%matplotlib inline
%load_ext autoreload
%autoreload 2
import numpy as np
import matplotlib.pyplot as plt
from pymks import MKSRegressionModel
from pymks import FiPyCHModel
The autoreload extension is already loaded. To reload it, use: %reload_ext autoreload
Make some actual data (400 samples) with a random seed of 101.
np.random.seed(101)
X = np.random.random((400, 21, 21))
fipymodel = FiPyCHModel()
y = fipymodel.predict(X)
The FiPy Cahn-Hilliard model has been packaged into the FiPyCHModel class within the fit, predict paradigm.
fipymodel.fit(None, None)
--------------------------------------------------------------------------- NotImplementedError Traceback (most recent call last) <ipython-input-26-2db047ef34e7> in <module>() ----> 1 fipymodel.fit(None, None) /home/wd15/git/pymks/pymks/fipyCHModel.pyc in fit(self, X, y) 15 16 def fit(self, X, y): ---> 17 raise NotImplementedError 18 19 def predict(self, X): NotImplementedError:
??fipymodel
A sanity check¶
Let's fit the data.
model = MKSRegressionModel(Nbin=10)
model.fit(X, y)
The model now knows its coefficients
model.Fcoeff.shape
(21, 21, 10)
Let's check that the fit sort of "looks right" with one test sample
np.random.seed(102)
X_test = np.random.random((1,) + X.shape[1:])
y_test = fipymodel.predict(X_test)
y_pred = model.predict(X_test)
np.random.seed(2)
index = np.random.randint(len(y_test.flatten()), size=10)
print y_test.flatten()[index]
print y_pred.flatten()[index]
[ 23950253.78282328 13425583.87216846 10370446.72304489 -5082255.45966469 -37232615.87479136 26787885.25872419 24692312.47186198 27598782.37350243 2548349.44343853 9095170.78582771] [ 23924081.38330398 13398981.55267717 10372689.12723125 -5103357.55708055 -37274867.56519653 26809449.81666655 24722045.53760792 27628866.17868118 2557932.70513815 9110175.33127277]
and check the "mean square error" using Sklearn's mean_squared_error function.
from sklearn import metrics
mse = metrics.mean_squared_error
'%1.3e' % np.sqrt(mse(y_test, y_pred))
'5.984e+04'
Seems okay. How do the coefficients look?
coeff = np.fft.ifftn(model.Fcoeff, axes=(0, 1)).real
Nroll = coeff.shape[0] / 2
coeff_rolled = np.roll(np.roll(coeff[:,:,0], Nroll, axis=0), Nroll, axis=1)
plt.contourf(coeff_rolled, 249)
plt.colorbar()
<matplotlib.colorbar.Colorbar instance at 0x4ddf248>
Training and testing¶
Now, let's use the Sklearn function train_test_split to split the data into training and test data sets. If we use the entire set
of data for the fitting and leave nothing for testing, we have no idea if we are simply "overfitting" the data. Think of the analogy of using a high order polynomial
to fit data points on a graph, while it may fit the data perfectly, we learn nothing as it is a useless model for fitting subsequently generated data.
The argument test_size=0.5 splits the data into equal sized chunks for training and testing.
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=3)
print X_train.shape
(200, 21, 21)
Let's refit with the training data set only.
model = MKSRegressionModel(Nbin=10)
model.fit(X_train, y_train)
How well does it predict?
'%1.3e' % mse(model.predict(X_test), y_test)
'4.018e+09'
The above is just one way to split the data. We may want to check with alternative splits of the data. This is easy using Sklearn's cross_validation module. Here we do 10 different splits and check the mean and standard deviation of the mean square error.
from sklearn import cross_validation
model = MKSRegressionModel(Nbin=10)
scores = cross_validation.cross_val_score(model, X, y, score_func=mse, cv=20)
print("MSE: %1.3e (+/- %1.3e)" % (scores.mean(), scores.std()))
MSE: 3.946e+09 (+/- 2.568e+08)
The MKSRegressionModel has methods inherited from the Sklearn's LinearRegressionModel that allows the use of cross_validation.
??MKSRegressionModel.set_params
The LinearRegressionModel may not be the best class to inherit from. Fitting MKS into Sklearn needs careful consideration.
Optimize the Nbin hyperparameter¶
Nbin is known as a hyperparameter. Hyperparameters are parameters that influence the fitting, but are separate from the data and the parameters used to generate the data.
Exercise 04-0¶
Use all the data (X and y) to call fit for various values of Nbin. Calculate the mean square error by comparing the predicted data against the original data, y.
- Do this for values of
Nbinstarting at 2 up to 100 with spacing of 10 usingnp.arange - What does the graph show?
Exercise 04-1¶
Use the training data (X_train, y_train) to fit the MKS model and compare it with the test data. Plot the mean square error versus Nbin. Do this for all values of Nbin between 1 and 20.
What is the optimum value of Nbin?
import matplotlib.pyplot as plt
mse = metrics.mean_squared_error
Nbins = np.arange(2, 100, 10)
errors = []
for Nbin in Nbins:
print Nbin
model = MKSRegressionModel(Nbin=Nbin)
model.fit(X, y)
errors.append(mse(model.predict(X), y))
plt.plot(Nbins, errors)
plt.xlabel('Nbin')
plt.ylabel('MSE')
2 12 22 32 42 52 62 72 82 92
<matplotlib.text.Text at 0x608d550>
errors = []
Nbins = np.arange(2, 20)
for Nbin in Nbins:
model = MKSRegressionModel(Nbin=Nbin)
model.fit(X_train, y_train)
errors.append(mse(model.predict(X_test), y_test))
plt.plot(Nbins, errors)
plt.xlabel('Nbin')
plt.ylabel('MSE')
argmin = np.argmin(errors)
print "optimal Nbin: {0}, mse: {1:1.3e}".format(Nbins[argmin], errors[argmin])
optimal Nbin: 6, mse: 3.946e+09
Training, validation and test data for hyperparameter optimization¶
Using the test data to optimize hyperparameters leads to test data
knowledge leaking into the model. A way to avoid this is to optimize
the hyperparamters with a validation data set separate from the test
data set. Sklearn provides GridSearchCV to automate the optimization
of hyperparamters using only the training data without using the test
data.
from sklearn.grid_search import GridSearchCV
parameters_to_tune = [{'Nbin': np.arange(2, 20)}]
Make a scoring function. Highest score is best.
def neg_mse(a, b):
return -mse(a, b)
Create a GridSearchCV instance.
gridSearch = GridSearchCV(MKSRegressionModel(Nbin=10), parameters_to_tune, cv=5, score_func=neg_mse)
Optimize with only the training data.
??gridSearch.fit
gridSearch.fit(X_train, y_train)
GridSearchCV(cv=5, estimator=MKSRegressionModel(Nbin=10), fit_params={},
iid=True, loss_func=None, n_jobs=1,
param_grid=[{'Nbin': array([ 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18,
19])}],
pre_dispatch='2*n_jobs', refit=True,
score_func=<function neg_mse at 0x5244848>, verbose=0)
Find the best estimator.
??gridSearch.score
print(gridSearch.best_estimator_)
print gridSearch.score(X_test, y_test)
MKSRegressionModel(Nbin=5) 0.999989464884
What were the MSEs for each value of Nbin.
for params, mean_score, scores in gridSearch.grid_scores_:
print("%1.3e (+/-%1.3e) for %r"% (mean_score, scores.std() / 2, params))
-4.149e+09 (+/-8.734e+07) for {'Nbin': 2}
-4.177e+09 (+/-8.521e+07) for {'Nbin': 3}
-3.979e+09 (+/-8.426e+07) for {'Nbin': 4}
-3.970e+09 (+/-8.234e+07) for {'Nbin': 5}
-3.990e+09 (+/-7.744e+07) for {'Nbin': 6}
-4.010e+09 (+/-7.878e+07) for {'Nbin': 7}
-4.032e+09 (+/-7.804e+07) for {'Nbin': 8}
-4.062e+09 (+/-7.806e+07) for {'Nbin': 9}
-4.089e+09 (+/-7.930e+07) for {'Nbin': 10}
-4.117e+09 (+/-7.846e+07) for {'Nbin': 11}
-4.144e+09 (+/-7.870e+07) for {'Nbin': 12}
-4.170e+09 (+/-8.034e+07) for {'Nbin': 13}
-4.199e+09 (+/-7.842e+07) for {'Nbin': 14}
-4.221e+09 (+/-7.922e+07) for {'Nbin': 15}
-4.245e+09 (+/-7.982e+07) for {'Nbin': 16}
-4.281e+09 (+/-8.383e+07) for {'Nbin': 17}
-4.313e+09 (+/-8.172e+07) for {'Nbin': 18}
-4.334e+09 (+/-8.032e+07) for {'Nbin': 19}
How does that match the test data?
y_true, y_pred = y_test, gridSearch.predict(X_test)
'%1.3e' % mse(y_true, y_pred)
'3.947e+09'
What about for other values of Nbin.
for Nbin in parameters_to_tune[0]['Nbin']:
model = MKSRegressionModel(Nbin=Nbin)
model.fit(X_train, y_train)
y_true, y_pred = y_test, model.predict(X_test)
print 'Nbin: {0}, mse: {1:1.3e}'.format(Nbin, mse(y_true, y_pred))
Another example from Sklearn's tutorial¶
The GridSearchCV class can be used to search over multiple hyperparameters. This following example is from Sklearn's turorial. The scoring is very different between a classification problem and a
from sklearn import datasets
digits = datasets.load_digits()
print digits.data.shape
for index, (image, label) in enumerate(zip(digits.images, digits.target)[:4]):
plt.subplot(2, 4, index + 1)
plt.axis('off')
plt.imshow(image, cmap=plt.cm.gray_r, interpolation='nearest')
plt.title('Training: %i' % label)
(1797, 64)
from IPython.display import HTML
HTML('<iframe src=http://scikit-learn.org/stable/auto_examples/grid_search_digits.html#example-grid-search-digits-py width=700 height=700></iframe>')
MKS is really prediciting a quantity. The above example is classification.
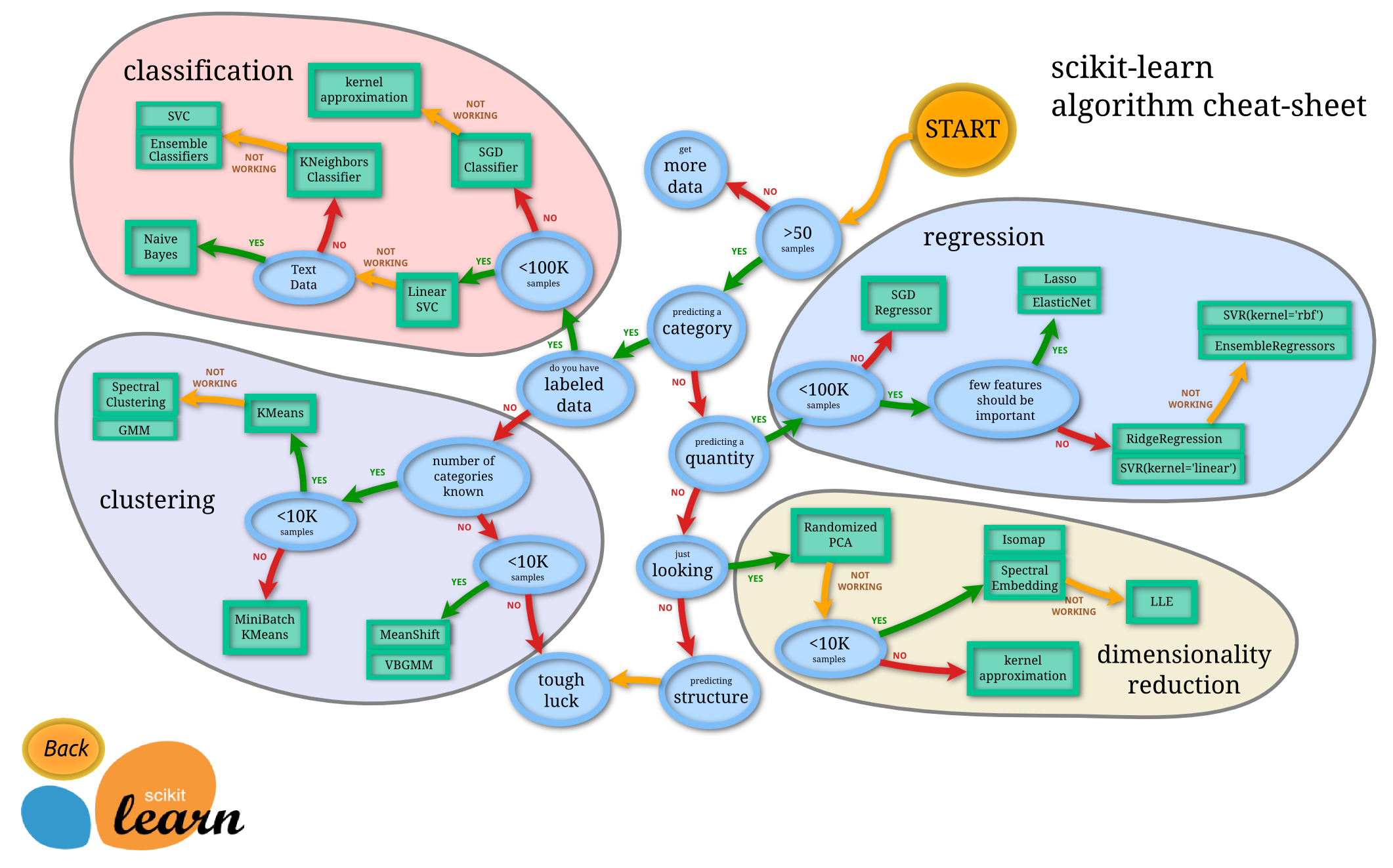
from IPython.display import Image
Image(url='http://scikit-learn.org/stable/_static/ml_map.png', width=900)