Goal: Larvae BS data (adaptor trimmed) - getting it analyzed.¶
This has morphed a bit based on data exploration and options for analysis.
In [ ]:
methratio files¶

In [3]:
!head /Volumes/web/Mollusk/bs_larvae_exp/AdapterTrimmed_methratio_files/methratio_out_M1.txt
chr pos strand context ratio eff_CT_count C_count CT_count rev_G_count rev_GA_count CI_lower CI_upper C12706 51 - CAGAG 0.000 1.00 0 1 0 0 0.000 0.793 C12706 53 - GAGAG 0.000 1.00 0 1 0 0 0.000 0.793 C12706 55 - GAGGG 0.000 1.00 0 1 0 0 0.000 0.793 C12706 60 - AAGAA 0.000 1.00 0 1 0 0 0.000 0.793 C12706 69 + TCCGC 0.000 1.00 0 1 1 1 0.000 0.793 C12706 71 + CGCGT 0.000 1.00 0 1 1 1 0.000 0.793 C12706 78 - TTGAA 0.000 1.00 0 1 0 0 0.000 0.793 C12706 82 - ATGTC 0.000 1.00 0 1 0 0 0.000 0.793 C12706 95 - TTGCA 0.000 1.00 0 1 0 0 0.000 0.793
Convert with awk¶
take above file and generate table as such
chr as seqname, 'methratio' as source, 'CpG' as feature, pos as start, pos + 1 as [end], ratio as score, strand, '.' as frame, '.' as attribute
where
context like '__CG_'
and
CT_Count > 3
example output
seqname source feature start end score strand frame attribute
C12706 methratio CpG 71 72 0.000 + . .
via grep / awk¶
In [16]:
cd /Volumes/web/cnidarian/BiGo_larvae_3
/Volumes/web/cnidarian/BiGo_larvae_3
In [17]:
ls
methratio_out_M1.txt methratio_out_T1D3.txt methratio_out_T3D3.txt methratio_out_M3.txt methratio_out_T1D5.txt methratio_out_T3D5.txt
In [18]:
samp="M1"
In [19]:
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_{samp}.txt> methratio_out_{samp}_CG.txt
#53x coverage
!awk '{if ($8 >= 3) print $1,"methratio","CpG",$2,$2+1,$5,".","."}' <methratio_out_{samp}_CG.txt> filt_methratio_out_{samp}_CG.gff
!tr ' ' "\t" <filt_methratio_out_{samp}_CG.gff> filt3_{samp}.gff
/bin/sh: methratio_out_{samp}_CG.txt: No such file or directory
/bin/sh: filt_methratio_out_M1_CG.gff: No such file or directory
In [21]:
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_M1.txt> methratio_out_M1_CG.txt
#3x coverage
!awk '{if ($8 >= 3) print $1,"methratio","CpG",$2,$2+1,$5,".","."}' <methratio_out_M1_CG.txt> filt_methratio_out_M1_CG.gff
!tr ' ' "\t" <filt_methratio_out_M1_CG.gff> filt3_M1.gff
In [22]:
!head filt3_M1.gff
C12768 methratio CpG 103 104 0.167 . . C12768 methratio CpG 119 120 0.250 . . C12768 methratio CpG 145 146 0.000 . . C12806 methratio CpG 56 57 0.000 . . C12806 methratio CpG 76 77 0.200 . . C12806 methratio CpG 78 79 0.200 . . C12806 methratio CpG 105 106 0.250 . . C12806 methratio CpG 142 143 0.375 . . C12924 methratio CpG 19 20 0.000 . . C12924 methratio CpG 30 31 0.000 . .
