Dheilly et al : Resources associated with Gametogenesis in the Pacific Oyster Crassostrea gigas: A Microarrays-Based Analysis Identifies Sex and Stage Specific Genes¶
quick links
Array Design¶
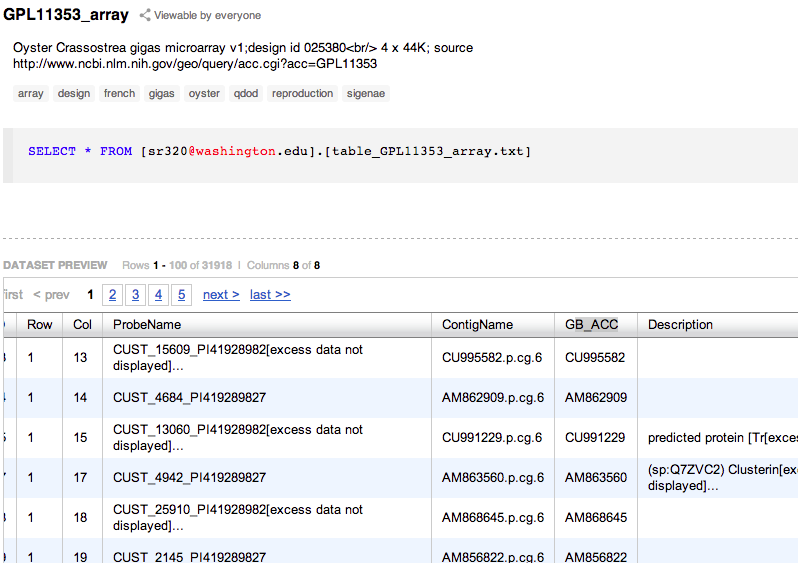
Sigenae Version 6¶
In [3]:
!wget http://dl.dropboxusercontent.com/u/115356/docs/annotable/Roberts_Sigenae6_transcriptome.fa
--2014-04-04 12:05:10-- http://dl.dropboxusercontent.com/u/115356/docs/annotable/Roberts_Sigenae6_transcriptome.fa Resolving dl.dropboxusercontent.com... 54.225.207.37, 54.243.164.243, 54.225.246.188, ... Connecting to dl.dropboxusercontent.com|54.225.207.37|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 25210642 (24M) [text/plain] Saving to: `Roberts_Sigenae6_transcriptome.fa' 100%[======================================>] 25,210,642 887K/s in 28s 2014-04-04 12:05:38 (890 KB/s) - `Roberts_Sigenae6_transcriptome.fa' saved [25210642/25210642]
In [4]:
#tab delimited version
!perl -e '$count=0; $len=0; while(<>) {s/\r?\n//; s/\t/ /g; if (s/^>//) { if ($. != 1) {print "\n"} s/ |$/\t/; $count++; $_ .= "\t";} else {s/ //g; $len += length($_)} print $_;} print "\n"; warn "\nConverted $count FASTA records in $. lines to tabular format\nTotal sequence length: $len\n\n";' Roberts_Sigenae6_transcriptome.fa > Roberts_Sigenae6_transcriptome.tab
Converted 31952 FASTA records in 451733 lines to tabular format Total sequence length: 24247683
In [1]:
!awk -F "," '{print ">"$1"\n"$2}' /Volumes/web/cnidarian/GPL11353_v6fasta.csv > /Volumes/web/cnidarian/GPL11353_v6fasta.fa
In [3]:
!tail /Volumes/web/cnidarian/GPL11353_v6fasta.fa
>AM860865.p.cg.6 >AM869341.p.cg.6 >EE677551.p.cg.6 >AM860135.p.cg.6 >BQ426856.p.cg.6
In [4]:
!head /Volumes/web/cnidarian/BiGoRNA_array_v6_tab
"Name" "Chromosome" "Region" "Expression value" "Gene length" "RPKM" "Unique gene reads" "Total gene reads" CU995582.p.cg.6 CU995582.p.cg.6 1..1092 4 1092 0.236 4 4 AM862909.p.cg.6 AM862909.p.cg.6 1..595 0 595 0 0 0 CU991229.p.cg.6 CU991229.p.cg.6 1..1230 148 1230 7.767 146 148 AM863560.p.cg.6 AM863560.p.cg.6 1..562 101 562 11.601 101 101 AM868645.p.cg.6 AM868645.p.cg.6 1..462 2 462 0.279 2 2 AM856822.p.cg.6 AM856822.p.cg.6 1..846 0 846 0 0 0 FP006184.p.cg.6 FP006184.p.cg.6 1..287 16 287 3.599 16 16 AM862004.p.cg.6 AM862004.p.cg.6 1..808 132 808 10.545 132 132 CU998134.p.cg.6 CU998134.p.cg.6 1..1001 772 1001 49.783 772 772
In [6]:
!head /Volumes/web/cnidarian/BiGoRNA_array_v6_tab.txt
ID Sig_No Gene length RPKM Unique gene reads Total gene reads CU995582 CU995582.p.cg.6 1092 0.236 4 4 AM862909 AM862909.p.cg.6 595 0 0 0 CU991229 CU991229.p.cg.6 1230 7.767 146 148 AM863560 AM863560.p.cg.6 562 11.601 101 101 AM868645 AM868645.p.cg.6 462 0.279 2 2 AM856822 AM856822.p.cg.6 846 0 0 0 FP006184 FP006184.p.cg.6 287 3.599 16 16 AM862004 AM862004.p.cg.6 808 10.545 132 132 CU998134 CU998134.p.cg.6 1001 49.783 772 772
In [8]:
spd="/Users/sr320/sqlshare-pythonclient/tools/"
In [9]:
#uploading to SQLSHARE
!python {spd}singleupload.py -d BiGoRNA_array_v6 /Volumes/web/cnidarian/BiGoRNA_array_v6_tab.txt
processing chunk line 0 to 31918 (1.10805797577 s elapsed) pushing /Volumes/web/cnidarian/BiGoRNA_array_v6_tab.txt... parsing F8132D65... finished BiGoRNA_array_v6
In [ ]:
!python {spd}fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[BiGoRNA_array_v6]rna left join [sr320@washington.edu].[GPL11353_array]arr on rna.Sig_No=arr.ContigName" -f tsv -o /Volumes/web/cnidarian/BiRNAseq_GPL11353_v6ref.txt
In [11]:
!head /Volumes/web/cnidarian/BiRNAseq_GPL11353_v6ref.txt
In [23]:
!python /Applications/q "SELECT * FROM /Volumes/web/cnidarian/BiRNAseq_GPL11353_v6ref.txt WHERE c1 like CU995582"
query error: no such column: CU995582
In [29]:
!grep "CU995582" /Volumes/web/cnidarian/BiRNAseq_GPL11353_v6ref.txt
In [37]:
!head -3 /Volumes/web/cnidarian/BiRNAseq_GPL11353_v6ref.txt
In [35]:
!egrep -wi --color 'CF369228|EF219426|EF219427|EF219428|EF219429' /Volumes/web/cnidarian/BiRNAseq_GPL11353_v6ref.txt
In [36]:
!egrep -wi --color 'CU998852' /Volumes/web/cnidarian/BiRNAseq_GPL11353_v6ref.txt
BiGo Sperm RNAseq v GPL11353 array¶
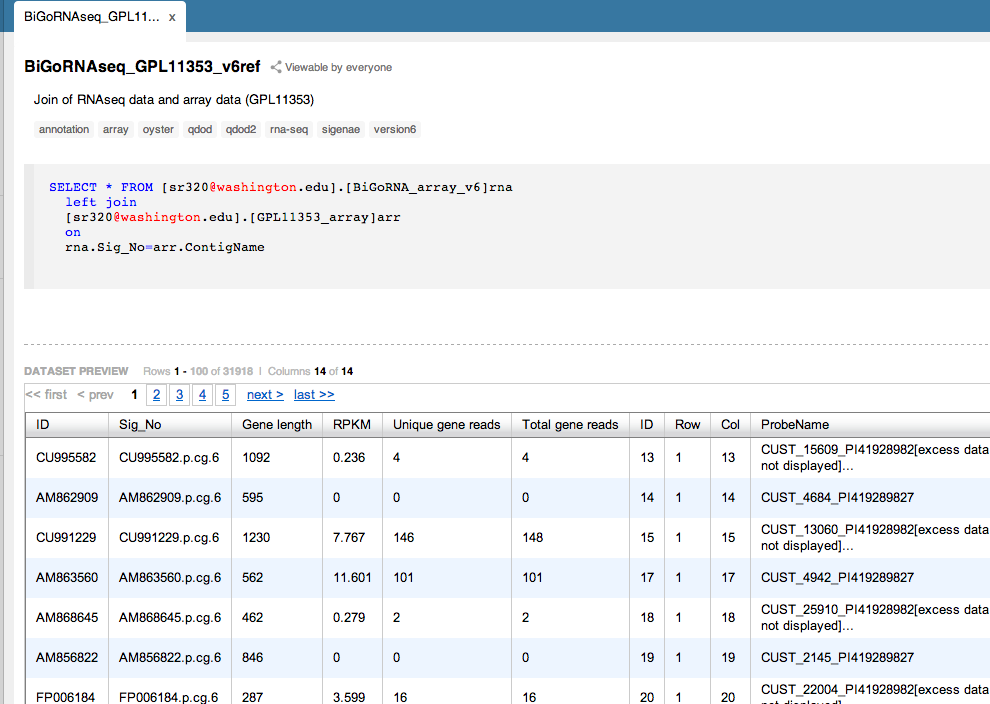
Supplementary tables from Dheilly et al¶
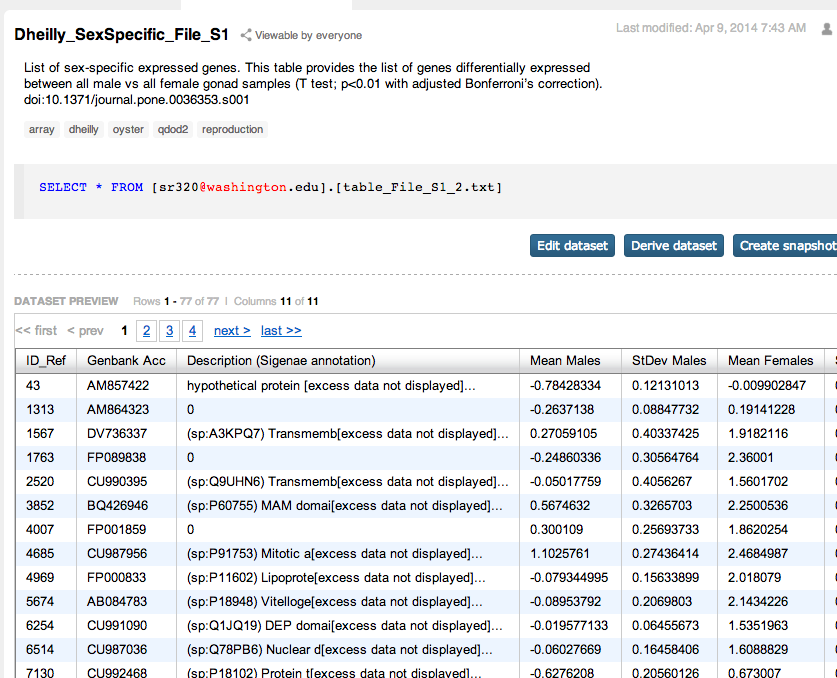
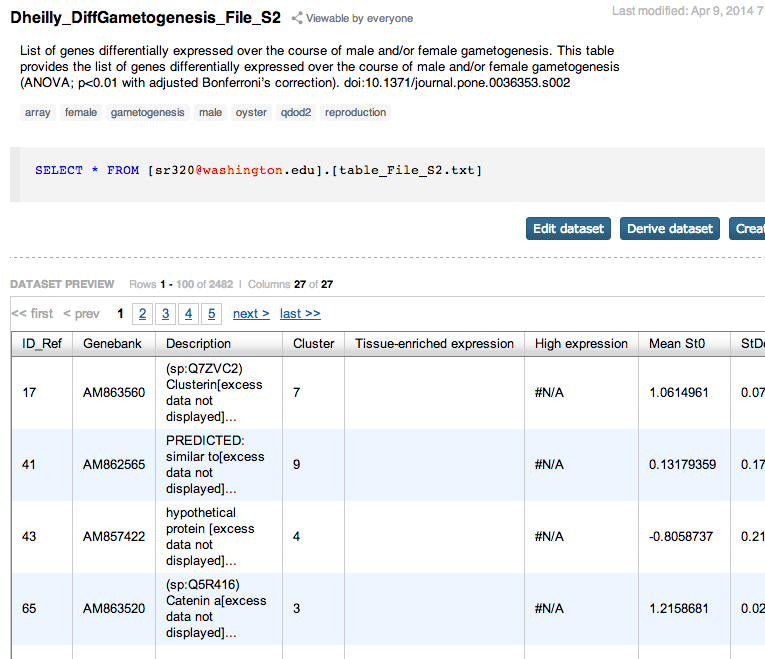
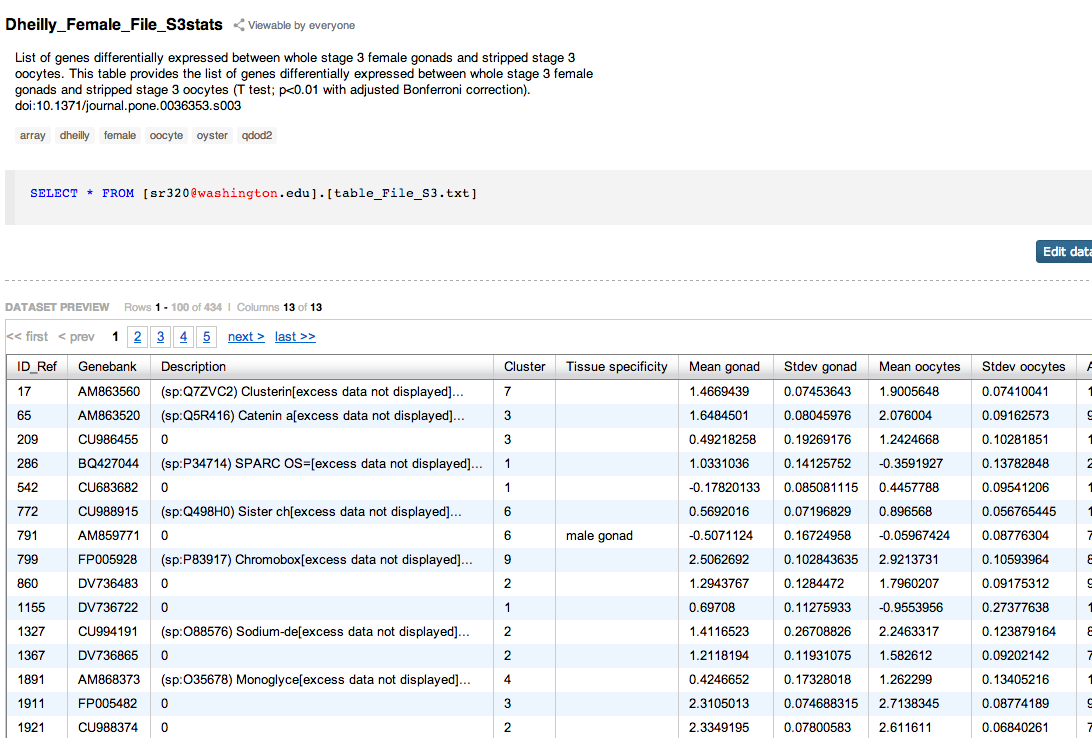
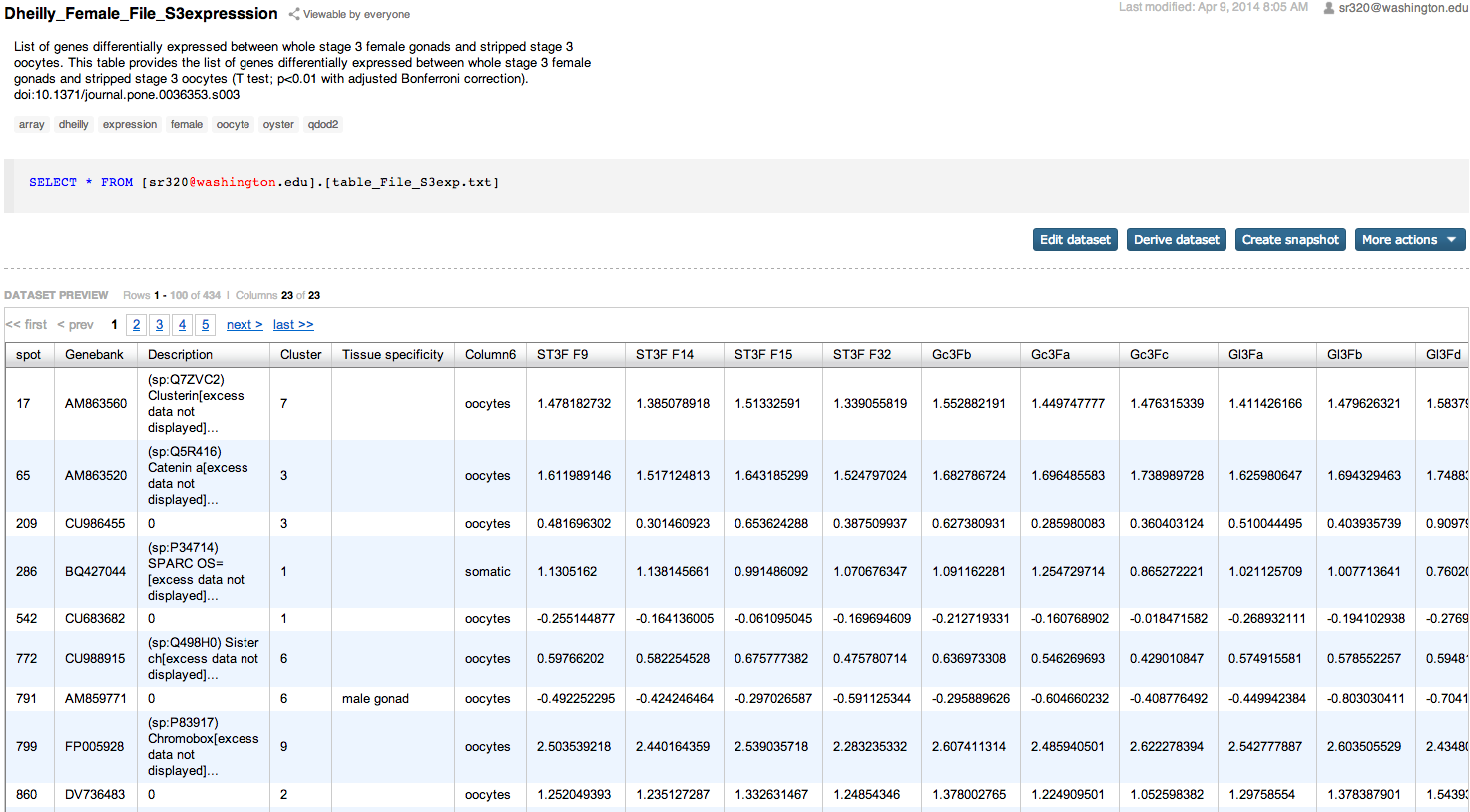
In [ ]: