7.0 Fire/ENSO teleconnections¶
7.1 Introduction¶
There is much public and scientific interest in monitoring and predicting the activity of wildfires and such topics are often in the media.
Part of this interest stems from the role fire plays in issues such as land cover change, deforestation and forest degradation and Carbon emissions from the land surface to the atmosphere, but also of concern are human health impacts. The impacts of fire should not however be considered as wholy negative, as it plays a significant role in natural ecosystem processes.
For many regions of the Earth, there are large inter-annual variations in the timing, frequency and severity of wildfires. Whilst anthropogenic activity accounts for a large and probably increasing proportion of fires started, this is not in itself a new phenomenon.
Fires spread where: (i) there is an ignition source (lightning or man, mostly); (ii) sufficient combustible fuel to maintain the fire. The latter is strongly dependent on fuel loads and mositure content, as well as meteorological conditions. Generally then, when conditions are drier (and there is sufficient fuel and appropriate weather conditions), we would expect fire spread to increase. If the number of ignitions remained approximately constant, this would mean more fires. Many models of fire activity predict increases in fire frequency in the coming decades, although there may well be different behaviours in different parts of the world.
Satellite data has been able to provide us with increasingly useful tools for monitoring wildfire activity, particularly since 2000 with the MODIS instruments on the NASA Terra and Aqua (2002) satellites. A suite of ‘fire’ products have been generated from these data that have been used in a large number of publications and practical/management projects.
There is growing evidence of ‘teleconnection’ links between fire occurence and large scale climate patterns, such as ENSO.
The proposed mechanisms are essentially that such climatic patterns are linked to local water status and temperature and thus affect the ability of fires to spread. For some regions of the Earth, empirical models built from such considerations have quite reasonable predictive skill, meaning that fire season severity might be predicted some months ahead of time.
7.2 A Practical Exercise¶
7.2.1 In This Session¶
In this session, you will be working in groups (of 3 or 4) to build a computer code in python to explore links between fire activity and Sea Surface Temperature anomalies.
This is a team exercise, but does not form part of your formal assessment for this course. You should be able to complete the exercise in the 3 hour session, if you work effectively as a team. Staff will be on hand to provide pointers.
You should be able to complete the exercise using coding skills and python modules that you have previously experience of, though we will also provide some pointers to get you started.
7.2.2 Statement of the problem¶
Using monthly fire count data from MODIS Terra, develop and test a predictive model for the number of fires per unit area per year driven by Sea Surface Temperature anomaly data.
The datasets should be created at 0.5 degree resolution on a latitude/longitude grid.
You should concentrate on building the model for the peak fire count in a particular year at a particular location, i.e. derive your model for annual peak fire count.
Reading datasets¶
You should use the datasets described below.
If you follow the naming conventions below, you should have:
fdata : shape e.g. (186, 36, 72) i.e. (nmonths,nlat,nlon) of fire count data
as a masked array.
fyear : shape e.g. (186,) for the years for which the Sea Surface Temperature anomaly are available.
fmonth: shape e.g. (186,) the associated month
cyears : shape e.g. (69,) for the years for which the Sea Surface Temperature anomaly are available.
cdata : shape e.g. (12,69) for each month (the 12) and each year for which the Sea Surface Temperature anomaly are available.
Derived Information¶
To do the modelling for this exercise, you should access:
peak_count : the peak fire count per year
peak_month : the month in which the peak fire count occured in a particular year
7.2.3 Datasets¶
We suggest that the datasets you use of this analysis, following Chen at al. (2011), are:
- MODIS Terra fire counts (2001-2011) (MOD14CMH). The particular dataset you will want from the file is ‘SUBDATASET_2 [360x720] CloudCorrFirePix (16-bit integer)’.
- Climate index data from NOAA
If you ever wish to take this study further, you can find various other useful datasets such as these.
Fire Data¶
The MOD14CMH CMG data are available from the UMD ftp server but have also been packaged for you as
If for any reason, you did want to copy or update them, use the following unix command:
cd Chapter7_ENSO/data;wget 'ftp://fire:burnt@fuoco.geog.umd.edu/modis/C5/cmg/monthly/hdf/*'
(though you may need to update the password). The data need to be placed in data and are in HDF format, so you should know how to read them into a numpy array in python.
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
!ls data/*hdf
data/MOD14CMH.200011.005.01.hdf data/MOD14CMH.201507.005.01.hdf data/MOD14CMH.200012.005.01.hdf data/MOD14CMH.201508.005.01.hdf data/MOD14CMH.200101.005.01.hdf data/MOD14CMH.201509.005.01.hdf data/MOD14CMH.200102.005.01.hdf data/MOD14CMH.201510.005.01.hdf data/MOD14CMH.200103.005.01.hdf data/MOD14CMH.201511.005.01.hdf data/MOD14CMH.200104.005.01.hdf data/MOD14CMH.201512.005.01.hdf data/MOD14CMH.200105.005.01.hdf data/MOD14CMH.201601.005.01.hdf data/MOD14CMH.200106.005.01.hdf data/MOD14CMH.201602.005.01.hdf data/MOD14CMH.200107.005.01.hdf data/MOD14CMH.201603.005.01.hdf data/MOD14CMH.200108.005.01.hdf data/MOD14CMH.201604.005.01.hdf data/MOD14CMH.200109.005.01.hdf data/MYD14CMH.200207.005.01.hdf data/MOD14CMH.200110.005.01.hdf data/MYD14CMH.200208.005.01.hdf data/MOD14CMH.200111.005.01.hdf data/MYD14CMH.200209.005.01.hdf data/MOD14CMH.200112.005.01.hdf data/MYD14CMH.200210.005.01.hdf data/MOD14CMH.200201.005.01.hdf data/MYD14CMH.200211.005.01.hdf data/MOD14CMH.200202.005.01.hdf data/MYD14CMH.200212.005.01.hdf data/MOD14CMH.200203.005.01.hdf data/MYD14CMH.200301.005.01.hdf data/MOD14CMH.200204.005.01.hdf data/MYD14CMH.200302.005.01.hdf data/MOD14CMH.200205.005.01.hdf data/MYD14CMH.200303.005.01.hdf data/MOD14CMH.200206.005.01.hdf data/MYD14CMH.200304.005.01.hdf data/MOD14CMH.200207.005.01.hdf data/MYD14CMH.200305.005.01.hdf data/MOD14CMH.200208.005.01.hdf data/MYD14CMH.200306.005.01.hdf data/MOD14CMH.200209.005.01.hdf data/MYD14CMH.200307.005.01.hdf data/MOD14CMH.200210.005.01.hdf data/MYD14CMH.200308.005.01.hdf data/MOD14CMH.200211.005.01.hdf data/MYD14CMH.200309.005.01.hdf data/MOD14CMH.200212.005.01.hdf data/MYD14CMH.200310.005.01.hdf data/MOD14CMH.200301.005.01.hdf data/MYD14CMH.200311.005.01.hdf data/MOD14CMH.200302.005.01.hdf data/MYD14CMH.200312.005.01.hdf data/MOD14CMH.200303.005.01.hdf data/MYD14CMH.200401.005.01.hdf data/MOD14CMH.200304.005.01.hdf data/MYD14CMH.200402.005.01.hdf data/MOD14CMH.200305.005.01.hdf data/MYD14CMH.200403.005.01.hdf data/MOD14CMH.200306.005.01.hdf data/MYD14CMH.200404.005.01.hdf data/MOD14CMH.200307.005.01.hdf data/MYD14CMH.200405.005.01.hdf data/MOD14CMH.200308.005.01.hdf data/MYD14CMH.200406.005.01.hdf data/MOD14CMH.200309.005.01.hdf data/MYD14CMH.200407.005.01.hdf data/MOD14CMH.200310.005.01.hdf data/MYD14CMH.200408.005.01.hdf data/MOD14CMH.200311.005.01.hdf data/MYD14CMH.200409.005.01.hdf data/MOD14CMH.200312.005.01.hdf data/MYD14CMH.200410.005.01.hdf data/MOD14CMH.200401.005.01.hdf data/MYD14CMH.200411.005.01.hdf data/MOD14CMH.200402.005.01.hdf data/MYD14CMH.200412.005.01.hdf data/MOD14CMH.200403.005.01.hdf data/MYD14CMH.200501.005.01.hdf data/MOD14CMH.200404.005.01.hdf data/MYD14CMH.200502.005.01.hdf data/MOD14CMH.200405.005.01.hdf data/MYD14CMH.200503.005.01.hdf data/MOD14CMH.200406.005.01.hdf data/MYD14CMH.200504.005.01.hdf data/MOD14CMH.200407.005.01.hdf data/MYD14CMH.200505.005.01.hdf data/MOD14CMH.200408.005.01.hdf data/MYD14CMH.200506.005.01.hdf data/MOD14CMH.200409.005.01.hdf data/MYD14CMH.200507.005.01.hdf data/MOD14CMH.200410.005.01.hdf data/MYD14CMH.200508.005.01.hdf data/MOD14CMH.200411.005.01.hdf data/MYD14CMH.200509.005.01.hdf data/MOD14CMH.200412.005.01.hdf data/MYD14CMH.200510.005.01.hdf data/MOD14CMH.200501.005.01.hdf data/MYD14CMH.200511.005.01.hdf data/MOD14CMH.200502.005.01.hdf data/MYD14CMH.200512.005.01.hdf data/MOD14CMH.200503.005.01.hdf data/MYD14CMH.200601.005.01.hdf data/MOD14CMH.200504.005.01.hdf data/MYD14CMH.200602.005.01.hdf data/MOD14CMH.200505.005.01.hdf data/MYD14CMH.200603.005.01.hdf data/MOD14CMH.200506.005.01.hdf data/MYD14CMH.200604.005.01.hdf data/MOD14CMH.200507.005.01.hdf data/MYD14CMH.200605.005.01.hdf data/MOD14CMH.200508.005.01.hdf data/MYD14CMH.200606.005.01.hdf data/MOD14CMH.200509.005.01.hdf data/MYD14CMH.200607.005.01.hdf data/MOD14CMH.200510.005.01.hdf data/MYD14CMH.200608.005.01.hdf data/MOD14CMH.200511.005.01.hdf data/MYD14CMH.200609.005.01.hdf data/MOD14CMH.200512.005.01.hdf data/MYD14CMH.200610.005.01.hdf data/MOD14CMH.200601.005.01.hdf data/MYD14CMH.200611.005.01.hdf data/MOD14CMH.200602.005.01.hdf data/MYD14CMH.200612.005.01.hdf data/MOD14CMH.200603.005.01.hdf data/MYD14CMH.200701.005.01.hdf data/MOD14CMH.200604.005.01.hdf data/MYD14CMH.200702.005.01.hdf data/MOD14CMH.200605.005.01.hdf data/MYD14CMH.200703.005.01.hdf data/MOD14CMH.200606.005.01.hdf data/MYD14CMH.200704.005.01.hdf data/MOD14CMH.200607.005.01.hdf data/MYD14CMH.200705.005.01.hdf data/MOD14CMH.200608.005.01.hdf data/MYD14CMH.200706.005.01.hdf data/MOD14CMH.200609.005.01.hdf data/MYD14CMH.200707.005.01.hdf data/MOD14CMH.200610.005.01.hdf data/MYD14CMH.200708.005.01.hdf data/MOD14CMH.200611.005.01.hdf data/MYD14CMH.200709.005.01.hdf data/MOD14CMH.200612.005.01.hdf data/MYD14CMH.200710.005.01.hdf data/MOD14CMH.200701.005.01.hdf data/MYD14CMH.200711.005.01.hdf data/MOD14CMH.200702.005.01.hdf data/MYD14CMH.200712.005.01.hdf data/MOD14CMH.200703.005.01.hdf data/MYD14CMH.200801.005.01.hdf data/MOD14CMH.200704.005.01.hdf data/MYD14CMH.200802.005.01.hdf data/MOD14CMH.200705.005.01.hdf data/MYD14CMH.200803.005.01.hdf data/MOD14CMH.200706.005.01.hdf data/MYD14CMH.200804.005.01.hdf data/MOD14CMH.200707.005.01.hdf data/MYD14CMH.200805.005.01.hdf data/MOD14CMH.200708.005.01.hdf data/MYD14CMH.200806.005.01.hdf data/MOD14CMH.200709.005.01.hdf data/MYD14CMH.200807.005.01.hdf data/MOD14CMH.200710.005.01.hdf data/MYD14CMH.200808.005.01.hdf data/MOD14CMH.200711.005.01.hdf data/MYD14CMH.200809.005.01.hdf data/MOD14CMH.200712.005.01.hdf data/MYD14CMH.200810.005.01.hdf data/MOD14CMH.200801.005.01.hdf data/MYD14CMH.200811.005.01.hdf data/MOD14CMH.200802.005.01.hdf data/MYD14CMH.200812.005.01.hdf data/MOD14CMH.200803.005.01.hdf data/MYD14CMH.200901.005.01.hdf data/MOD14CMH.200804.005.01.hdf data/MYD14CMH.200902.005.01.hdf data/MOD14CMH.200805.005.01.hdf data/MYD14CMH.200903.005.01.hdf data/MOD14CMH.200806.005.01.hdf data/MYD14CMH.200904.005.01.hdf data/MOD14CMH.200807.005.01.hdf data/MYD14CMH.200905.005.01.hdf data/MOD14CMH.200808.005.01.hdf data/MYD14CMH.200906.005.01.hdf data/MOD14CMH.200809.005.01.hdf data/MYD14CMH.200907.005.01.hdf data/MOD14CMH.200810.005.01.hdf data/MYD14CMH.200908.005.01.hdf data/MOD14CMH.200811.005.01.hdf data/MYD14CMH.200909.005.01.hdf data/MOD14CMH.200812.005.01.hdf data/MYD14CMH.200910.005.01.hdf data/MOD14CMH.200901.005.01.hdf data/MYD14CMH.200911.005.01.hdf data/MOD14CMH.200902.005.01.hdf data/MYD14CMH.200912.005.01.hdf data/MOD14CMH.200903.005.01.hdf data/MYD14CMH.201001.005.01.hdf data/MOD14CMH.200904.005.01.hdf data/MYD14CMH.201002.005.01.hdf data/MOD14CMH.200905.005.01.hdf data/MYD14CMH.201003.005.01.hdf data/MOD14CMH.200906.005.01.hdf data/MYD14CMH.201004.005.01.hdf data/MOD14CMH.200907.005.01.hdf data/MYD14CMH.201005.005.01.hdf data/MOD14CMH.200908.005.01.hdf data/MYD14CMH.201006.005.01.hdf data/MOD14CMH.200909.005.01.hdf data/MYD14CMH.201007.005.01.hdf data/MOD14CMH.200910.005.01.hdf data/MYD14CMH.201008.005.01.hdf data/MOD14CMH.200911.005.01.hdf data/MYD14CMH.201009.005.01.hdf data/MOD14CMH.200912.005.01.hdf data/MYD14CMH.201010.005.01.hdf data/MOD14CMH.201001.005.01.hdf data/MYD14CMH.201011.005.01.hdf data/MOD14CMH.201002.005.01.hdf data/MYD14CMH.201012.005.01.hdf data/MOD14CMH.201003.005.01.hdf data/MYD14CMH.201101.005.01.hdf data/MOD14CMH.201004.005.01.hdf data/MYD14CMH.201102.005.01.hdf data/MOD14CMH.201005.005.01.hdf data/MYD14CMH.201103.005.01.hdf data/MOD14CMH.201006.005.01.hdf data/MYD14CMH.201104.005.01.hdf data/MOD14CMH.201007.005.01.hdf data/MYD14CMH.201105.005.01.hdf data/MOD14CMH.201008.005.01.hdf data/MYD14CMH.201106.005.01.hdf data/MOD14CMH.201009.005.01.hdf data/MYD14CMH.201107.005.01.hdf data/MOD14CMH.201010.005.01.hdf data/MYD14CMH.201108.005.01.hdf data/MOD14CMH.201011.005.01.hdf data/MYD14CMH.201109.005.01.hdf data/MOD14CMH.201012.005.01.hdf data/MYD14CMH.201110.005.01.hdf data/MOD14CMH.201101.005.01.hdf data/MYD14CMH.201111.005.01.hdf data/MOD14CMH.201102.005.01.hdf data/MYD14CMH.201112.005.01.hdf data/MOD14CMH.201103.005.01.hdf data/MYD14CMH.201201.005.01.hdf data/MOD14CMH.201104.005.01.hdf data/MYD14CMH.201202.005.01.hdf data/MOD14CMH.201105.005.01.hdf data/MYD14CMH.201203.005.01.hdf data/MOD14CMH.201106.005.01.hdf data/MYD14CMH.201204.005.01.hdf data/MOD14CMH.201107.005.01.hdf data/MYD14CMH.201205.005.01.hdf data/MOD14CMH.201108.005.01.hdf data/MYD14CMH.201206.005.01.hdf data/MOD14CMH.201109.005.01.hdf data/MYD14CMH.201207.005.01.hdf data/MOD14CMH.201110.005.01.hdf data/MYD14CMH.201208.005.01.hdf data/MOD14CMH.201111.005.01.hdf data/MYD14CMH.201209.005.01.hdf data/MOD14CMH.201112.005.01.hdf data/MYD14CMH.201210.005.01.hdf data/MOD14CMH.201201.005.01.hdf data/MYD14CMH.201211.005.01.hdf data/MOD14CMH.201202.005.01.hdf data/MYD14CMH.201212.005.01.hdf data/MOD14CMH.201203.005.01.hdf data/MYD14CMH.201301.005.01.hdf data/MOD14CMH.201204.005.01.hdf data/MYD14CMH.201302.005.01.hdf data/MOD14CMH.201205.005.01.hdf data/MYD14CMH.201303.005.01.hdf data/MOD14CMH.201206.005.01.hdf data/MYD14CMH.201304.005.01.hdf data/MOD14CMH.201207.005.01.hdf data/MYD14CMH.201305.005.01.hdf data/MOD14CMH.201208.005.01.hdf data/MYD14CMH.201306.005.01.hdf data/MOD14CMH.201209.005.01.hdf data/MYD14CMH.201307.005.01.hdf data/MOD14CMH.201210.005.01.hdf data/MYD14CMH.201308.005.01.hdf data/MOD14CMH.201211.005.01.hdf data/MYD14CMH.201309.005.01.hdf data/MOD14CMH.201212.005.01.hdf data/MYD14CMH.201310.005.01.hdf data/MOD14CMH.201301.005.01.hdf data/MYD14CMH.201311.005.01.hdf data/MOD14CMH.201302.005.01.hdf data/MYD14CMH.201312.005.01.hdf data/MOD14CMH.201303.005.01.hdf data/MYD14CMH.201401.005.01.hdf data/MOD14CMH.201304.005.01.hdf data/MYD14CMH.201402.005.01.hdf data/MOD14CMH.201305.005.01.hdf data/MYD14CMH.201403.005.01.hdf data/MOD14CMH.201306.005.01.hdf data/MYD14CMH.201404.005.01.hdf data/MOD14CMH.201307.005.01.hdf data/MYD14CMH.201405.005.01.hdf data/MOD14CMH.201308.005.01.hdf data/MYD14CMH.201406.005.01.hdf data/MOD14CMH.201309.005.01.hdf data/MYD14CMH.201407.005.01.hdf data/MOD14CMH.201310.005.01.hdf data/MYD14CMH.201408.005.01.hdf data/MOD14CMH.201311.005.01.hdf data/MYD14CMH.201409.005.01.hdf data/MOD14CMH.201312.005.01.hdf data/MYD14CMH.201410.005.01.hdf data/MOD14CMH.201401.005.01.hdf data/MYD14CMH.201411.005.01.hdf data/MOD14CMH.201402.005.01.hdf data/MYD14CMH.201412.005.01.hdf data/MOD14CMH.201403.005.01.hdf data/MYD14CMH.201501.005.01.hdf data/MOD14CMH.201404.005.01.hdf data/MYD14CMH.201502.005.01.hdf data/MOD14CMH.201405.005.01.hdf data/MYD14CMH.201503.005.01.hdf data/MOD14CMH.201406.005.01.hdf data/MYD14CMH.201504.005.01.hdf data/MOD14CMH.201407.005.01.hdf data/MYD14CMH.201505.005.01.hdf data/MOD14CMH.201408.005.01.hdf data/MYD14CMH.201506.005.01.hdf data/MOD14CMH.201409.005.01.hdf data/MYD14CMH.201507.005.01.hdf data/MOD14CMH.201410.005.01.hdf data/MYD14CMH.201508.005.01.hdf data/MOD14CMH.201411.005.01.hdf data/MYD14CMH.201509.005.01.hdf data/MOD14CMH.201412.005.01.hdf data/MYD14CMH.201510.005.01.hdf data/MOD14CMH.201501.005.01.hdf data/MYD14CMH.201511.005.01.hdf data/MOD14CMH.201502.005.01.hdf data/MYD14CMH.201512.005.01.hdf data/MOD14CMH.201503.005.01.hdf data/MYD14CMH.201601.005.01.hdf data/MOD14CMH.201504.005.01.hdf data/MYD14CMH.201602.005.01.hdf data/MOD14CMH.201505.005.01.hdf data/MYD14CMH.201603.005.01.hdf data/MOD14CMH.201506.005.01.hdf data/MYD14CMH.201604.005.01.hdf
If you are really stuck on reading the data, or just want to move on to the next parts, you can use python/reader.py which will create a masked array in data, and an array of years (year) and months (month):
run python/reader
# refine the data mask
data.mask = data.data.sum(axis=0) == 0
# the reader script loads the
# firecount data into the data array
# shape: (nmonths,nlat,nlon)
print 'data shape',data.shape
# data is a masked array:
print 'data type',type(data)
# so has a mask:
print 'mask shape',data.mask.shape
# also: year and month
print 'year and month shape',year.shape,month.shape
# display the firecount data
plt.figure(figsize=(10,6))
x = year + month/12.
y = np.sum(data,axis=(1,2))
plt.plot(x,y)
plt.ylabel('global fire count')
plt.xlabel('time')
plt.title('global fire count')
plt.xlim(x[0],x[-1])
fyear,fmonth = year,month
data shape (186, 360, 720) data type <class 'numpy.ma.core.MaskedArray'> mask shape (186, 360, 720) year and month shape (186,) (186,)
To use the dataset here, we will need to shrink the dataset by a factor of 10 then, to a 5 degrees dataset.
There are different ways to achive this, but one way would be to reoganise the data:
plt.figure(figsize=(10,5))
plt.imshow(data.mask[0])
<matplotlib.image.AxesImage at 0x13703ee10>
plt.figure(figsize=(10,5))
plt.imshow(rdata.sum(axis=0)[0],interpolation='none')
<matplotlib.image.AxesImage at 0x12e5ec7d0>
# how does this work: do it in stages to try to understand this
# NB -- all we are trying to achieve is a reduction in the
# dataset resolution by summingover each sun 10 x 10 cells
rdata = np.array([data[:,i::10,j::10] for i in xrange(10) for j in xrange(10)])
# sum over 1st axis to achieve reduced resolution summation
rdata = rdata.sum(axis=0)
nslots,nlat,nlon = rdata.shape
# set a min count
mincount = 1000
#build a mask
all_mask = rdata.sum(axis=0) <= mincount
# repeat it for each frame
rmask = np.repeat(all_mask.T,nslots).reshape((nlon,nlat,nslots)).T
plt.figure(figsize=(10,5))
print rmask.shape
plt.imshow(rmask[0],interpolation='none')
fdata = ma.array(rdata,mask=rmask)
plt.figure(figsize=(10,5))
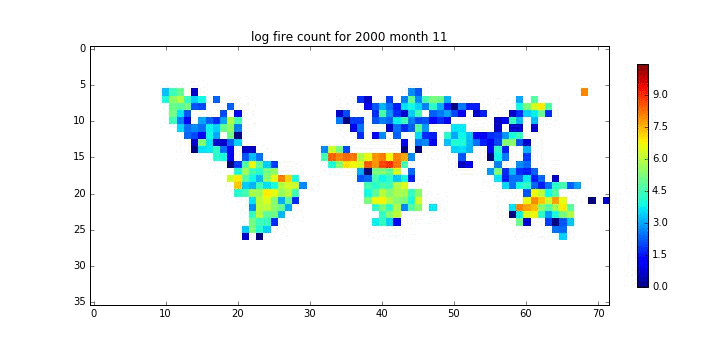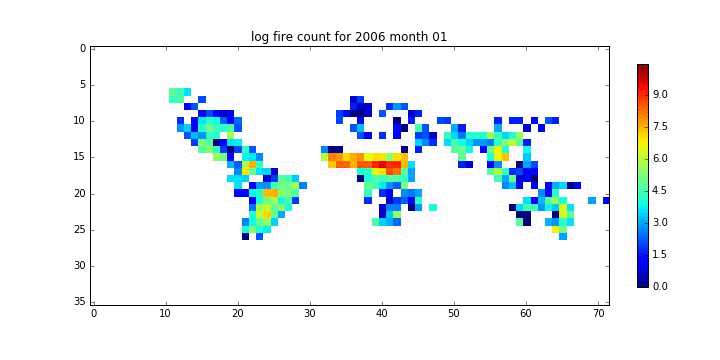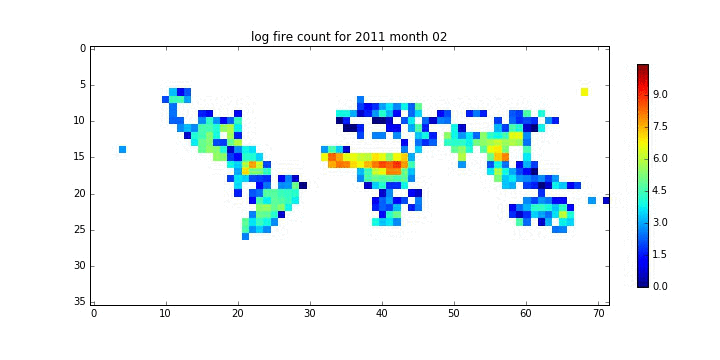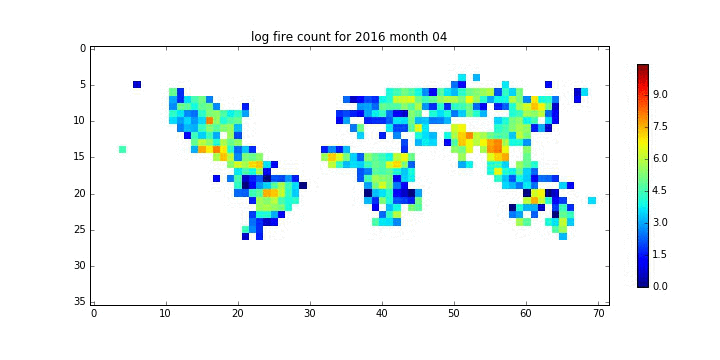
plt.title('log fire count for %d month %02d'%(year[10],month[10]))
plt.imshow(np.ma.log(rdata[10]),interpolation='none')
plt.colorbar(fraction=0.02)
(186, 36, 72)
<matplotlib.colorbar.Colorbar at 0x135ef1150>
# or even make a movie
lf = np.ma.log(fdata)
vmax = np.max(lf[lf>0])
root = 'images/'
print lf.shape
# loop over lf.shape[0] (time samples)
for i in xrange(lf.shape[0]):
print i,
fig = plt.figure(figsize=(10,5))
plt.imshow(np.ma.log(fdata[i]),interpolation='nearest',vmax=vmax)
plt.colorbar(fraction=0.02)
file_id = '%d month %02d'%(year[i],month[i])
plt.title('log fire count for %s'%file_id)
plt.savefig('%s_%s.jpg'%(root,file_id.replace(' ','_')))
plt.close(fig)
(186, 36, 72)
# make a movie from the files
cmd = 'convert -delay 100 -loop 0 {0}_*month*.jpg {0}fire_movie3.gif'.format(root)
os.system(cmd)
0
A. Sort the information required¶
What information do we need to derive from the fire dataset?
There are several ways we could approach the problem, but a simple method might be to first find the 'typical' peak fire month per geographical location. We could call that the 'fire month'.
To calculate that, we could find which month in each (full) year the maximum fire count occurs in.
A useful method for that will be np.argmax() that gives the argument ('index' if you like) of the maximum value in an array.
data = np.random.rand(100)
amax = np.argmax(data)
print 'the maxiumum value in the data array is',data[amax],'that occurs at sample index',amax
the maxiumum value in the data array is 0.977845430318 that occurs at sample index 34
Once we have defined the spatial distribution of fire month, we can calculate an annual dataset containing the firecount (for the fire month) for each year and location.
plt.figure(figsize=(10,5))
plt.imshow(fdata.mask[0],interpolation='nearest')
plt.colorbar(fraction=0.02)
<matplotlib.colorbar.Colorbar at 0x175908dd0>
# find complete years
# the list of years
good_year = []
for y in np.unique(fyear):
if (fyear == y).sum() == 12:
# then we have a full year
good_year.append(y)
good_year = np.array(good_year)
print good_year
[2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 2012 2013 2014 2015]
# so for these years, find the max fire count
# and the associated month
# the list of years
good_year = []
month_data = []
for y in np.unique(fyear):
if (fyear == y).sum() == 12:
# then we have a full year
good_year.append(y)
# this gives the month in this year where firecount is maximum
year_data = np.ma.array(np.ma.argmax(fdata[fyear == y],axis=0),\
mask=fdata.mask[0])
month_data.append(year_data)
# now find the e.g. median of these to get th erepresetative fire month ...
month_data = np.median(np.ma.array(month_data),axis=0).astype(int)
plt.figure(figsize=(10,5))
plt.imshow(month_data,interpolation='nearest',vmax=11)
plt.colorbar(fraction=0.02)
plt.title('fire month')
<matplotlib.text.Text at 0x1a0fae590>
# now generate the fire_count data for each year in an array fire_count
fire_count = np.zeros((len(good_year),nlat,nlon))
for iy,y in enumerate(good_year):
year_data = np.ma.array(fdata[fyear == y])
for m in np.arange(12):
r,c = np.where(month_data == m)
fire_count[iy,r,c] = year_data[m,r,c]
fire_count_mask = fire_count == 0
fire_count = np.ma.array(fire_count,mask=fire_count_mask)
# plot a few years data
for y in xrange(2):
plt.figure(figsize=(10,5))
plt.imshow(np.ma.log(fire_count[y]),interpolation='nearest')
plt.colorbar(fraction=0.02)
plt.title('log max fire count %d'%good_year[y])
# plot for one location
# the biggest one ...
y,r,c = np.where(fire_count == np.ma.max(fire_count))
plt.figure(figsize=(10,5))
plt.title('log fire count for pixel %d %d'%(r,c))
plt.plot(good_year,np.log(fire_count[:,r,c]))
plt.xlabel('year')
plt.ylabel('log fire count')
<matplotlib.text.Text at 0x1a3e30fd0>
print fire_count.shape
print len(good_year)
print month_data.shape
(15, 36, 72) 15 (36, 72)
In summary, we now have the fire month stored in month_data, and, for each year (in good_year), the fire count for that month in fire_count.
This is the core information we require for modelling.
We now consider the associated climate datasets.
Climate Data¶
The climate data you will want will be some form of Sea Surface Temperature (SST) anomaly measure. There is a long list of such measures on http://www.esrl.noaa.gov/psd/data/climateindices/list.
Examples would be AMO or ONI. Note that some of these measures are smoothed and others not.
When you see a new dataset, you should investigate the formatting and other issues before trying to interpret it.
Whilst learning coding, it is often worthwhile doing the 'low level' reading and formatting yourself. We provide 2 examples of this below.
There are simpler ways of reading in tabular ('csv') datasets, so we show two of these as well.
First, lets explore the dataset format:
Suppose we had selected AMO and we want to read directly from the url:
import urllib2
url = 'http://www.esrl.noaa.gov/psd/data/correlation/amon.us.data'
# as a backup, in case the noaa server is not responding,
# you can access the data from the data directory or
# https://github.com/profLewis/geogg122/blob/master/Chapter7_ENSO/data/amon.us.data
req = urllib2.Request ( url )
raw_data = urllib2.urlopen(req).readlines()
Now examine the data format and work out how to access the information we want.
# look at the first 2 lines of the data we pull from the url
#
# The first line gives the dates range
#
# The data values appear on subsequent lines
print 'first 2 lines:'
print raw_data[:2]
# years: integer array
year_range = np.array(raw_data[0].split()).astype(int)
print 'years range:',year_range
# look at the end of the data:
print 'last 5 lines:'
print raw_data[-5:]
# so the last 4 lines of the data contains
# dataset information
fill_value = float(raw_data[-4])
print 'fill value:',fill_value
data_name = raw_data[-3]
print'data name:',data_name
data_info = raw_data[-2]
print'data info:',data_info
data_url = raw_data[-1]
print'data url:',data_url
#
# we notice from inspection that
# we want data from rows 1 to -4
# so raw_data[1:-4]
# set this as data_str, the string representation of the data
data_str = raw_data[1:-4]
print 'length of data_str:',len(data_str)
first 2 lines: [' 1948 2016\n', ' 1948 -0.009 -0.021 0.034 -0.064 0.002 0.061 -0.033 -0.016 -0.046 0.014 0.140 0.069\n'] years range: [1948 2016] last 5 lines: [' 2016 0.243 0.167 0.200 0.189 0.356 0.421 0.444 0.468 -99.990 -99.990 -99.990 -99.990\n', ' -99.99\n', ' AMO unsmoothed, detrended from the Kaplan SST V2\n', ' Calculated at NOAA/ESRL/PSD1\n', ' http://www.esrl.noaa.gov/psd/data/timeseries/AMO/\n'] fill value: -99.99 data name: AMO unsmoothed, detrended from the Kaplan SST V2 data info: Calculated at NOAA/ESRL/PSD1 data url: http://www.esrl.noaa.gov/psd/data/timeseries/AMO/ length of data_str: 69
# so to read the data we loop over each line
# Version 1:
#
# we could do this in a loop of this sort:
# We want the data in an array of shape (nmonths,nyears)
# so initialise this
nyears = year_range[-1] - year_range[0] + 1
nmonths = 12
first_year = year_range[0]
print 'nyears, nmonths:',nyears,nmonths
print 'first year:',first_year
cdata = np.zeros((nmonths,nyears))
cyears = np.zeros((nyears))
# loop over all lines in data_str
for i in xrange(len(data_str)):
# load the string data for thatr line
line_data = data_str[i]
# split it into an array of (13) floats
fline_data = np.array(line_data.split()).astype(float)
# the first entry is the year
this_year = int(fline_data[0])
# give the user info on which years we read
print this_year,
cyears[i] = this_year
# the rest of the line is the 12 data entries per month
year_data = fline_data[1:]
# the year index is this_year minus first_year
year_index = this_year - first_year
print '(%d),'%year_index,
# store that in cdata
cdata[:,year_index] = year_data
# now, generate a mask of the fill values
cmask = (cdata == fill_value)
# where are the duff (fill) values?
# (why would this be?)
print np.where(cmask)
nyears, nmonths: 69 12 first year: 1948 1948 (0), 1949 (1), 1950 (2), 1951 (3), 1952 (4), 1953 (5), 1954 (6), 1955 (7), 1956 (8), 1957 (9), 1958 (10), 1959 (11), 1960 (12), 1961 (13), 1962 (14), 1963 (15), 1964 (16), 1965 (17), 1966 (18), 1967 (19), 1968 (20), 1969 (21), 1970 (22), 1971 (23), 1972 (24), 1973 (25), 1974 (26), 1975 (27), 1976 (28), 1977 (29), 1978 (30), 1979 (31), 1980 (32), 1981 (33), 1982 (34), 1983 (35), 1984 (36), 1985 (37), 1986 (38), 1987 (39), 1988 (40), 1989 (41), 1990 (42), 1991 (43), 1992 (44), 1993 (45), 1994 (46), 1995 (47), 1996 (48), 1997 (49), 1998 (50), 1999 (51), 2000 (52), 2001 (53), 2002 (54), 2003 (55), 2004 (56), 2005 (57), 2006 (58), 2007 (59), 2008 (60), 2009 (61), 2010 (62), 2011 (63), 2012 (64), 2013 (65), 2014 (66), 2015 (67), 2016 (68), (array([ 8, 9, 10, 11]), array([68, 68, 68, 68]))
We can see that the dataset has no gaps in year, so instead of working out year_index in the loop, we could more simply use the index i.
Actually, we could read the data in in a more 'pythonic' manner with a much simpler loop:
# so to read the data we loop over each line
# Version 2:
# form float array of all 13 data columns
cdata = np.array([r.split() for r in raw_data[1:-4]]).astype(float).T
# check the shape is how we want it
print 'shape:',cdata.shape
# separate the data and year values
cyears = cdata[0]
cdata = cdata[1:]
# make a mask for fill values
cmask = (cdata == fill_value)
# put cdata in a masked array
cdata = ma.array(cdata,mask=cmask)
print cdata.shape
print 'fill values:',fill_value,np.where(cmask)
shape: (13, 69) (12, 69) fill values: -99.99 (array([ 8, 9, 10, 11]), array([68, 68, 68, 68]))
# so to read the data we loop over each line
# Version 3:
# use numpy.genfromtxt
# see http://docs.scipy.org/doc/numpy/reference/generated/numpy.genfromtxt.html
cdata = np.genfromtxt(raw_data,skip_header=1,skip_footer=4,unpack=True,usemask=True)
# In this case, need to specify and fix fill value
cdata.fill_value=-99.99
cdata.mask[cdata.data==cdata.fill_value] = True
# separate the years column and the data
cyears = cdata[0]
cdata = cdata[1:]
print cdata.shape
(12, 69)
fig = plt.figure(figsize=(15,3))
plt.plot(cyears,cdata[0])
plt.xlabel('year')
plt.ylabel('January AMO')
plt.xlim(year_range[0],year_range[1])
(1948, 2016)
# so to read the data we loop over each line
# Version 4:
#
# use pandas to do the whole thing from the url
# pandas is a useful package for dealing with and plotting tabular data
# that you may like to consider
import pandas as pd
# see http://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html
# and http://pandas.pydata.org/pandas-docs/stable/tutorials.html
c=pd.read_csv(url,skiprows=1,\
header=None,delim_whitespace=True,skipfooter=4,index_col=0,engine='python').T
c[c == -99.99] = np.nan
print c.shape
# now display the dataset in a table
#print c.info()
print 'keys:',c.keys()
# display option
pd.set_option('display.max_columns', 10)
# show table
c
(12, 69)
keys: Int64Index([1948, 1949, 1950, 1951, 1952, 1953, 1954, 1955, 1956, 1957, 1958,
1959, 1960, 1961, 1962, 1963, 1964, 1965, 1966, 1967, 1968, 1969,
1970, 1971, 1972, 1973, 1974, 1975, 1976, 1977, 1978, 1979, 1980,
1981, 1982, 1983, 1984, 1985, 1986, 1987, 1988, 1989, 1990, 1991,
1992, 1993, 1994, 1995, 1996, 1997, 1998, 1999, 2000, 2001, 2002,
2003, 2004, 2005, 2006, 2007, 2008, 2009, 2010, 2011, 2012, 2013,
2014, 2015, 2016],
dtype='int64', name=0)
| 1948 | 1949 | 1950 | 1951 | 1952 | ... | 2012 | 2013 | 2014 | 2015 | 2016 | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | -0.009 | 0.159 | 0.117 | 0.109 | 0.178 | ... | -0.053 | 0.140 | -0.050 | 0.004 | 0.243 |
| 2 | -0.021 | 0.166 | -0.027 | 0.005 | 0.189 | ... | 0.016 | 0.127 | -0.031 | 0.008 | 0.167 |
| 3 | 0.034 | 0.045 | -0.099 | 0.019 | 0.237 | ... | 0.037 | 0.170 | -0.069 | -0.117 | 0.200 |
| 4 | -0.064 | 0.105 | -0.124 | 0.175 | 0.198 | ... | 0.092 | 0.148 | -0.082 | -0.060 | 0.189 |
| 5 | 0.002 | -0.018 | -0.052 | 0.179 | 0.187 | ... | 0.176 | 0.113 | 0.010 | 0.056 | 0.356 |
| 6 | 0.061 | 0.007 | -0.036 | 0.296 | 0.389 | ... | 0.312 | 0.058 | 0.074 | 0.041 | 0.421 |
| 7 | -0.033 | 0.078 | -0.049 | 0.428 | 0.379 | ... | 0.387 | 0.202 | 0.233 | 0.144 | 0.444 |
| 8 | -0.016 | 0.109 | 0.027 | 0.310 | 0.410 | ... | 0.443 | 0.206 | 0.346 | 0.190 | 0.468 |
| 9 | -0.046 | 0.076 | 0.016 | 0.256 | 0.368 | ... | 0.460 | 0.267 | 0.322 | 0.312 | NaN |
| 10 | 0.014 | 0.110 | -0.084 | 0.266 | 0.361 | ... | 0.341 | 0.359 | 0.304 | 0.336 | NaN |
| 11 | 0.140 | 0.114 | 0.086 | 0.181 | 0.257 | ... | 0.177 | 0.140 | 0.077 | 0.198 | NaN |
| 12 | 0.069 | 0.124 | 0.095 | 0.181 | 0.347 | ... | 0.153 | 0.048 | 0.070 | 0.241 | NaN |
12 rows × 69 columns
ax = c.T.plot.line(figsize=(15,5),title='ENSO AMO')
ax.set_xlabel('years')
<matplotlib.text.Text at 0x185db1610>
# to just dump it as a numpy array if thats all you want to do ...
cdata = np.array(pd.DataFrame(c))
cyears = np.array(c.keys()).astype(int)
year_range = np.array([cyears[0],cyears[-1]])
fig = plt.figure(figsize=(15,3))
plt.plot(cyears,cdata[0])
plt.xlabel('year')
plt.ylabel('January AMO')
plt.xlim(year_range[0],year_range[1])
(1948, 2016)
7.2.4 Data Processing¶
The fire information we have is the peak fire count and the month this occurred in, for each location and year.
The climate data gives the monthly ENSO-like variable for each month and year.
# relevant datasets
print '------- Fire -----------'
print fire_count.shape
print len(good_year)
print month_data.shape
print '------- Climate --------'
print cyears.shape
print cdata.shape
------- Fire ----------- (15, 36, 72) 15 (36, 72) ------- Climate -------- (69,) (12, 69)
We now want to form datasets matching the ENSO index for N months prior to the fire month to the peak fire activity.
We can prototype this for N = 0 to test the idea.
We appreciate in our design that N months before any month may lie in the previous year, so have to build a case for this:
# what offset to use
N = 0
enso = np.zeros_like(fire_count)
for iy,y in enumerate(good_year):
for m in np.arange(12):
r,c = np.where(month_data==m)
# which month in the climate dataset
climate_month = m - N
# account for possibility of month being in
# previous year
if climate_month < 0:
climate_month += 12
climate_year = y-1
else:
climate_year = y
yy = np.where(cyears == y)[0]
# pull the climate value for this month and year
climate_data = cdata[climate_month,yy]
# store in enso
enso[y-good_year[0],r,c] = climate_data
plt.figure(figsize=(10,5))
plt.imshow(enso[0],interpolation='nearest')
plt.colorbar(fraction=0.02)
plt.title('enso value for year %d at lag %d'%(good_year[0],N))
# these should match up
print enso.shape,fire_count.shape
(15, 36, 72) (15, 36, 72)
So, now we have the ENSO index dataset for lag month N and the fire count data.
Lets look at a plot of this two for a particular location:
import scipy.stats
# see http://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.linregress.html
y,r,c = np.where(fire_count == np.ma.max(fire_count))
enso_ = np.array(enso[:,r,c]).flatten()
fire_count_ = np.array(fire_count[:,r,c]).flatten()
plt.figure(figsize=(5,5))
plt.plot(enso_,fire_count_,'+')
plt.xlabel('ENSO')
plt.ylabel('fire count')
# linear regression
slope, intercept, r_value, p_value, std_err = scipy.stats.linregress(enso_,fire_count_)
print 'slope, intercept, r_value, p_value, std_err',slope, intercept, r_value, p_value, std_err
slope, intercept, r_value, p_value, std_err 1668.5039503 15592.6186334 0.0838460646882 0.766401291507 5499.72415493
We see there is a rather weak relationship between the two here, but that may just be the particular sample we have selected
Let's make a method to help wiith this:
def enso_data(N,cdata=cdata,fire_count=fire_count,good_year=good_year,\
month_data=month_data,cyears=cyears):
enso = np.zeros_like(fire_count)
for iy,y in enumerate(good_year):
for m in np.arange(12):
r,c = np.where(month_data==m)
# which month in the climate dataset
climate_month = m - N
# account for possibility of month being in
# previous year
if climate_month < 0:
climate_month += 12
climate_year = y-1
else:
climate_year = y
yy = np.where(cyears == y)[0]
# pull the climate value for this month and year
climate_data = cdata[climate_month,yy]
# store in enso
enso[y-good_year[0],r,c] = climate_data
return enso
y,r,c = np.where(fire_count == np.ma.max(fire_count))
fire_count_ = np.array(fire_count[:,r,c]).flatten()
r_2 = np.zeros(12)
for N in xrange(12):
enso = enso_data(N)
enso_ = np.array(enso[:,r,c]).flatten()
slope, intercept, r_value, p_value, std_err = scipy.stats.linregress(enso_,fire_count_)
r_2[N] = r_value**2
print np.argmax(r_2),r_2.max()
plt.plot(r_2)
plt.xlabel('lag month')
plt.ylabel('R^2')
8 0.312752086836
<matplotlib.text.Text at 0x12990e4d0>
So the highest correlation is achieved for N=8 for this location.
# do this spatially now
# so
r_2 = np.zeros((12,nlat,nlon))
for N in xrange(12):
enso = enso_data(N)
rs,cs = np.where(~fire_count.mask[0])
for i in xrange(len(rs)):
r,c = rs[i],cs[i]
enso_ = np.array(enso[:,r,c]).flatten()
fire_count_ = np.array(fire_count[:,r,c]).flatten()
# regress
slope, intercept, r_value, p_value, std_err = scipy.stats.linregress(enso_,fire_count_)
r_2[N,r,c] = r_value**2
lag_month = np.argmax(r_2,axis=0)
max_r2 = np.max(r_2,axis=0)
plt.figure(figsize=(10,5))
plt.imshow(lag_month,interpolation='nearest')
plt.colorbar(fraction=0.02)
plt.title('enso optimal lag month')
plt.figure(figsize=(10,5))
plt.imshow(np.sqrt(max_r2),interpolation='nearest')
plt.colorbar(fraction=0.02)
plt.title('enso/fire -r')
####### NEED TO EDIT BEYOND HERE
<matplotlib.text.Text at 0x17024de50>
7.2.4 Code to perform correlation analysis¶
The idea here is, for a particular (or set of) SST anomaly measures, work out which ‘lag’ month gives the highest correlation coefficient with fire count.
By ‘lag’ month, we mean that e.g. if the peak fire month for a particular pixel was September, which month prior to that has a set of SST anomalies over the sample years that is most strongly correlated with fire count.
So, if we were using a single SST anomaly measure (e.g. AMO or ONI) and sample years 2001 to 2009 to build our model, then we would do a linear regression of fire count for a particular pixel over these years against e.g. AMO data for September (lag 0) then August (lag 1) then July (lag 2) etc. and see which produced the highest $R^2$.
Before we get into that, let's look again at the data structure we have:
# climate data
print 'cdata',cdata.shape
print 'cyears',cyears.shape
# From the fire data
print 'peak_count',peak_count.shape
print 'av_fire_month',av_fire_month.shape
print 'min_year',min_year
So, if we want to select data for particular years:
# which years (inclusive)
years = [2001,2010]
ypeak_count = peak_count[years[0]-min_year:years[1] - min_year + 1]
ycdata = cdata[:,years[0] - cyears[0]:years[1] - cyears[0] + 1]
# check the shape
print ycdata.shape,ypeak_count.shape,av_fire_month.shape
We need to consider a little carefully the implementation of lag ...
# we will need to access ycdata[month - n][year]
# which is a bit fiddly as e.g. -3 will be interpreted as
# October for that same year, rather than the previous year
y = 2001 - min_year
m = 2
lag = 5
print m - lag,y
# so one way to fix this is to decrease y by one
# if m - lag is -ve
Y = y - (m - lag < 0)
print m-lag,Y
from scipy.stats import linregress
# examine an example row col
# for a given month over all years
c = 24
r = 19
m = av_fire_month[r,c]
# pull the data
yyears = np.arange(years[1]-years[0]+1)
R2 = np.array([linregress(\
ycdata[m-n,yyears - (m - n < 0)],\
ypeak_count[yyears - (m - n < 0),r,c]\
)[2] for n in xrange(12)])
n = np.argmax(R2)
x = ycdata[m-n,yyears - (m - n < 0)]
y = ypeak_count[yyears - (m - n < 0),r,c]
slope,intercept,R,p,err = linregress(x,y)
print slope,intercept,p,err
plt.plot(ycdata[m-n],y,'r+')
plt.xlabel('Climate Index')
plt.ylabel('Fire count')
plt.plot([x.min(),x.max()],\
[intercept+slope*x.min(),intercept+slope*x.max()],'k--')
plt.title('Fire count at r %03d c %03d: R^2 = %.3f: lag %d'%(r,c,R2[n],n))
# looper
data_mask = ypeak_count.sum(axis=0)>100
rs,cs = np.where(data_mask)
results = {'intercept':0,'slope':0,'p':0,'R':0,'stderr':0,'lag':0}
for k in results.keys():
results[k] = np.zeros_like(av_fire_month).astype(float)
results[k] = ma.array(results[k],mask=~data_mask)
for r,c in zip(rs,cs):
m = av_fire_month[r,c]
# pull the data
yyears = np.arange(years[1]-years[0]+1)
R2 = np.array([\
linregress(\
ycdata[m-n,yyears - (m - n < 0)],\
ypeak_count[yyears - (m - n < 0),r,c]\
)[2] for n in xrange(12)])
n = np.argmax(R2)
results['lag'][r,c] = n
x = ycdata[m-n,yyears - (m - n < 0)]
y = ypeak_count[yyears - (m - n < 0),r,c]
results['slope'][r,c],results['intercept'][r,c],\
results['R'][r,c],results['p'][r,c],\
results['stderr'][r,c] = linregress(x,y)
plt.figure(figsize=(10,4))
plt.imshow(results['R'],interpolation='nearest')
plt.colorbar()
plt.title('R')
plt.figure(figsize=(10,4))
plt.imshow(results['p'],interpolation='nearest')
plt.colorbar()
plt.title('p')
plt.figure(figsize=(10,4))
plt.imshow(results['slope'],interpolation='nearest')
plt.colorbar()
plt.title('slope')
plt.figure(figsize=(10,4))
plt.imshow(results['lag'],interpolation='nearest')
plt.colorbar()
plt.title('lag')
which we can now predict:
# prediction year
pyear = 2012
# which month?
M = av_fire_month - results['lag']
Y = np.zeros_like(M) + pyear
Y[M<0] -= 1
# lets look at that ...
plt.imshow(Y,interpolation='nearest')
# climate data
scdata = np.zeros_like(Y).astype(float)
for y in [pyear,pyear-1]:
for m in xrange(12):
scdata[(Y == y) & (M == m)] = cdata[m,y-cyears[0]]
plt.imshow(scdata,interpolation='nearest')
# now predict
fc_predict = results['intercept'] + results['slope'] * scdata
plt.figure(figsize=(10,4))
plt.imshow(fc_predict,interpolation='nearest',vmin=0,vmax=10000)
plt.colorbar()
plt.title('predicted peak fire count for %d'%pyear)
plt.figure(figsize=(10,4))
plt.imshow(peak_count[pyear-min_year],\
interpolation='nearest',vmin=0,vmax=10000)
plt.colorbar()
plt.title('actual peak fire count for %d'%pyear)
x = peak_count[pyear-min_year].flatten()
y = fc_predict.flatten()
slope,intercept,R,p,err = linregress(x,y)
plt.plot(x,y,'+')
plt.xlabel('measured fire count')
plt.ylabel('predicted fire count')
cc = np.array([0.,x.max()])
plt.plot(cc,cc,'k--')
plt.plot(cc,slope*cc+intercept,'k-')
plt.title('fire count predictions')
print slope,intercept,R,p,err

