Larval Transcriptome of OsHV exposed oysters¶
A re-examination of Cgigas larval transcriptome using genome.
Solid Files -- http://eagle.fish.washington.edu/whale/index.php?dir=GE%2Freads%2F
Importing into CLCv6
Quality trim¶
Ambiguous trim = Yes Ambiguous limit = 2 Quality trim = Yes Quality limit = 0.05 Create report = Yes Save discarded sequences = No Remove 5' terminal nucleotides = No Minimum number of nucleotides in reads = 20 Discard short reads = Yes Remove 3' terminal nucleotides = No Discard long reads = No Save broken pairs = No

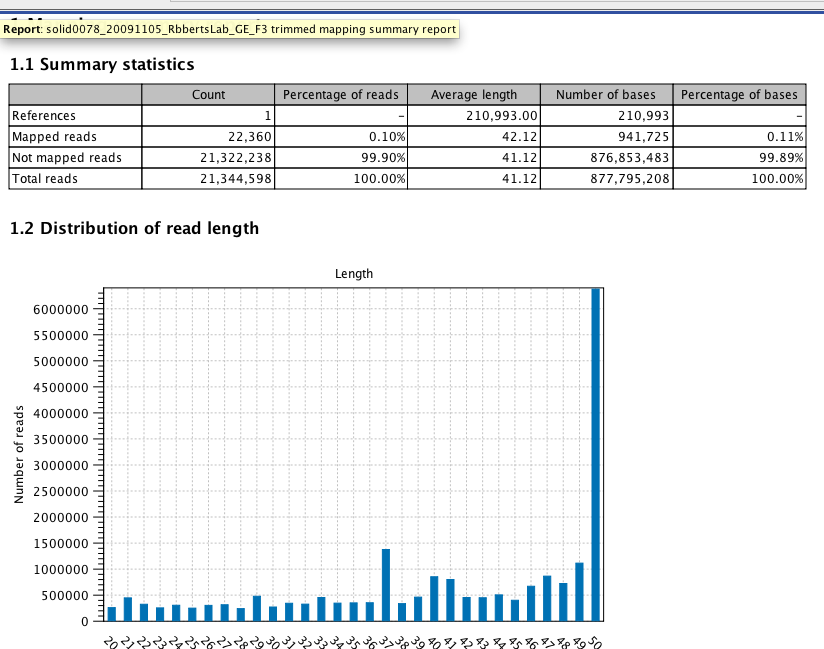
QC Report¶
https://docs.google.com/file/d/0B9V_gF766XZAQXp4OE45VXc0U00/preview
Mapping to Transcriptome¶
RNA-seq to genes
Parameters
Use strand specific assembly = No Create report = Yes References = oyster_v9_gene Count paired reads as two = No Use colorspace encoding = Yes Minimum number of reads = 10 Additional upstream bases = 0 Minimum read count fusion gene table = 5 Minimum length of putative exons = 25 Minimum exon coverage fraction = 0.2 Minimum length fraction (long reads) = 0.9 Use annotations for gene and transcript identification = No Create fusion gene table = No Expression value = TOTAL_GENE_COUNT Minimum similarity fraction = 0.8 Expression level = Genes Create list of unmapped reads = No Unspecific match limit = 5 Exon discovery = No Organism type = PROKARYOTE Additional downstream bases = 0 Maximum paired distance = 250 Minimum paired distance = 180 Strand = Forward Maximum number of mismatches allowed (applies to short reads) = 2 Expression value = Number of reads mapped to the gene

#tab delim RNA-Seq file
!head /Volumes/web/cnidarian/solid0078_20091105_RobertsLab_GE_F3\ trimmed\ RNA-Seq.txt
"Feature ID" "Expression values" "Gene length" "Unique gene reads" "Total gene reads" "RPKM" CGI_10000780 2 1350 2 2 0.272 CGI_10000456 28 438 18 28 11.735 CGI_10000457 5 603 4 5 1.522 CGI_10000774 1 375 1 1 0.49 CGI_10000917 2 426 2 2 0.862 CGI_10000861 7 2004 7 7 0.641 CGI_10000994 96 1635 74 96 10.779 CGI_10000643 0 552 0 0 0 CGI_10000763 4 249 4 4 2.949
#SAM output
!head /Volumes/web/cnidarian/solid0078_20091105_RobertsLab_GE_F3_tr_RNA-Seq.sam
@HD VN:1.0 SO:unsorted @SQ SN:CGI_10000780 LN:1350 @SQ SN:CGI_10000456 LN:438 @SQ SN:CGI_10000457 LN:603 @SQ SN:CGI_10000774 LN:375 @SQ SN:CGI_10000917 LN:426 @SQ SN:CGI_10000861 LN:2004 @SQ SN:CGI_10000994 LN:1635 @SQ SN:CGI_10000643 LN:552 @SQ SN:CGI_10000763 LN:249
#SAM output
!tail /Volumes/web/cnidarian/solid0078_20091105_RobertsLab_GE_F3_tr_RNA-Seq.sam
read_5447380 0 CGI_10003117 263 60 50M * 0 0 CCTCGCATTGGAAAACCCCAGTGTTTGGGTGGGTGTCACAAGAGGAAGAA @A@?B=AA;@BB@:@<BA?:?B=:92?3;@?>:;<=2;>6:?98>:77/7 NH:i:1 CS:Z:T20223313010200010001211100100110011121110222020220 read_5447381 0 CGI_10003117 357 60 48M * 0 0 ACCATTGAATCGATTTCCAAATGTGATGTTCCATCTGCAGTCCCTCTT B@B@BA?ABBAAABAABBA@5A@?@ABA>AAB@=?AB@?@7@@8@?@5 NH:i:1 CS:Z:T310130120323230020100311123110201322131212002220 read_5447382 0 CGI_10003117 375 60 44M * 0 0 AAATGTGATGTTCCATCTGCAGTCCCTCTTTAAATGGTCACAGT @?BB@@BABB>B@A??BBB9=B:B@@AAB<59>@><@>A?>?<: NH:i:1 CS:Z:T30031112311020132213121200222003003101211121 read_5447383 0 CGI_10003117 375 60 44M * 0 0 AAATGTGATGTTCCATCTGCAGTCCCTCTTTAAATGGTCACAGT >@BB@?BBBA<BAA=ABAB<=@;>>@B>@<78?@=9@?A>>?;; NH:i:1 CS:Z:T30031112311020132213121200222003003101211121 read_5447384 0 CGI_10003117 375 60 44M * 0 0 AAATGTGATGTTCCATCTGCAGTCCCTCTTTAAATGGTCACAGT @BB@@>@BAA=@@B?=B@B::@>5==?@@=/*?B81>;AA?@:7 NH:i:1 CS:Z:T30031112311020132213121200222003003101211121 read_5447385 0 CGI_10003117 507 60 29M * 0 0 GATGAATTGGTATAACATTGTCAACTCTT BBB?<@BBA=@BA?>=AA@;6@?<:8AA/ NH:i:1 CS:Z:T12312030101333011301121012220 read_5447386 0 CGI_10003117 609 60 3S35M * 0 0 AGCACAGATCCAAACTGGAATTTACCAGAACCGCCAGA BBBABBBABBBB>BA>@??BB@?AB@B<3@A>=9AA>= NH:i:1 CS:Z:T32311122320100121020300310122010330122 read_5447387 0 CGI_10003117 609 60 3S47M * 0 0 AGCACAAATCCAAACTGGAATTTACCAGAACCGCCAGAAGAATACATCCC BBBABAA=BBBA@?B@=@?AB??BA?@<;A@:>=@>8:7@=><=@=<5;? NH:i:1 CS:Z:T32311100320100121020300310122010330122022033113200 read_5447388 0 CGI_10003117 609 60 3S35M * 0 0 AGCACAGATCCAAACTGGAATTTACCAGAACCGCCAGA BBBBBBB@BBBBBB@B?BBBBB@BB@@==BA;A>AA@= NH:i:1 CS:Z:T32311122320100121020300310122010330122 read_5447389 0 CGI_10003117 609 60 48M2S * 0 0 ACAGATCCAAACTGGAATTTACCAGAACCGCCAGAAGAATACATCCCCCC BA@BBBBAB@BBAAB>ABB?BAA??>;=??>A>=97<?>@@AA537<;@= NH:i:1 CS:Z:T31122320100121020300310122010330122022033113200000
#Lets figure out a way to visualize what genes are expressed
#Take RNA-seq file import into SQLShare
#imported /Volumes/web/cnidarian/solid0078_20091105_RobertsLab_GE_F3\ trimmed\ RNA-Seq.txt
#cleaned up
#joining with annotation data.
#create generic SQLShare Wiki workflow.
!head /Volumes/web/cnidarian/Cgigas_larvae_RNAseq_OsHV_GO.csv
!wc /Volumes/web/cnidarian/Cgigas_larvae_RNAseq_OsHV_GO.csv
121979 315335 6935565 /Volumes/web/cnidarian/Cgigas_larvae_RNAseq_OsHV_GO.csv
#into GoCategorizer then ManyEyes

Enrichment Analysis¶
Background - all SPIDs associated with oyster transcriptome Gene list - SPID of gene with at least 10 unique reads
Kegg
;%20National%20Institute%20of%20Allergies%20and%20Infectious%20Diseases%20(NIAID);%20Science%20Applications%20International%20Corporation%20(SAIC).png)
http://eagle.fish.washington.edu/cnidarian/chart_5EB4CB21C87A1374501610998.txt
BP-Fat
;%20National%20Institute%20of%20Allergies%20and%20Infectious%20Diseases%20(NIAID);%20Science%20Applications%20International%20Corporation%20(SAIC)%20and.png)
http://eagle.fish.washington.edu/cnidarian/chart_5EB4CB21C87A1374501882704.txt
pvalue <.05 into Revigo
R script for treemap http://eagle.fish.washington.edu/cnidarian/OsHV_Cg_REVIGO_treemap.r
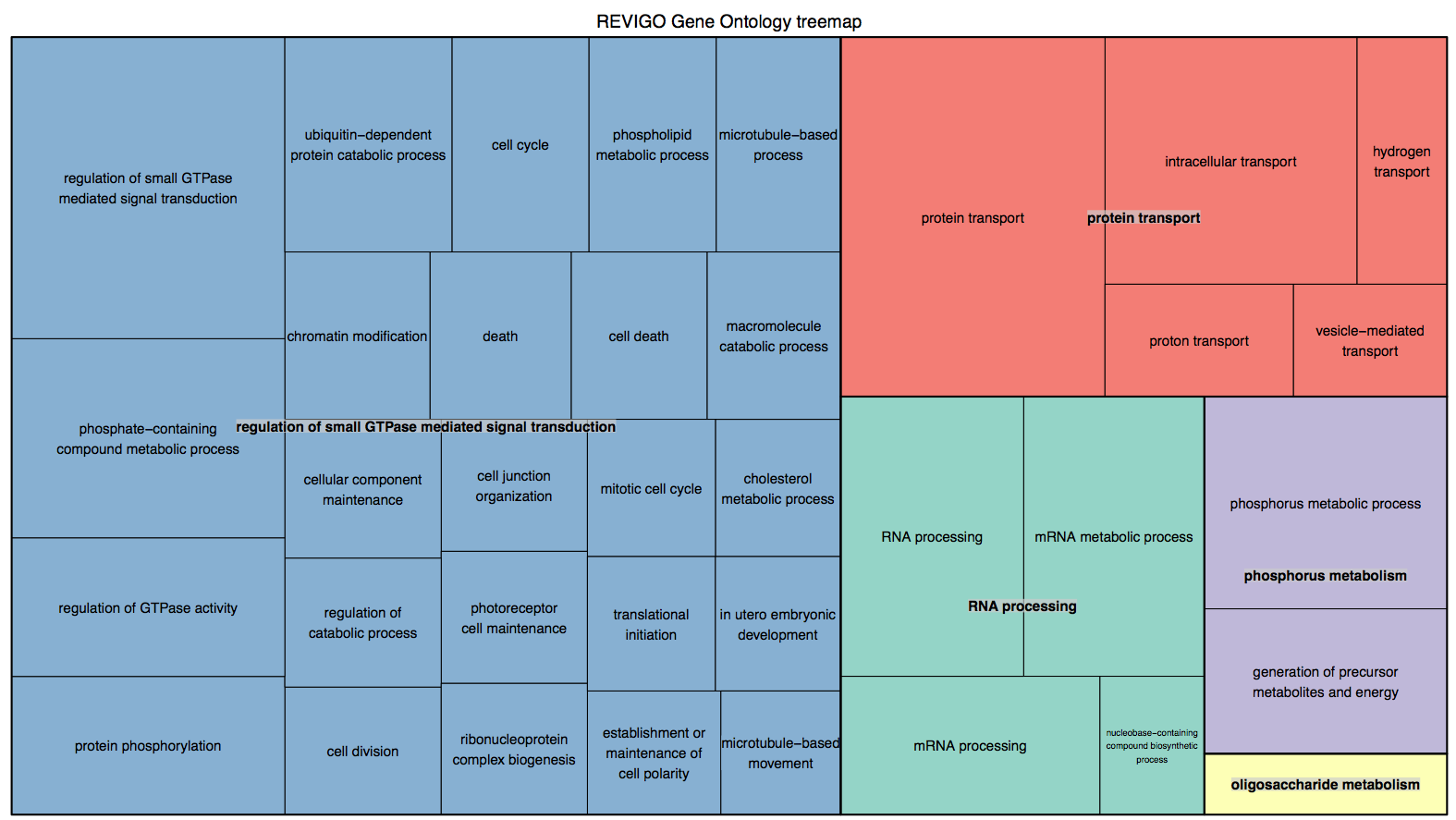 mm
mm
iPath¶
Mapping to OsHV¶
http://www.ncbi.nlm.nih.gov/nuccore/NC_005881.1
imported genbank format to hav CDS, gene information..
started in CLC using same parameters..
Minimum number of reads = 10
Minimum exon coverage fraction = 0.2
Additional downstream bases = 0
Use colorspace encoding = Yes
Create report = Yes
Use strand specific assembly = No
Count paired reads as two = No
Minimum length fraction (long reads) = 0.9
Additional upstream bases = 0
Unspecific match limit = 5
Expression value = RPKM
Minimum read count fusion gene table = 5
Create fusion gene table = No
Minimum paired distance = 180
Use annotations for gene and transcript identification = Yes
Strand = Forward
Organism type = PROKARYOTE
Maximum number of mismatches allowed (applies to short reads) = 2
Minimum similarity fraction = 0.8
References = NC_005881
Expression level = Genes
Minimum length of putative exons = 25
Maximum paired distance = 250
Create list of unmapped reads = No
Exon discovery = No
Expression value = Read Per Kilobase of exon Model value
Found: 127 genes. Total number of reads : 21344598 ( single reads: 21344598, paired reads: 0 ) Total number of mapped reads : 21135 ( single reads: 21135, paired reads: 0 ) Total number of unmapped reads : 21323463 ( single reads: 21323463, paired reads: 0 )


!head /Volumes/web/cnidarian/solid0078_20091105_RobertsLab_GE_F3trim_RNAseqOSHV.csv
"Feature ID","Expression values","Gene length","Unique gene reads","Total gene reads","RPKM","Chromosome region start","Chromosome region end" "OsHV1_gp001","423.399","447","0","4","423.399","115","561" "OsHV1_gp002","751.03","504","0","8","751.03","680","1183" "OsHV1_gp003","61.85","765","0","1","61.85","1890","2654" "OsHV1_gp004","2748.769","1050","0","61","2748.769","3384","4433" "OsHV1_gp005","3168.303","2031","70","136","3168.303","6421","8451" "OsHV1_gp006","1534.465","3546","115","115","1534.465","8628","12173" "OsHV1_gp007","1429.304","960","29","29","1429.304","12211","13170" "OsHV1_gp008","4016.047","1779","151","151","4016.047","13258","15036" "OsHV1_gp009","6345.436","1029","138","138","6345.436","15297","16325"
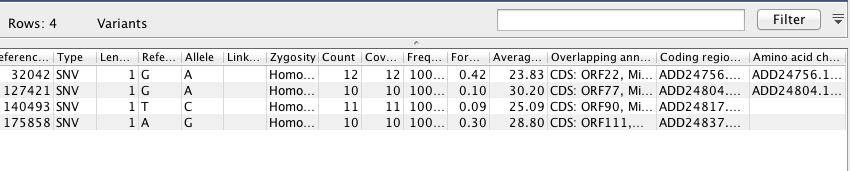
SNP ID based on Reference Mapping¶
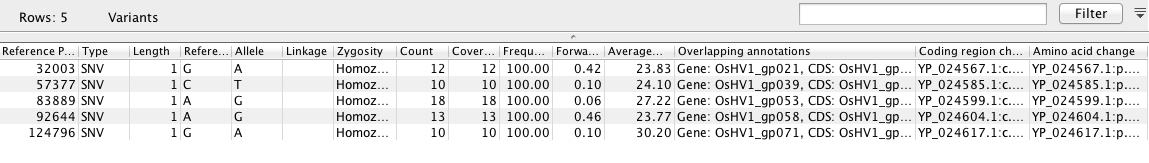
!cat /Volumes/web/cnidarian/OsHV_snp_table
"Reference Position" "Type" "Length" "Reference" "Allele" "Linkage" "Zygosity" "Count" "Coverage" "Frequency" "Forward/reverse balance" "Average quality" "Overlapping annotations" "Coding region change" "Amino acid change" 32003 SNV 1 G A Homozygous 12 12 100 0.417 23.833 Gene: OsHV1_gp021, CDS: OsHV1_gp021, Misc. feature: UL YP_024567.1:c.1533G>A YP_024567.1:p.Met511Ile 57377 SNV 1 C T Homozygous 10 10 100 0.1 24.1 Gene: OsHV1_gp039, CDS: OsHV1_gp039, Misc. feature: UL YP_024585.1:c.1468C>T YP_024585.1:p.Pro490Ser 83889 SNV 1 A G Homozygous 18 18 100 0.056 27.222 Gene: OsHV1_gp053, CDS: OsHV1_gp053, Misc. feature: UL YP_024599.1:c.554A>G YP_024599.1:p.Glu185Gly 92644 SNV 1 A G Homozygous 13 13 100 0.462 23.769 Gene: OsHV1_gp058, CDS: OsHV1_gp058, Misc. feature: UL YP_024604.1:c.1349A>G YP_024604.1:p.Tyr450Cys 124796 SNV 1 G A Homozygous 10 10 100 0.1 30.2 Gene: OsHV1_gp071, CDS: OsHV1_gp071, Misc. feature: UL YP_024617.1:c.2083G>A YP_024617.1:p.Ala695Thr
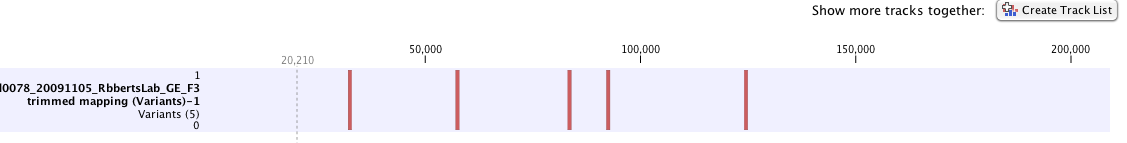
IGV Browser for OsHV¶
!head /Volumes/web/cnidarian/solid0078_20091105_RbbertsLab_GE_F3\ trimmed\ mapping.sam
@HD VN:1.0 SO:unsorted @SQ SN:NC_005881 LN:207439 SP:Ostreid herpesvirus 1 @PG ID:0 PN:clcgenomicswb VN:7.0 read_1 0 NC_005881 2764 60 2S18M * 0 0 CATGGGGGGGGGGGGGGGGG BB@??979/12.=<+9-;22 NH:i:1 CS:Z:T21312000000000000000 read_2 16 NC_005881 7624 60 32M * 0 0 CACTTGACAACATCCATACATATCGTCATCCA >9;3,@5>/<@7?=8BA>B:B?A<=@AA;7B@ NH:i:1 CS:Z:T01023121323331133102311011200211 read_3 16 NC_005881 7624 60 26M * 0 0 CACTTGACAACATCCATACATATCGT ?>;>2?5>?><AA@=ABA>BBBBABB NH:i:1 CS:Z:T31323331133102311011210211 read_4 16 NC_005881 7638 60 46M * 0 0 CATACATATCGTCATCCAGGGTCTCTCCGTGTTTAGGATATAGCGT <?A>0=??6&A8:?9>>>@9??>@@B6@ABBA?ABBBB>BBBBBAB NH:i:1 CS:Z:T3133233332023001113022222100210231211233311331 read_5 0 NC_005881 7640 60 34M * 0 0 TACATATCGTCATCCAGGGTCTCTCCGTGTTTAG BBAABBBBBABBBBABBBBAB@BB@BBA@+>BBB NH:i:1 CS:Z:T0311333231213201200122222031110032 read_6 16 NC_005881 7640 60 44M * 0 0 TACATATCGTCATCCAGGGTCTCTCCGTGTTTAGGATATAGCGT =26?+:0/:04;=87232=1-1<A:<:@=@8=@BAA<@@1BA=B NH:i:1 CS:Z:T31332333320230011130222221002102312132323113 read_7 0 NC_005881 7640 60 46M3S * 0 0 TACATATCGTCATCCAGGGTCTCTCCGTGTTTAGGATATAGCGTATATA BBBBBBBBB=BBBBBABB@AB@BA@BB=B/>ABA+@BB?%B@B;??6B> NH:i:1 CS:Z:T0311333231213201200122222031110032023333233133333
!head /Volumes/web/cnidarian/NC_005881
>NC_005881 Ostreid herpesvirus 1, complete genome. CCCCCCACCTCCCCAACACCTCCCCCATCCTCCCCACCTCCTCCCCCTCCTCTCTTCCGC CCGCGATCCCGCCAATACCCATAATGCACCTGGGCACTCTCTTTTTTCCTTTCCTTATCC AAGATGTCCGCCCATTGCCAGGTACAGCCTTCCCACCGTGTGAACAATGTCCATCCTCTT CTCCGAGACTTCCCTGACCAGATTGTCGTAATCCAATTGACACATTCTCGTCAATGCCCT CCTCATATTCTCCATCGGCCAACTGTCGTCTCTACTCATGGTCATAAACAATCCCAATCC ACTCTTGGCATCCCGCAACCTTTCCAATAGCCTCCCGAATTCGTCTACCGCCGCCTTATC GTCGTCCGTGCTGCAATGTGGTCTTACCGCATTTTTAAGCAATGCGCACGCCACTCTCAA TTCCTGACAGGTAATCTCCTCCACCGGTTTCCTATCGTGTAATAGACTGACCACGGCGGC CATGTCTCTCAGTTCCTTGCTCATCTCACCACCGCCAACCAATTCAGCAGTGACAGTTAC
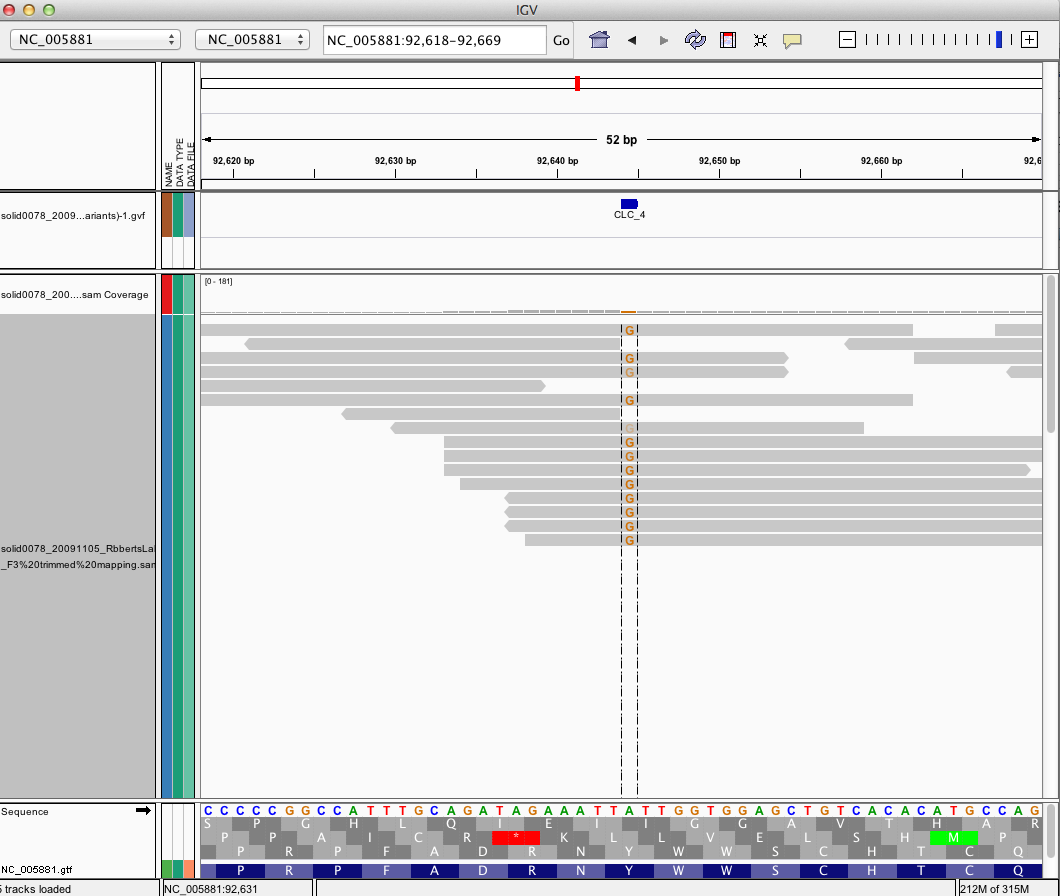
LINKOUT to xml http://eagle.fish.washington.edu/cnidarian/OsHV_igv_session.xml
Mapping to GQ153938¶