Re-defining canonical C gigas Genome Tracks¶
via Ensembl
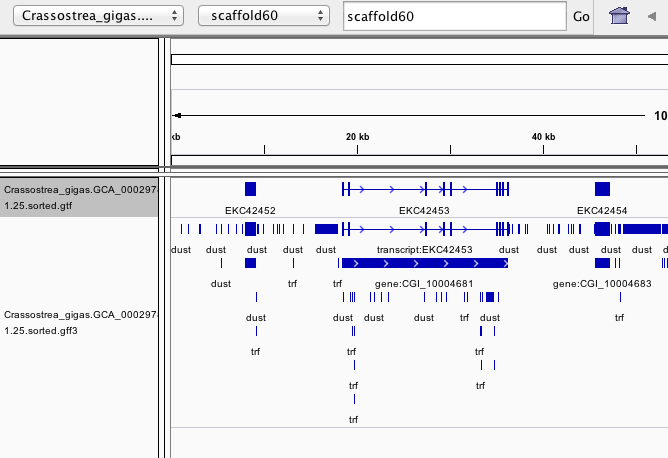
In [19]:
!tail -3 /Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gff3
scaffold44098 dust repeat_region 518076 518099 . . . Name=dust;class=dust;type=Dust scaffold44098 dust repeat_region 519261 519281 . . . Name=dust;class=dust;type=Dust scaffold44098 trf repeat_region 519261 519281 . . . Name=trf;class=trf;repeat_consensus=AT;type=Tandem repeats
In [8]:
!cut -f 3 \
/Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gff3 \
| sort | uniq -c | sed '/#/d'
186890 CDS 5 RNA 189468 exon 26114 gene 28 miRNA 28 miRNA_gene 1410 pseudogenic_tRNA 13 rRNA 13 rRNA_gene 875275 repeat_region 47 snRNA 47 snRNA_gene 20 snoRNA 20 snoRNA_gene 994 tRNA_gene 28523 transcript
In [10]:
!cut -f 2,3 \
/Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gff3 \
| sort | uniq -c | sed '/#/d'
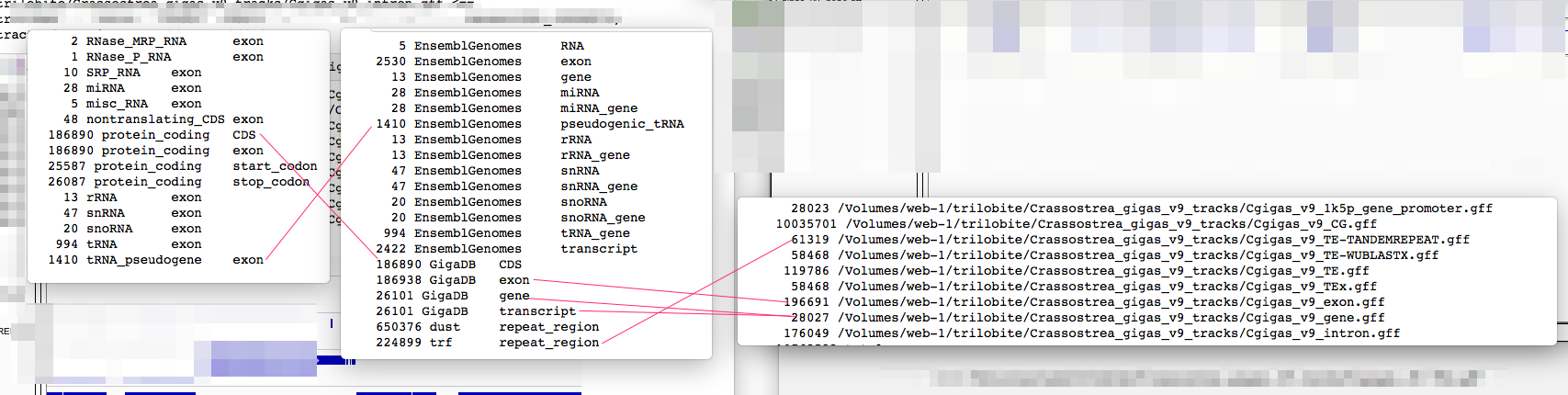
5 EnsemblGenomes RNA 2530 EnsemblGenomes exon 13 EnsemblGenomes gene 28 EnsemblGenomes miRNA 28 EnsemblGenomes miRNA_gene 1410 EnsemblGenomes pseudogenic_tRNA 13 EnsemblGenomes rRNA 13 EnsemblGenomes rRNA_gene 47 EnsemblGenomes snRNA 47 EnsemblGenomes snRNA_gene 20 EnsemblGenomes snoRNA 20 EnsemblGenomes snoRNA_gene 994 EnsemblGenomes tRNA_gene 2422 EnsemblGenomes transcript 186890 GigaDB CDS 186938 GigaDB exon 26101 GigaDB gene 26101 GigaDB transcript 650376 dust repeat_region 224899 trf repeat_region
In [3]:
!tail /Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gtf
scaffold44098 protein_coding CDS 509746 510288 . - 0 gene_id "CGI_10017729"; gene_version "1"; transcript_id "EKC17988"; transcript_version "1"; exon_number "2"; protein_id "EKC17988"; scaffold44098 protein_coding exon 514550 514690 . - . gene_id "CGI_10017729"; gene_version "1"; transcript_id "EKC17988"; transcript_version "1"; exon_number "1"; seqedit "false"; scaffold44098 protein_coding CDS 514550 514690 . - 0 gene_id "CGI_10017729"; gene_version "1"; transcript_id "EKC17988"; transcript_version "1"; exon_number "1"; protein_id "EKC17988"; scaffold44098 protein_coding start_codon 514688 514690 . - 0 gene_id "CGI_10017729"; gene_version "1"; transcript_id "EKC17988"; transcript_version "1"; exon_number "1"; scaffold44098 protein_coding exon 514859 515511 . - . gene_id "CGI_10017730"; gene_version "1"; transcript_id "EKC17989"; transcript_version "1"; exon_number "2"; seqedit "false"; scaffold44098 protein_coding stop_codon 514859 514861 . - 0 gene_id "CGI_10017730"; gene_version "1"; transcript_id "EKC17989"; transcript_version "1"; exon_number "2"; scaffold44098 protein_coding CDS 514862 515511 . - 2 gene_id "CGI_10017730"; gene_version "1"; transcript_id "EKC17989"; transcript_version "1"; exon_number "2"; protein_id "EKC17989"; scaffold44098 protein_coding exon 515871 515877 . - . gene_id "CGI_10017730"; gene_version "1"; transcript_id "EKC17989"; transcript_version "1"; exon_number "1"; seqedit "false"; scaffold44098 protein_coding CDS 515871 515877 . - 0 gene_id "CGI_10017730"; gene_version "1"; transcript_id "EKC17989"; transcript_version "1"; exon_number "1"; protein_id "EKC17989"; scaffold44098 protein_coding start_codon 515875 515877 . - 0 gene_id "CGI_10017730"; gene_version "1"; transcript_id "EKC17989"; transcript_version "1"; exon_number "1";
In [12]:
!cut -f 2,3 \
/Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gtf \
| sort | uniq -c | sed '/#/d'
2 RNase_MRP_RNA exon 1 RNase_P_RNA exon 10 SRP_RNA exon 28 miRNA exon 5 misc_RNA exon 48 nontranslating_CDS exon 186890 protein_coding CDS 186890 protein_coding exon 25587 protein_coding start_codon 26087 protein_coding stop_codon 13 rRNA exon 47 snRNA exon 20 snoRNA exon 994 tRNA exon 1410 tRNA_pseudogene exon
via GigaDB aka version9¶
In [18]:
!tail -2 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_*.gff
==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_1k5p_gene_promoter.gff <== ==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_CG.gff <== scaffold38980 fuzznuc nucleotide_motif 63903 63904 2 + . ID=scaffold38980.744;note=*pat pattern:CG scaffold38980 fuzznuc nucleotide_motif 64051 64052 2 + . ID=scaffold38980.745;note=*pat pattern:CG ==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE-TANDEMREPEAT.gff <== scaffold999 TRF Tandem_Repeat 153009 153196 189 + . . scaffold999 TRF Tandem_Repeat 166754 166792 69 + . . ==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE-WUBLASTX.gff <== scaffold1009 WUBlastX DNA_TcMar-Tc2 1790325 1790603 20 + . . scaffold983 WUBlastX DNA_TcMar-Tc1 369636 369770 26 - . . ==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE.gff <== scaffold1009 WUBlastX DNA_TcMar-Tc2 1790325 1790603 20 + . . scaffold983 WUBlastX DNA_TcMar-Tc1 369636 369770 26 - . . ==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TEx.gff <== scaffold1009 WUBlastX DNA_TcMar-Tc2 1790325 1790603 20 + . . scaffold983 WUBlastX DNA_TcMar-Tc1 369636 369770 26 - . . ==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_exon.gff <== scaffold22 GLEAN CDS 1870289 1870360 . - 0 Parent=CGI_10028939; scaffold22 GLEAN CDS 1869336 1869428 . - 0 Parent=CGI_10028939; ==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_gene.gff <== scaffold22 GLEAN mRNA 1863760 1864161 0.544455 + . ID=CGI_10028938; scaffold22 GLEAN mRNA 1869336 1885890 0.999933 - . ID=CGI_10028939; ==> /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_intron.gff <==
In [15]:
!wc -l /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_*.gff
28023 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_1k5p_gene_promoter.gff 10035701 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_CG.gff 61319 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE-TANDEMREPEAT.gff 58468 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE-WUBLASTX.gff 119786 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE.gff 58468 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TEx.gff 196691 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_exon.gff 28027 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_gene.gff 176049 /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_intron.gff 10762532 total
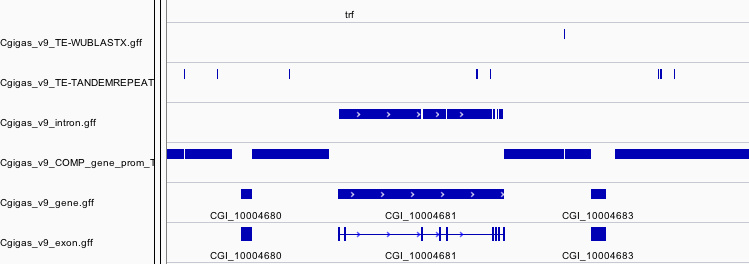
Comparison¶
Lets see if can take all array and intersect with Ensembl gff¶
In [27]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.2M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gff3 \
| cut -f 6,7 \
| sort | uniq -c | sed '/#/d'
1373 GigaDB CDS 1373 GigaDB exon 8468 GigaDB gene 8468 GigaDB transcript 1240 dust repeat_region 975 trf repeat_region
In [28]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.4M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gff3 \
| cut -f 6,7 \
| sort | uniq -c | sed '/#/d'
2 EnsemblGenomes exon 1 EnsemblGenomes pseudogenic_tRNA 1 EnsemblGenomes tRNA_gene 2 EnsemblGenomes transcript 1177 GigaDB CDS 1177 GigaDB exon 8491 GigaDB gene 8491 GigaDB transcript 1320 dust repeat_region 873 trf repeat_region
In [29]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.6M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gff3 \
| cut -f 6,7 \
| sort | uniq -c | sed '/#/d'
1 EnsemblGenomes exon 1 EnsemblGenomes snRNA 1 EnsemblGenomes snRNA_gene 947 GigaDB CDS 948 GigaDB exon 9689 GigaDB gene 9689 GigaDB transcript 1591 dust repeat_region 864 trf repeat_region
In [3]:
!intersectbed \
-wb \
-a /Users/sr320/git-repos/paper-Temp-stress/ipynb/data/array-design/OID40453_probe_locations.gff \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.25.sorted.gff3 \
| cut -f 11,12 \
| sort | uniq -c | sed '/#/d'
5 EnsemblGenomes RNA 444 EnsemblGenomes exon 6 EnsemblGenomes gene 2 EnsemblGenomes miRNA 2 EnsemblGenomes miRNA_gene 259 EnsemblGenomes pseudogenic_tRNA 14 EnsemblGenomes snRNA 14 EnsemblGenomes snRNA_gene 6 EnsemblGenomes snoRNA 6 EnsemblGenomes snoRNA_gene 152 EnsemblGenomes tRNA_gene 422 EnsemblGenomes transcript 157279 GigaDB CDS 157307 GigaDB exon 600445 GigaDB gene 600445 GigaDB transcript 56210 dust repeat_region 42390 trf repeat_region
TEs¶
In [6]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.2M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE-WUBLASTX.gff \
| cut -f 6 \
| sort | uniq -c | sed '/#/d'
383 WUBlastX
In [9]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.4M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE-WUBLASTX.gff \
| cut -f 6 \
| sort | uniq -c | sed '/#/d'
254 WUBlastX
In [8]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.6M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE-WUBLASTX.gff \
| cut -f 6 \
| sort | uniq -c | sed '/#/d'
168 WUBlastX
In [10]:
!intersectbed \
-wb \
-a /Users/sr320/git-repos/paper-Temp-stress/ipynb/data/array-design/OID40453_probe_locations.gff \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE-WUBLASTX.gff \
| cut -f 11 \
| sort | uniq -c | sed '/#/d'
10322 WUBlastX
Promoters¶
In [12]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.2M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_1k5p_gene_promoter.gff \
| cut -f 6,7 \
| sort | uniq -c | sed '/#/d'
976 flankbed promoter
In [13]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.4M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_1k5p_gene_promoter.gff \
| cut -f 6,7 \
| sort | uniq -c | sed '/#/d'
992 flankbed promoter
In [14]:
!intersectbed \
-wb \
-a ./data/2014.07.02.colson/genomeBrowserTracks/logFC_HS-preHS/2014.07.02.6M_sig.bedGraph \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_1k5p_gene_promoter.gff \
| cut -f 6,7 \
| sort | uniq -c | sed '/#/d'
1248 flankbed promoter
In [15]:
!intersectbed \
-wb \
-a /Users/sr320/git-repos/paper-Temp-stress/ipynb/data/array-design/OID40453_probe_locations.gff \
-b /Volumes/web-1/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_1k5p_gene_promoter.gff \
| cut -f 11 \
| sort | uniq -c | sed '/#/d'
66368 flankbed
Plot¶
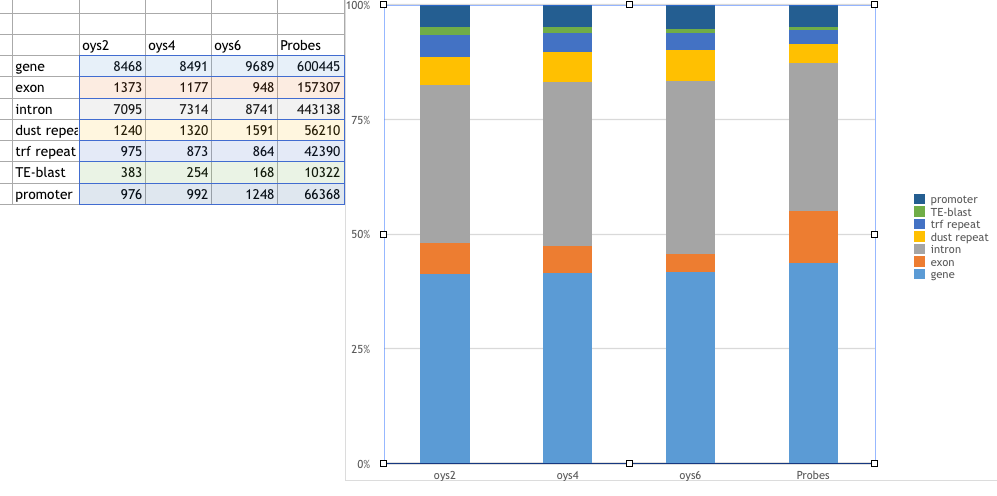
oys2 oys4 oys6 Probes
gene 8468 8491 9689 600445
exon 1373 1177 948 157307
intron 7095 7314 8741 443138
dust repeat 1240 1320 1591 56210
trf repeat 975 873 864 42390
TE-blast 383 254 168 10322
promoter 976 992 1248 66368
Analysis of one proportion¶
In [2]:
%pylab inline
import scipy.stats as stats
Populating the interactive namespace from numpy and matplotlib
In [3]:
# Get the data Probes exon
numTotal = 697753
numPositive = 157307
# Calculate the confidence intervals
p = float(numPositive)/numTotal
se = sqrt(p*(1-p)/numTotal)
td = stats.t(numTotal-1)
ci = p + array([-1,1])*td.isf(0.025)*se
# Print them
print('ONE PROPORTION')
print('The confidence interval for the given sample is {0:5.3f} to {1:5.3f}'.format(
ci[0], ci[1]))
ONE PROPORTION The confidence interval for the given sample is 0.224 to 0.226
Chi-square test to a 2x2 table¶
In [4]:
# Enter the data comparing Oyster 2; probes at intron
obs = array([[7095, 10028], [443138, 697753]])
# Calculate the chi-square test
chi2_corrected = stats.chi2_contingency(obs, correction=True)
chi2_uncorrected = stats.chi2_contingency(obs, correction=False)
# Print the result
print('CHI SQUARE')
print('The corrected chi2 value is {0:5.3f}, with p={1:5.3f}'.format(chi2_corrected[0], chi2_corrected[1]))
print('The uncorrected chi2 value is {0:5.3f}, with p={1:5.3f}'.format(chi2_uncorrected[0], chi2_uncorrected[1]))
CHI SQUARE The corrected chi2 value is 47.663, with p=0.000 The uncorrected chi2 value is 47.772, with p=0.000
In [5]:
# Enter the data comparing Oyster 2; probes at gene
obs = array([[8468, 10028], [600445, 697753]])
# Calculate the chi-square test
chi2_corrected = stats.chi2_contingency(obs, correction=True)
chi2_uncorrected = stats.chi2_contingency(obs, correction=False)
# Print the result
print('CHI SQUARE')
print('The corrected chi2 value is {0:5.3f}, with p={1:5.3f}'.format(chi2_corrected[0], chi2_corrected[1]))
print('The uncorrected chi2 value is {0:5.3f}, with p={1:5.3f}'.format(chi2_uncorrected[0], chi2_uncorrected[1]))
CHI SQUARE The corrected chi2 value is 1.597, with p=0.206 The uncorrected chi2 value is 1.616, with p=0.204
In [ ]: