import numpy as np
import pandas as pd
import patsy
import seaborn as sns
import statsmodels.formula.api as smf
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
import itertools
%matplotlib inline
sns.set_style("white")
Introduction¶
In this lecture we will cover:
- interaction effects and what they mean
- link functions
In next week's lecture we will look at ANOVA and repeated measures designs.
Interactions¶
An interaction between two variables is when the effect of one variable depends on (is modified by) another variable.
For example,
- the relationship beween drug dosage and disease outcome depends on weight
- the effect of attentional load on reaction time depends on whether the observer is depressed or not
Interactions are ubiquitous in experimental designs in neuroscience, and sometimes they aren't tested appropriately:
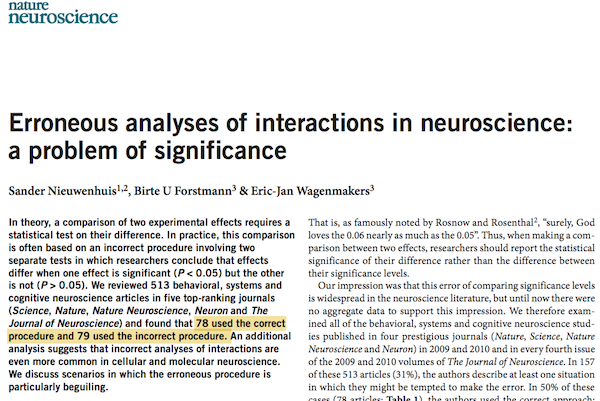

First let's give a simple example of an interaction between two continuous variables.
Interaction example¶
A model with no interaction is a plane in a multidimensional space. Adding an interaction term lets the model prediction bend – that is, the influence of one variable on the response is no longer constant, but depends on the level of the other variable.
We can consider numerous types of interactions that make sense in the real world. The sign of the interaction's coefficient will determine the shape of the bend in the model surface.
One example of a strong interaction (from John Kruschke's book, Doing Bayesian Data Analysis) occurs for the lift (how fast an object rises) of lighter-than-air craft (i.e. balloons). For hot air balloons, lift increases as a function of fire.

For hydrogen balloons, lift increases as a function of hydrogen. But if we combine fire and hydrogen, we get a massive decrease in lift.
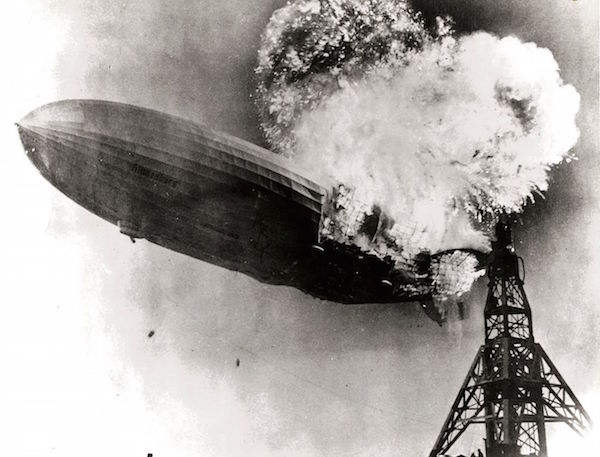

That is, the two interact. We can visualise this as a curved 3D surface:
betas = np.array([1, 1, -2.])
x = np.linspace(0, 1) # vector for x
y = np.linspace(0, 1) # vector for y
X, Y = np.meshgrid(x, y) # grid of X and Y
# the height Z is the sum of the weighted X and Ys (weighted by our betas):
Z = X*betas[0] + \
Y*betas[1] + \
X*Y*betas[2]
# create a matplotlib figure window, add 3D axes
# http://matplotlib.org/examples/mplot3d/subplot3d_demo.html
fig = plt.figure(figsize=(12, 4))
# fig = plt.figure(figsize=plt.figaspect(0.3))
# first view:
ax = fig.add_subplot(1, 3, 1, projection='3d')
ax.plot_surface(X, Y, Z, alpha=1, cmap=cm.coolwarm)
ax.view_init(elev=15, azim=20)
ax.set_xlabel('Fire')
ax.set_xticks([0, .5, 1])
ax.set_ylabel('Hydrogen')
ax.set_yticks([0, .5, 1])
ax.set_zlabel('Lift')
ax.set_zticks([])
# second view:
ax = fig.add_subplot(1, 3, 2, projection='3d')
ax.plot_surface(X, Y, Z, alpha=1, cmap=cm.coolwarm)
ax.view_init(elev=20, azim=55)
ax.set_xlabel('Fire')
ax.set_xticks([0, .5, 1])
ax.set_ylabel('Hydrogen')
ax.set_yticks([0, .5, 1])
ax.set_zlabel('Lift')
ax.set_zticks([])
# third view:
ax = fig.add_subplot(1, 3, 3, projection='3d')
ax.plot_surface(X, Y, Z, alpha=1, cmap=cm.coolwarm)
ax.view_init(elev=10, azim=80)
ax.set_xlabel('Fire')
ax.set_xticks([0, .5, 1])
ax.set_ylabel('Hydrogen')
ax.set_yticks([0, .5, 1])
ax.set_zlabel('Lift')
ax.set_zticks([])
plt.show()
While 3D plots can be nifty, usually it's nicer to visualise this in 2D as a coloured plane (equivalent to looking down on the surface above). Seaborn can fit interaction models between two continuous covariates in a single step, and plot:
# make a z vector
z = x*betas[0] + \
y*betas[1] + \
x*y*betas[2]
# put all the vectors into a new dataframe:
df = pd.DataFrame({'fire': x,
'hydrogen': y,
'lift': z})
# fit regression with interaction, plot:
sns.interactplot('fire', 'hydrogen', 'lift', df,
cmap="coolwarm", filled=True, levels=25);
Interactions in the design matrix: two continuous variables example¶
How are interaction terms implemented in a design matrix? It's very simple: they are the product of single-variable design matrix columns. The interaction columns are then weighted by a coefficient, as for any other variable. Let's look at how this works, and what it means.
# import the dataset csv file into a Pandas dataframe object:
sleep = pd.read_csv('../Datasets/sleep.txt', sep='\t')
sleep = sleep.dropna(axis=0) # drop on axis 0 (i.e. rows)
# variable creation:
sleep['log_BodyWt'] = np.log(sleep['BodyWt'])
sleep['danger_cat'] = 'low' # set as a string, "low" for all.
sleep.loc[sleep['Danger'] > 3, 'danger_cat'] = 'high' # set values of 4 and 5 to "high"
sleep['life_cat'] = pd.qcut(sleep['LifeSpan'], [0, .33, .66, 1])
sleep['life_cat'] = sleep['life_cat'].cat.rename_categories(['short', 'med', 'long']) # rename categories
sleep['life_cat'] = sleep['life_cat'].astype('object') # make it an object, rather than a category
sleep.info()
<class 'pandas.core.frame.DataFrame'> Int64Index: 42 entries, 1 to 60 Data columns (total 14 columns): Species 42 non-null object BodyWt 42 non-null float64 BrainWt 42 non-null float64 NonDreaming 42 non-null float64 Dreaming 42 non-null float64 TotalSleep 42 non-null float64 LifeSpan 42 non-null float64 Gestation 42 non-null float64 Predation 42 non-null int64 Exposure 42 non-null int64 Danger 42 non-null int64 log_BodyWt 42 non-null float64 danger_cat 42 non-null object life_cat 42 non-null object dtypes: float64(8), int64(3), object(3) memory usage: 4.9+ KB
An interaction hypothesis for sleep¶
Let's imagine that we want to know if the relationship between bodyweight and total sleep changes depending on the length of the animal's gestation.
In patsy formulae, we can include an interaction term using the ":" character. So BodyWt:Gestation is an interaction between those two variables. A shorthand for including both main effects (the variables alone) and their interaction is the "*" character. So writing BodyWt * Gestation in the formula is the same as writing BodyWt + Gestation + BodyWt:Gestation.
X = patsy.dmatrix('log_BodyWt * Gestation', sleep)
X
DesignMatrix with shape (42, 4)
Intercept log_BodyWt Gestation log_BodyWt:Gestation
1 0.00000 42.0 0.00000
1 7.84267 624.0 4893.82700
1 2.35613 180.0 424.10265
1 -3.77226 35.0 -132.02914
1 5.07517 392.0 1989.46814
1 1.19392 63.0 75.21712
1 3.95432 230.0 909.49266
1 -0.85567 112.0 -95.83460
1 6.14204 281.0 1725.91251
1 -2.59027 42.0 -108.79122
1 1.09861 28.0 30.76114
1 -0.24207 42.0 -10.16701
1 -1.60944 120.0 -193.13255
1 3.31999 148.0 491.35812
1 -2.12026 16.0 -33.92422
1 4.44265 310.0 1377.22189
1 -2.29263 28.0 -64.19377
1 0.03922 68.0 2.66701
1 6.25575 336.0 2101.93201
1 -5.29832 21.5 -113.91382
1 -4.60517 50.0 -230.25851
1 4.12713 267.0 1101.94488
1 -3.77226 19.0 -71.67296
1 -3.03655 30.0 -91.09663
1 0.53063 12.0 6.36754
1 1.25276 120.0 150.33156
1 -0.73397 140.0 -102.75568
1 2.30259 170.0 391.43947
1 0.48243 17.0 8.20124
1 5.25750 115.0 604.61197
[12 rows omitted]
Terms:
'Intercept' (column 0)
'log_BodyWt' (column 1)
'Gestation' (column 2)
'log_BodyWt:Gestation' (column 3)
(to view full data, use np.asarray(this_obj))
So you can see we get a new column that's the product of the other two. Let's think about what this column means.
First, let's look at the relationship between these variables by using a pairplot (also called a scatterplot matrix).
# convert to pandas df so seaborn is happy:
tmp = pd.DataFrame(X, columns = ['intercept', 'log_bodyweight', 'gestation', 'interaction'] )
# plot pairplot:
sns.pairplot(tmp, vars=['log_bodyweight', 'gestation', 'interaction']);
Incidentally, note how gestation is also skewed (and consequently the interaction term, too). This would be a good candidate to log-scale as well, but for simplicity we'll leave it be.
The interaction term means that we're adding some value ($\beta_4 \times \mathrm{bodyweight} \times \mathrm{gestation}$) into the linear predictor. When both bodyweight and gestation are large, this value is large.
What happens when we fit this model?
fit_2 = smf.glm('TotalSleep ~ log_BodyWt * Gestation', sleep).fit()
print(fit_2.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: TotalSleep No. Observations: 42
Model: GLM Df Residuals: 38
Model Family: Gaussian Df Model: 3
Link Function: identity Scale: 13.3387035372
Method: IRLS Log-Likelihood: -111.90
Date: Tue, 24 Feb 2015 Deviance: 506.87
Time: 17:11:27 Pearson chi2: 507.
No. Iterations: 4
========================================================================================
coef std err z P>|z| [95.0% Conf. Int.]
----------------------------------------------------------------------------------------
Intercept 13.1770 1.056 12.474 0.000 11.107 15.248
log_BodyWt -0.6223 0.298 -2.086 0.037 -1.207 -0.038
Gestation -0.0196 0.010 -1.873 0.061 -0.040 0.001
log_BodyWt:Gestation 0.0013 0.001 0.876 0.381 -0.002 0.004
========================================================================================
The interaction term is not significant (its confidence interval borders zero), so we could probably exclude it from the model. For demonstration, let's look at the model surface.
x = np.linspace(sleep['log_BodyWt'].min(), sleep['log_BodyWt'].max()) # vector for x (log bodyweight)
y = np.linspace(sleep['Gestation'].min(), sleep['Gestation'].max()) # vector for y (gestation)
X, Y = np.meshgrid(x, y) # grid of X and Y
# the height Z is the sum of the weighted X and Ys (weighted by our betas):
Z = fit_2.params[0] + \
X*fit_2.params[1] + \
Y*fit_2.params[2] + \
X*Y*fit_2.params[3] # note the addition of the interaction term here!
# create a matplotlib figure window, add 3D axes
# http://matplotlib.org/examples/mplot3d/subplot3d_demo.html
fig = plt.figure(figsize=(12, 4))
# first view:
ax = fig.add_subplot(1, 3, 1, projection='3d')
ax.plot_surface(X, Y, Z, cmap=cm.coolwarm)
ax.view_init(elev=10, azim=20)
ax.set_xlabel('log bodyweight (kg)')
ax.set_ylabel('gestation (days)')
ax.set_zlabel('Total sleep (hours)')
# second view:
ax = fig.add_subplot(1, 3, 2, projection='3d')
ax.plot_surface(X, Y, Z, cmap=cm.coolwarm)
ax.view_init(elev=10, azim=45)
ax.set_xlabel('log bodyweight (kg)')
ax.set_ylabel('gestation (days)')
ax.set_zlabel('Total sleep (hours)')
# third view:
ax = fig.add_subplot(1, 3, 3, projection='3d')
ax.plot_surface(X, Y, Z, cmap=cm.coolwarm)
ax.view_init(elev=10, azim=70)
ax.set_xlabel('log bodyweight (kg)')
ax.set_ylabel('gestation (days)')
ax.set_zlabel('Total sleep (hours)')
plt.show()
sns.interactplot("log_BodyWt", "Gestation", "TotalSleep", sleep,
cmap="coolwarm", filled=True, levels=25);
Interactions between categorical variables¶
We saw in the last lecture how categorical variables (e.g. gender, religion) can be included into design matrices, using dummy coding. How are interaction terms calculated for dummy coded variables?
It's exactly the same: the product of the relevant design matrix columns. This allows the effect of one variable to depend on another. Let's look at an example in the context of a $2 \times 2$ design.
Paper planes experiment¶
How far does a paper plane fly, depending on its design and the weight of the paper used?
Some undergraduate statistics students from Australia wanted to find out. So they ran a study in which they threw paper planes down a hallway in a university building.
We're going to use this dataset to demonstrate an interaction in a $2 \times 2$ factorial experiment. Note that for the purposes of this demo, we will ignore some of the variables in the dataset.
# read in the data, rename some values:
planes = pd.read_csv('../Datasets/planes.txt', sep='\t')
# column names to lowercase:
planes.columns = [c.lower() for c in planes.columns]
# turn "paper" variable into an object, not int64:
planes['paper'] = planes['paper'].astype(object)
planes.replace(to_replace={'paper': {1: '80',
2: '50'},
'angle': {1: 'horz',
2: '45deg '},
'plane': {1: 'sleek',
2: 'simple'}},
inplace=True)
# rename "plane" to "design":
planes = planes.rename(columns={'plane': 'design'})
print(planes)
distance paper angle design order 0 2160 80 horz sleek 12 1 1511 80 horz sleek 11 2 4596 80 horz simple 8 3 3706 80 horz simple 6 4 3854 80 45deg sleek 4 5 1690 80 45deg sleek 2 6 5088 80 45deg simple 1 7 4255 80 45deg simple 7 8 6520 50 horz sleek 3 9 4091 50 horz sleek 9 10 2130 50 horz simple 14 11 3150 50 horz simple 5 12 6348 50 45deg sleek 16 13 4550 50 45deg sleek 15 14 2730 50 45deg simple 13 15 2585 50 45deg simple 10
fit = smf.glm('distance ~ design * paper', planes).fit()
print(fit.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: distance No. Observations: 16
Model: GLM Df Residuals: 12
Model Family: Gaussian Df Model: 3
Link Function: identity Scale: 796752.416667
Method: IRLS Log-Likelihood: -129.11
Date: Tue, 24 Feb 2015 Deviance: 9.5610e+06
Time: 17:11:29 Pearson chi2: 9.56e+06
No. Iterations: 4
===============================================================================================
coef std err z P>|z| [95.0% Conf. Int.]
-----------------------------------------------------------------------------------------------
Intercept 2648.7500 446.305 5.935 0.000 1774.008 3523.492
design[T.sleek] 2728.5000 631.171 4.323 0.000 1491.429 3965.571
paper[T.80] 1762.5000 631.171 2.792 0.005 525.429 2999.571
design[T.sleek]:paper[T.80] -4836.0000 892.610 -5.418 0.000 -6585.483 -3086.517
===============================================================================================
In this model, there are significant effects of plane design and paper weight, and also a significant interaction term. What do these mean?
First, let's look at the design matrix.
X = patsy.dmatrix('~ design * paper', planes)
X
DesignMatrix with shape (16, 4)
Intercept design[T.sleek] paper[T.80] design[T.sleek]:paper[T.80]
1 1 1 1
1 1 1 1
1 0 1 0
1 0 1 0
1 1 1 1
1 1 1 1
1 0 1 0
1 0 1 0
1 1 0 0
1 1 0 0
1 0 0 0
1 0 0 0
1 1 0 0
1 1 0 0
1 0 0 0
1 0 0 0
Terms:
'Intercept' (column 0)
'design' (column 1)
'paper' (column 2)
'design:paper' (column 3)
We have our intercept column of ones as before. The second and third columns are the "switches" for whether the plane is "sleek", and whether a paper weight of 80 grams was used.
So the intercept $\beta_1$ weight corresponds to the distance travelled when design == simple and paper == 50. $\beta_2$ is the difference between the distance travelled for sleek and simple planes, when the paper weight is 50. $\beta_3$ is the difference in distance travelled between paper of weight 50 and paper of 80 grams, for simple planes.
What about the interaction column? We can see that again, this is just the product of the two simple effect columns. When both design[T.sleek] and paper[T.80] are 1, the interaction is 1. Otherwise, it's zero.
Let's think about what this means.
If we throw a sleek-designed plane, made of 80 gram paper, then we add the interaction coefficient $\beta_4$ to the prediction. Otherwise, no coefficient is added.
Let's visualise the data and the model predictions, using Seaborn. To do this, we need to merge some data frames together so that we can plot the predictions with the data. The reason for this is that Seaborn plotting commands accept one and only one data frame. There are probably other ways to make this plot; I got this way from the answer to my question here.
# make a data frame containing the predictions, using the expand grid function.
# define expand grid function:
def expand_grid(data_dict):
""" A port of R's expand.grid function for use with Pandas dataframes.
from http://pandas.pydata.org/pandas-docs/stable/cookbook.html?highlight=expand%20grid
"""
rows = itertools.product(*data_dict.values())
return pd.DataFrame.from_records(rows, columns=data_dict.keys())
# build a new matrix with expand grid:
preds = expand_grid({'paper': ['80', '50'],
'design': ['sleek', 'simple']})
preds_2 = preds.copy()
preds['distance'] = fit.predict(preds)
# also fit a model with no interaction for comparison:
fit_2 = smf.glm('distance ~ paper + design', planes).fit()
preds_2['distance'] = fit_2.predict(preds_2)
# to plot model predictions against data, we should merge them together:
# see answer (https://stackoverflow.com/questions/28239359/showing-data-and-model-predictions-in-one-plot-using-seaborn-and-statsmodels)
merged = pd.concat(dict(data=planes,
additive=preds_2,
interaction=preds),
names=['type']).reset_index()
print(merged)
type level_1 angle design distance order paper 0 additive 0 NaN sleek 3512.75 NaN 80 1 additive 1 NaN sleek 4168.25 NaN 50 2 additive 2 NaN simple 3202.25 NaN 80 3 additive 3 NaN simple 3857.75 NaN 50 4 data 0 horz sleek 2160.00 12 80 5 data 1 horz sleek 1511.00 11 80 6 data 2 horz simple 4596.00 8 80 7 data 3 horz simple 3706.00 6 80 8 data 4 45deg sleek 3854.00 4 80 9 data 5 45deg sleek 1690.00 2 80 10 data 6 45deg simple 5088.00 1 80 11 data 7 45deg simple 4255.00 7 80 12 data 8 horz sleek 6520.00 3 50 13 data 9 horz sleek 4091.00 9 50 14 data 10 horz simple 2130.00 14 50 15 data 11 horz simple 3150.00 5 50 16 data 12 45deg sleek 6348.00 16 50 17 data 13 45deg sleek 4550.00 15 50 18 data 14 45deg simple 2730.00 13 50 19 data 15 45deg simple 2585.00 10 50 20 interaction 0 NaN sleek 2303.75 NaN 80 21 interaction 1 NaN sleek 5377.25 NaN 50 22 interaction 2 NaN simple 4411.25 NaN 80 23 interaction 3 NaN simple 2648.75 NaN 50
# plot with seaborn:
g = sns.factorplot('design', 'distance', 'paper',
data=merged,
col='type',
kind='point',
size=3.5)
g.fig.subplots_adjust(wspace=0.3)
sns.despine(offset=10);
print(fit_2.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: distance No. Observations: 16
Model: GLM Df Residuals: 13
Model Family: Gaussian Df Model: 2
Link Function: identity Scale: 2534455.76923
Method: IRLS Log-Likelihood: -139.01
Date: Tue, 24 Feb 2015 Deviance: 3.2948e+07
Time: 17:11:30 Pearson chi2: 3.29e+07
No. Iterations: 4
===================================================================================
coef std err z P>|z| [95.0% Conf. Int.]
-----------------------------------------------------------------------------------
Intercept 3857.7500 689.355 5.596 0.000 2506.639 5208.861
paper[T.80] -655.5000 795.999 -0.823 0.410 -2215.629 904.629
design[T.sleek] 310.5000 795.999 0.390 0.696 -1249.629 1870.629
===================================================================================
The three panels above show the data (with bootstrapped 95% confidence intervals around the means), the predictions of the additive model and the predictions of the interaction model.
By comparing the above plots (predictions with and without an interaction), it becomes clear that the effect of paper depends strongly on the plane design. That is, the relationship between design and distance depends on the paper weight.
This is also a good example of when it's not very meaningful to talk about a significant main effect if an interaction involving that factor is significant.
For example, in the interaction model, there is a signficant simple effect of design such that the sleek design travels further on average than the simple design (for a paper weight of 50). However, the interaction means that the sleek design with heavy paper actually travels a shorter distance than the simple design (whether the simple plane is made of heavy or light paper). So writing something like "the sleek design travels further than the simple design, since we had a significant effect of design" is misleading.
Interaction between categorical and continuous variables¶
What happens when we include an interaction term between a categorical and a continuous variable? In the design matrix, it's exactly the same operation: we take the products of the relevant columns.
To demonstrate what this means, let's return to the sleep dataset, and consider an interaction between bodyweight and our categorical danger variable.
X = patsy.dmatrix('~ log_BodyWt * danger_cat', sleep)
X
DesignMatrix with shape (42, 4)
Intercept danger_cat[T.low] log_BodyWt log_BodyWt:danger_cat[T.low]
1 1 0.00000 0.00000
1 0 7.84267 0.00000
1 0 2.35613 0.00000
1 1 -3.77226 -3.77226
1 0 5.07517 0.00000
1 1 1.19392 1.19392
1 1 3.95432 3.95432
1 0 -0.85567 -0.00000
1 0 6.14204 0.00000
1 1 -2.59027 -2.59027
1 1 1.09861 1.09861
1 1 -0.24207 -0.24207
1 1 -1.60944 -1.60944
1 0 3.31999 0.00000
1 1 -2.12026 -2.12026
1 1 4.44265 4.44265
1 1 -2.29263 -2.29263
1 0 0.03922 0.00000
1 0 6.25575 0.00000
1 0 -5.29832 -0.00000
1 1 -4.60517 -4.60517
1 1 4.12713 4.12713
1 1 -3.77226 -3.77226
1 1 -3.03655 -3.03655
1 1 0.53063 0.53063
1 1 1.25276 1.25276
1 1 -0.73397 -0.73397
1 0 2.30259 0.00000
1 1 0.48243 0.48243
1 0 5.25750 0.00000
[12 rows omitted]
Terms:
'Intercept' (column 0)
'danger_cat' (column 1)
'log_BodyWt' (column 2)
'log_BodyWt:danger_cat' (column 3)
(to view full data, use np.asarray(this_obj))
This model can be written as:
$\hat y = \beta_1 + \beta_2 \mathrm{danger} + \beta_3 \mathrm{bodyweight} + \beta_4 \mathrm{interaction}$
So when danger is "high", our model reduces to the intercept plus the log_BodyWeight:
$\hat y = \beta_1 + \beta_3 \mathrm{bodyweight}$
Whereas when danger is "low", we
- add $\beta_2$ to the predictions, irrespective of bodyweight (i.e. a change in intercept)
- change the slope of bodyweight by $\beta_4$
So what our interaction term effectively does is allows the danger = low condition to have a different intercept and a different slope than the danger = high condition.
What happens when we fit this model?
fit_2 = smf.glm('TotalSleep ~ log_BodyWt * danger_cat', sleep).fit() # interaction model
print(fit_2.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: TotalSleep No. Observations: 42
Model: GLM Df Residuals: 38
Model Family: Gaussian Df Model: 3
Link Function: identity Scale: 13.1208381942
Method: IRLS Log-Likelihood: -111.55
Date: Tue, 24 Feb 2015 Deviance: 498.59
Time: 17:11:30 Pearson chi2: 499.
No. Iterations: 4
================================================================================================
coef std err z P>|z| [95.0% Conf. Int.]
------------------------------------------------------------------------------------------------
Intercept 8.9987 1.262 7.132 0.000 6.526 11.472
danger_cat[T.low] 3.1610 1.436 2.201 0.028 0.346 5.976
log_BodyWt -0.6155 0.292 -2.108 0.035 -1.188 -0.043
log_BodyWt:danger_cat[T.low] -0.2637 0.415 -0.636 0.525 -1.077 0.549
================================================================================================
We can see that the z-test for the coefficient of interaction term in the table is not significant. This means that from these data, we can't conclude that the relationship between bodyweight and sleep depends on danger. That is, the slopes of the line for the low and high danger groups do not significantly differ.
# Plot the two models against each other.
# we do this slightly differently to the categorical examples above,
# due to the need for some data re-arranging to plot the data as points
# and the model predictions as lines.
preds = expand_grid(
{'log_BodyWt': np.linspace(-6, 9, 2),
'life_cat': ['short', 'med', 'long'],
'danger_cat': ['low', 'high']})
fit = smf.glm('TotalSleep ~ log_BodyWt + danger_cat', sleep).fit() # additive model
fit_2 = smf.glm('TotalSleep ~ log_BodyWt * danger_cat', sleep).fit() # interaction model
# add model predictions to data frames:
preds['yhat'] = fit.predict(preds)
preds['yhat_2'] = fit_2.predict(preds)
merged = sleep.append(preds)
# Plot additive model:
g = sns.FacetGrid(merged, hue='danger_cat', size=4)
# use the `map` method to add stuff to the facetgrid axes:
g.map(plt.scatter, "log_BodyWt", "TotalSleep")
g.map(plt.plot, "log_BodyWt", "yhat")
sns.despine(offset=10)
g.set_axis_labels('Bodyweight [log kg]', 'Sleep [hours]')
g.add_legend(title='Danger')
# plot interaction model:
g = sns.FacetGrid(merged, hue='danger_cat', size=4)
# use the `map` method to add stuff to the facetgrid axes:
g.map(plt.scatter, "log_BodyWt", "TotalSleep")
g.map(plt.plot, "log_BodyWt", "yhat_2")
sns.despine(offset=10)
g.set_axis_labels('Bodyweight [log kg]', 'Sleep [hours]')
g.add_legend(title='Danger')
Conclusion¶
Interactions let you model how the value of one variable changes the relationship between other variables. That is, we ask "does the relationship between X and Y depend on Z"?
In terms of the design matrix, interaction terms are simply the products of the relative columns.
from IPython.core.display import HTML
def css_styling():
styles = open("../custom_style.css", "r").read()
return HTML(styles)
css_styling()