from url to local location
#also curl -O works
!wget http://eagle.fish.washington.edu/cnidarian/Ab_4denovo_CLC6_a.fa
--2014-01-08 11:40:14-- http://eagle.fish.washington.edu/cnidarian/Ab_4denovo_CLC6_a.fa Resolving eagle.fish.washington.edu... 128.95.149.81 Connecting to eagle.fish.washington.edu|128.95.149.81|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 2030182 (1.9M) [text/plain] Saving to: `Ab_4denovo_CLC6_a.fa' 100%[======================================>] 2,030,182 --.-K/s in 0.03s 2014-01-08 11:40:14 (68.2 MB/s) - `Ab_4denovo_CLC6_a.fa' saved [2030182/2030182]
In [87]:
#lets get a preview
!head Ab_4denovo_CLC6_a.fa
>solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_1 ACACCCCACCCCAACGCACCCTCACCCCCACCCCAACAATCCATGATTGAATACTTCATC TATCCAAGACAAACTCCTCCTACAATCCATGATAGAATTCCTCCAAAAATAATTTCACAC TGAAACTCCGGTATCCGAGTTATTTTGTTCCCAGTAAAATGGCATCAACAAAAGTAGGTC TGGATTAACGAACCAATGTTGCTGCGTAATATCCCATTGACATATCTTGTCGATTCCTAC CAGGATCCGGACTGACGAGATTTCACTGTACGTTTATGCAAGTCATTTCCATATATAAAA TTGGATCTTATTTGCACAGTTAAATGTCTCTATGCTTATTTATAAATCAATGCCCGTAAG CTCCTAATATTTCTCTTTTCGTCCGACGAGCAAACAGTGAGTTTACTGTGGCCTTCAGCA AAAGTATTGATGTTGTAAATCTCAGTTGTGATTGAACAATTTGCCTCACTAGAAGTAGCC TTC
In [88]:
#word count
!wc Ab_4denovo_CLC6_a.fa
35092 35092 2030182 Ab_4denovo_CLC6_a.fa
In [90]:
#how many sequences? lets count ">" as we know each contig has 1
!grep -c ">" Ab_4denovo_CLC6_a.fa
5490
Run Blast¶
In [ ]:
#will use full paths..
!blastx \
-query /Volumes/Bay3/Software/ncbi-blast-2.2.29\+/query/Ab_4denovo_CLC6_a.fa \
-db /Volumes/Bay3/Software/ncbi-blast-2.2.29\+/db/uniprot_sprot_r2013_12 \
-out /Volumes/Bay3/Software/ncbi-blast-2.2.29\+/out/Ab_4denovo_CLC6_a_uniprot_blastx.tab \
-evalue 1E-20 \
-max_target_seqs 1 \
-outfmt 6
In [94]:
!head /Volumes/Bay3/Software/ncbi-blast-2.2.29\+/out/Ab_4denovo_CLC6_a_uniprot_blastx.tab
solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_3 sp|O42248|GBLP_DANRE 82.46 171 30 0 1 513 35 205 1e-101 301 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_5 sp|Q08013|SSRG_RAT 75.38 65 16 0 3 197 121 185 1e-27 104 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_6 sp|P12234|MPCP_BOVIN 76.62 77 18 0 2 232 286 362 2e-23 98.6 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_9 sp|Q41629|ADT1_WHEAT 82.26 62 11 0 3 188 170 231 3e-27 104 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_13 sp|Q32NG4|PDDC1_XENLA 54.44 90 40 1 1 270 140 228 1e-27 106 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_23 sp|Q9GNE2|RL23_AEDAE 97.22 72 2 0 67 282 14 85 1e-42 142 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_31 sp|Q3V1H3|HPHL1_MOUSE 53.38 133 59 1 2 391 23 155 5e-42 153 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_32 sp|Q641Y2|NDUS2_RAT 88.03 117 14 0 2 352 334 450 1e-70 224 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_37 sp|Q9D3D9|ATPD_MOUSE 56.10 123 54 0 2 370 46 168 7e-42 144 solid0078_20110412_FRAG_BC_WHITE_WHITE_F3_QV_SE_trimmed_contig_39 sp|Q39613|CYPH_CATRO 75.00 120 23 1 55 393 1 120 7e-49 160
In [95]:
!wc /Volumes/Bay3/Software/ncbi-blast-2.2.29\+/out/Ab_4denovo_CLC6_a_uniprot_blastx.tab
664 7968 84910 /Volumes/Bay3/Software/ncbi-blast-2.2.29+/out/Ab_4denovo_CLC6_a_uniprot_blastx.tab
In [ ]: