Redux on data QC¶
In [1]:
!date
Fri May 30 08:28:32 PDT 2014
In [3]:
!say "Back at it"

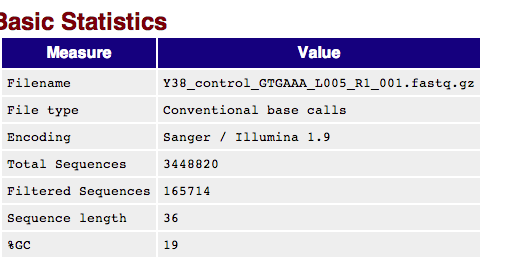
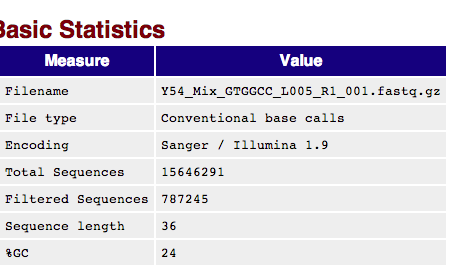
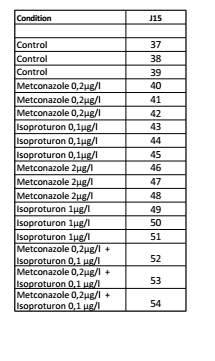
Point- Controls are low- November run only one filtered data set was produced.¶
In [ ]:
In [21]:
!python methratio.py -d /Volumes/Monarch/cnidary/oyster.v9.fa -z -u -g -o /Volumes/Monarch/cnidary/EY_mixmethratio.txt -s /Applications/bsmap-2.74/samtools /Volumes/Monarch/cnidary/EY_Mixbsmapoutput.sam
@ Wed Nov 13 03:47:15 2013: reading reference /Volumes/Monarch/cnidary/oyster.v9.fa ... @ Wed Nov 13 03:48:03 2013: reading /Volumes/Monarch/cnidary/EY_Mixbsmapoutput.sam ... sh: /Applications/bsmap-2.74/samtools/samtools: No such file or directory @ Wed Nov 13 03:48:03 2013: combining CpG methylation from both strands ... @ Wed Nov 13 03:49:05 2013: writing /Volumes/Monarch/cnidary/EY_mixmethratio.txt ... @ Wed Nov 13 03:51:30 2013: done.
In [22]:
!head /Volumes/Monarch/cnidary/EY_mixmethratio.txt
chr pos strand context ratio eff_CT_count C_count CT_count rev_G_count rev_GA_count CI_lower CI_upper
In [23]:
!python methratio.py -d /Volumes/Monarch/cnidary/oyster.v9.fa -z -u -g -o /Volumes/Monarch/cnidary/EY_mixmethratio_2.txt -s /Applications/bsmap-2.74/ /Volumes/Monarch/cnidary/EY_Mixbsmapoutput.sam
@ Wed Nov 13 03:51:42 2013: reading reference /Volumes/Monarch/cnidary/oyster.v9.fa ... @ Wed Nov 13 03:52:30 2013: reading /Volumes/Monarch/cnidary/EY_Mixbsmapoutput.sam ... sh: /Applications/bsmap-2.74/samtools: is a directory @ Wed Nov 13 03:52:30 2013: combining CpG methylation from both strands ... @ Wed Nov 13 03:53:35 2013: writing /Volumes/Monarch/cnidary/EY_mixmethratio_2.txt ... @ Wed Nov 13 03:56:01 2013: done.
In [2]:
#greenbird
!python /Volumes/Bay3/Software/BSMAP/bsmap-2.74/methratio.py -d /Volumes/web/cnidarian/oyster.v9.fa -z -u -g -o /Volumes/web/cnidarian/EY_mixmethratio_3.txt -s /Volumes/Bay3/Software/BSMAP/bsmap-2.73/samtools /Volumes/web/scaphapoda/Yanouk/Mixbsmapoutput.sam
@ Wed Nov 13 04:07:28 2013: reading reference /Volumes/web/cnidarian/oyster.v9.fa ... @ Wed Nov 13 04:08:37 2013: reading /Volumes/web/scaphapoda/Yanouk/Mixbsmapoutput.sam ... [samopen] SAM header is present: 11969 sequences. @ Wed Nov 13 04:14:44 2013: combining CpG methylation from both strands ... @ Wed Nov 13 04:15:56 2013: writing /Volumes/web/cnidarian/EY_mixmethratio_3.txt ... @ Wed Nov 13 04:22:10 2013: done. total 3279697 valid mappings, 16154541 covered cytosines, average coverage: 1.19 fold.
In [3]:
!head /Volumes/web/cnidarian/EY_mixmethratio_3.txt
chr pos strand context ratio eff_CT_count C_count CT_count rev_G_count rev_GA_count CI_lower CI_upper C10005 102 + GTCTC 0.000 1.00 0 1 0 0 0.000 0.793 C10005 104 + CTCAA 0.000 1.00 0 1 0 0 0.000 0.793 C10005 117 + GACTT 0.000 1.00 0 1 0 0 0.000 0.793 C10007 45 - CAGAG 0.000 1.00 0 1 0 0 0.000 0.793 C10007 47 - GAGCT 0.000 1.00 0 1 0 0 0.000 0.793 C10007 56 - AAGCA 0.000 1.00 0 1 0 0 0.000 0.793 C10007 66 + CCCGG 0.000 1.00 0 1 1 1 0.000 0.793 C10007 68 - CGGAG 0.000 1.00 0 1 0 0 0.000 0.793 C10007 70 - GAGTT 0.000 1.00 0 1 0 0 0.000 0.793
In [4]:
!wc /Volumes/web/cnidarian/EY_mixmethratio_3.txt
16154542 193854471 939011992 /Volumes/web/cnidarian/EY_mixmethratio_3.txt
In [ ]: