Workflows for processing raw RNAseq file for integration into qDOD platform¶
Solid Data¶
Step 1: Import into Solid data (csfasta and qual) into CLC
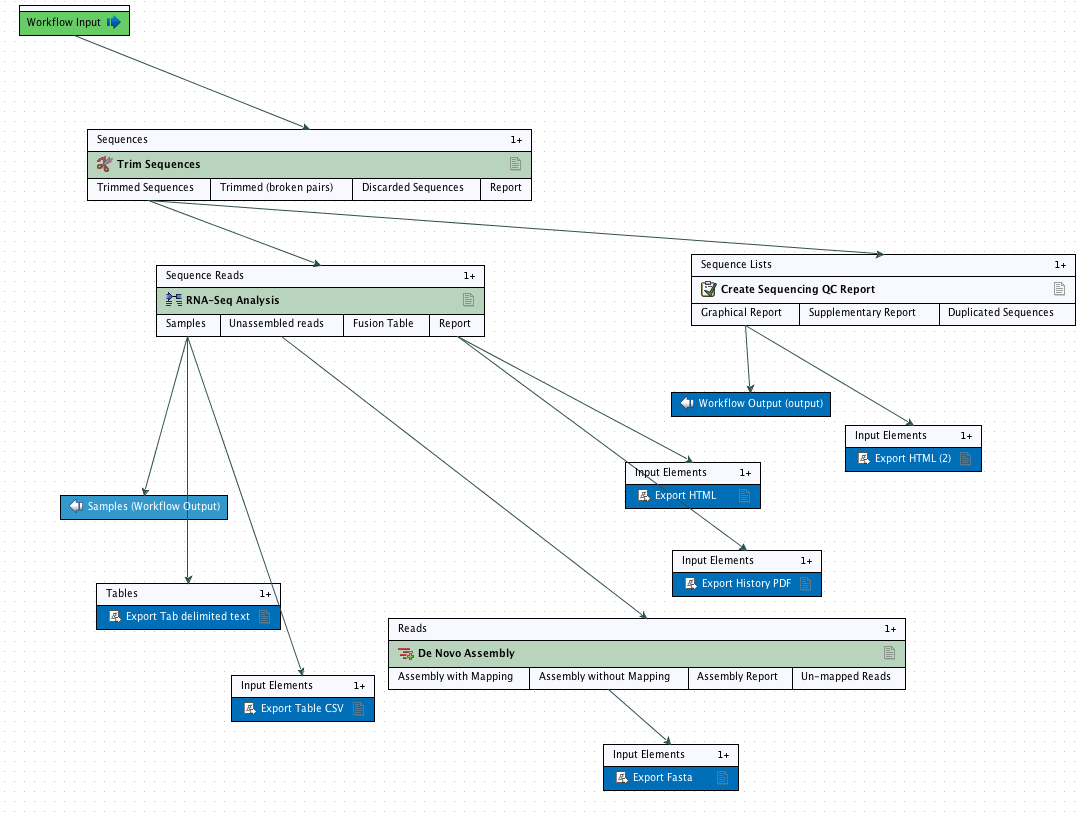
In [22]:
ls /Volumes/web/cnidarian/qdod2
RNA-Seq_DH2.csv RNA-Seq_DH2.csv.sqlshare.restart RNA-Seq_DH2.sqlshare.restart solid0078_20091105_BB3 trimmed RNA-Seq (Workflow Output).csv solid0078_20091105_BB3 trimmed RNA-Seq (Workflow Output).txt solid0078_20091105_BB3 trimmed report.pdf solid0078_20091105_BB3 trimmed un-mapped reads [no read group] (single) contig list.fa solid0078_20091105_DH2 trimmed RNA-Seq (Workflow Output).csv solid0078_20091105_DH2 trimmed RNA-Seq (Workflow Output).txt solid0078_20091105_DH2 trimmed report.pdf solid0078_20091105_DH2 trimmed un-mapped reads [no read group] (single) contig list.fa solid0078_20091105_DH3 trimmed RNA-Seq (Workflow Output).csv solid0078_20091105_DH3 trimmed RNA-Seq (Workflow Output).txt solid0078_20091105_DH3 trimmed report.pdf solid0078_20091105_DH3 trimmed un-mapped reads [no read group] (single) contig list.fa
In [2]:
#solid0078_20091105_DH2
!head /Volumes/web/cnidarian/qdod2/solid0078_20091105_DH2\ trimmed\ RNA-Seq\ \(Workflow\ Output\).csv
"Feature ID","Expression values","Transcripts annotated","Detected transcripts","Exon length","Unique gene reads","Total gene reads","Unique exon reads","Total exon reads","Ratio of unique to total (exon reads)","Unique exon-exon reads","Total exon-exon reads","Unique intron-exon reads","Total intron-exon reads","Exons","Putative exons","RPKM","Median coverage","Chromosome","Chromosome region start","Chromosome region end" "CGI_10000780","0.148","2","1","1350","1","2","1","2","0.5","0","0","0","0","2","0","0.148","0","scaffold350","1105","3206" "CGI_10000456","12.346","4","4","438","44","61","41","54","0.759","0","0","2","6","4","0","12.346","2","scaffold28720","349","2038" "CGI_10000457","0","7","0","603","0","0","0","0","?","0","0","0","0","7","0","0","0","scaffold28720","3692","5692" "CGI_10000774","31.244","2","2","375","117","117","117","117","1","0","0","1","1","2","0","31.244","9","scaffold31684","6549","7563" "CGI_10000917","0","1","0","426","0","0","0","0","?","0","0","0","0","1","0","0","0","scaffold32586","1736","2161" "CGI_10000861","0.6","1","1","2004","12","12","12","12","1","0","0","0","0","1","0","0.6","0","scaffold900","6516","8519" "CGI_10000994","27.133","11","11","1635","389","477","361","443","0.815","0","0","31","35","11","0","27.133","7","scaffold33132","5089","12821" "CGI_10000643","0.181","4","1","552","2","2","1","1","1","0","0","0","0","4","0","0.181","0","scaffold740","109","2125" "CGI_10000763","0.804","3","2","249","4","4","2","2","1","0","0","0","0","3","0","0.804","0","scaffold31534","1194","3724"
In [10]:
#copy and rename RNA-seq_DH2
cp /Volumes/web/cnidarian/qdod2/solid0078_20091105_DH2\ trimmed\ RNA-Seq\ \(Workflow\ Output\).csv /Volumes/web/cnidarian/qdod2/RNA-Seq_DH2
SQLShare Upload Pipeline¶
In [10]:
#code to upload file to SQLShare
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d RNA-Seq_BB3 /Volumes/web/cnidarian/qdod2/solid0078_20091105_BB3\ trimmed\ RNA-Seq\ \(Workflow\ Output\).csv
processing chunk line 0 to 28028 (0.138172864914 s elapsed) pushing /Volumes/web/cnidarian/qdod2/solid0078_20091105_BB3 trimmed RNA-Seq (Workflow Output).csv... parsing B4AB743A... finished RNA-Seq_BB3
In [11]:
#code to set tags
!python /Users/sr320/sqlshare-pythonclient/tools/settags.py RNA-Seq_BB3 "qdod2 RNAseq gill"
send: 'GET /REST.svc/v1/user/sr320@washington.edu HTTP/1.1\r\nHost: rest.sqlshare.escience.washington.edu\r\nAccept-Encoding: identity\r\nAuthorization: ss_apikey sr320@washington.edu : c8c52036bdecbbc798d3248bcc15c15c\r\n\r\n'
reply: 'HTTP/1.1 200 OK\r\n'
send: 'PUT /REST.svc/v2/db/dataset/sr320%40washington.edu/RNA-Seq_BB3/tags HTTP/1.1\r\nHost: rest.sqlshare.escience.washington.edu\r\nAccept-Encoding: identity\r\nContent-Length: 71\r\nContent-Type: application/json\r\nAuthorization: ss_apikey sr320@washington.edu : c8c52036bdecbbc798d3248bcc15c15c\r\nAccept: application/json\r\n\r\n[{"name": "sr320@washington.edu", "tags": ["qdod2", "RNAseq", "gill"]}]'
reply: 'HTTP/1.1 200 OK\r\n'
tags ['qdod2', 'RNAseq', 'gill'] added to dataset RNA-Seq_BB3
In [8]:
!python /Users/sr320/sqlshare-pythonclient/tools/permissions.py -h
Usage: permissions.py [options] (add|remove|print) <user1> <user2> ... <userN>
Options:
-h, --help show this help message and exit
-u USERNAME, --user=USERNAME
SQLshare user name
-p PASSWORD, --password=PASSWORD
SQLshare password
-t TABLES, --table=TABLES
SQLshare dataset