In [23]:
#make igv
!awk '{if ($8 >= 3) print $1,$2-1,$2+1,"CpG",$5}' <methratio_out_M1_CG.txt> filt_methratio_out_M1_CG.igv
!tr ' ' "\t" <filt_methratio_out_M1_CG.igv> filt3_M1.igv
In [24]:
#make igv
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_M3.txt> methratio_out_M3_CG.txt
!awk '{if ($8 >= 3) print $1,$2-1,$2+1,"CpG",$5}' <methratio_out_M3_CG.txt> filt_methratio_out_M3_CG.igv
!tr ' ' "\t" <filt_methratio_out_M3_CG.igv> filt3_M3.igv
In [42]:
#make igv with join column
!awk '{if ($8 >= 3) print $1"_"$2-1,$1,$2-1,$2+1,"CpG",$5,$8}' <methratio_out_M3_CG.txt> filt_methratio_out_M3_CGj.txt
!tr ' ' "\t" <filt_methratio_out_M3_CGj.txt> filt3_M3j.txt
In [41]:
#make igv with join column
!awk '{if ($8 >= 3) print $1"_"$2-1,$1,$2-1,$2+1,"CpG",$5,$8}' <methratio_out_M1_CG.txt> filt_methratio_out_M1_CGj.txt
!tr ' ' "\t" <filt_methratio_out_M1_CGj.txt> filt3_M1j.txt
In [39]:
!head filt3_M1j.txt
C12768_102 C12768 102 104 CpG 0.167 C12768_118 C12768 118 120 CpG 0.250 C12768_144 C12768 144 146 CpG 0.000 C12806_55 C12806 55 57 CpG 0.000 C12806_75 C12806 75 77 CpG 0.200 C12806_77 C12806 77 79 CpG 0.200 C12806_104 C12806 104 106 CpG 0.250 C12806_141 C12806 141 143 CpG 0.375 C12924_18 C12924 18 20 CpG 0.000 C12924_29 C12924 29 31 CpG 0.000
In [77]:
!echo "M3ID M3scaffold M3start M3end M3feat M3ratio M3coverage" >> m3head
In [78]:
!head m1head
M1ID M1scaffold M1start M1end M1feat M1ratio M1coverage
In [69]:
ls
filt3_M1.gff hdfilt_methratio_out_M1_CGj.txt filt3_M1.igv m1head filt3_M1j.txt methratio_out_M1.txt filt3_M3.igv methratio_out_M1_CG.txt filt3_M3j.igv methratio_out_M3.txt filt3_M3j.txt methratio_out_M3_CG.txt filt_methratio_out_M1_CG.gff methratio_out_T1D3.txt filt_methratio_out_M1_CG.igv methratio_out_T1D5.txt filt_methratio_out_M1_CGj.txt methratio_out_T3D3.txt filt_methratio_out_M3_CG.igv methratio_out_T3D5.txt filt_methratio_out_M3_CGj.igv test filt_methratio_out_M3_CGj.txt test2
In [79]:
!cat m3head filt_methratio_out_M3_CGj.txt > hdfilt_methratio_out_M3_CGj.txt
In [82]:
!head hdfilt_methratio_out_M3_CGj.txt
M3ID M3scaffold M3start M3end M3feat M3ratio M3coverage C12806_141 C12806 141 143 CpG 0.000 4 C12866_161 C12866 161 163 CpG 0.000 3 C12960_122 C12960 122 124 CpG 0.000 3 C12994_133 C12994 133 135 CpG 0.000 3 C13046_75 C13046 75 77 CpG 0.000 3 C13074_145 C13074 145 147 CpG 0.000 3 C13080_101 C13080 101 103 CpG 0.000 3 C13080_110 C13080 110 112 CpG 0.000 3 C13080_129 C13080 129 131 CpG 0.000 3
In [75]:
!tr ' ' "\t" <hdfilt_methratio_out_M1_CGj.txt> filt3_M1j.txt
In [84]:
!tr ' ' "\t" <hdfilt_methratio_out_M3_CGj.txt> filt3_M3j.txt
In [85]:
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d filt3_M3 filt3_M3j.txt
processing chunk line 0 to 1364437 (0.66103887558 s elapsed) pushing filt3_M3j.txt... parsing B0EB1C71... finished filt3_M3
In [76]:
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d filt3_M1 filt3_M1j.txt
processing chunk line 0 to 1218606 (0.596918106079 s elapsed) pushing filt3_M1j.txt... parsing B993D808... finished filt3_M1
Clean convert¶
In [130]:
smp="M1"
#Need to manually change id in awk line.
In [132]:
#cg only
#!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_{smp}.txt> methratio_out_{smp}_CG.txt
#!sed '/NA/d' methratio_out_{smp}_CG.txt > methratio_out_{smp}_CGna.txt
#make join column, 3x etc
!awk '{if ($8 >= 3) print $1"_"$2,$1,$2,$2+1,"CpG",$5,$8}' <methratio_out_M1_CGna.txt> filt_methratio_out_M1_CGj.txt
#print header
!echo "{smp}ID {smp}scaffold {smp}start {smp}end {smp}feat {smp}ratio {smp}coverage" >> {smp}head
#add header
!cat {smp}head filt_methratio_out_{smp}_CGj.txt > hdfilt_methratio_out_{smp}_CGj.txt
!tr ' ' "\t" <hdfilt_methratio_out_{smp}_CGj.txt> filt3_{smp}j.txt
#upload to sqlshare
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d filt3_{smp} filt3_{smp}j.txt
processing chunk line 0 to 1212751 (0.757140159607 s elapsed) pushing filt3_M1j.txt... parsing 57755226... finished filt3_M1
In [133]:
smp="M3"
In [134]:
#cg only
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_{smp}.txt> methratio_out_{smp}_CG.txt
!sed '/NA/d' methratio_out_{smp}_CG.txt > methratio_out_{smp}_CGna.txt
#make join column, 3x etc
!awk '{if ($8 >= 3) print $1"_"$2,$1,$2,$2+1,"CpG",$5,$8}' <methratio_out_M3_CGna.txt> filt_methratio_out_M3_CGj.txt
#print header
!echo "{smp}ID {smp}scaffold {smp}start {smp}end {smp}feat {smp}ratio {smp}coverage" >> {smp}head
#add header
!cat {smp}head filt_methratio_out_{smp}_CGj.txt > hdfilt_methratio_out_{smp}_CGj.txt
!tr ' ' "\t" <hdfilt_methratio_out_{smp}_CGj.txt> filt3_{smp}j.txt
#upload to sqlshare
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d filt3_{smp} filt3_{smp}j.txt
processing chunk line 0 to 1360752 (0.855348110199 s elapsed) pushing filt3_M3j.txt... parsing 831E6928... finished filt3_M3
In [135]:
smp="T1D3"
In [136]:
#cg only
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_{smp}.txt> methratio_out_{smp}_CG.txt
!sed '/NA/d' methratio_out_{smp}_CG.txt > methratio_out_{smp}_CGna.txt
#make join column, 3x etc
!awk '{if ($8 >= 3) print $1"_"$2,$1,$2,$2+1,"CpG",$5,$8}' <methratio_out_T1D3_CGna.txt> filt_methratio_out_T1D3_CGj.txt
#print header
!echo "{smp}ID {smp}scaffold {smp}start {smp}end {smp}feat {smp}ratio {smp}coverage" >> {smp}head
#add header
!cat {smp}head filt_methratio_out_{smp}_CGj.txt > hdfilt_methratio_out_{smp}_CGj.txt
!tr ' ' "\t" <hdfilt_methratio_out_{smp}_CGj.txt> filt3_{smp}j.txt
#upload to sqlshare
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d filt3_{smp} filt3_{smp}j.txt
processing chunk line 0 to 251459 (0.165581941605 s elapsed) pushing filt3_T1D3j.txt... parsing 582E2A55... finished filt3_T1D3
In [137]:
smp="T1D5"
In [138]:
#cg only
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_{smp}.txt> methratio_out_{smp}_CG.txt
!sed '/NA/d' methratio_out_{smp}_CG.txt > methratio_out_{smp}_CGna.txt
#make join column, 3x etc
!awk '{if ($8 >= 3) print $1"_"$2,$1,$2,$2+1,"CpG",$5,$8}' <methratio_out_T1D5_CGna.txt> filt_methratio_out_T1D5_CGj.txt
#print header
!echo "{smp}ID {smp}scaffold {smp}start {smp}end {smp}feat {smp}ratio {smp}coverage" >> {smp}head
#add header
!cat {smp}head filt_methratio_out_{smp}_CGj.txt > hdfilt_methratio_out_{smp}_CGj.txt
!tr ' ' "\t" <hdfilt_methratio_out_{smp}_CGj.txt> filt3_{smp}j.txt
#upload to sqlshare
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d filt3_{smp} filt3_{smp}j.txt
processing chunk line 0 to 873151 (0.512318849564 s elapsed) pushing filt3_T1D5j.txt... parsing C5B90BB9... finished filt3_T1D5
In [139]:
smp="T3D3"
In [140]:
#cg only
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_{smp}.txt> methratio_out_{smp}_CG.txt
!sed '/NA/d' methratio_out_{smp}_CG.txt > methratio_out_{smp}_CGna.txt
#make join column, 3x etc
!awk '{if ($8 >= 3) print $1"_"$2,$1,$2,$2+1,"CpG",$5,$8}' <methratio_out_T3D3_CGna.txt> filt_methratio_out_T3D3_CGj.txt
#print header
!echo "{smp}ID {smp}scaffold {smp}start {smp}end {smp}feat {smp}ratio {smp}coverage" >> {smp}head
#add header
!cat {smp}head filt_methratio_out_{smp}_CGj.txt > hdfilt_methratio_out_{smp}_CGj.txt
!tr ' ' "\t" <hdfilt_methratio_out_{smp}_CGj.txt> filt3_{smp}j.txt
#upload to sqlshare
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d filt3_{smp} filt3_{smp}j.txt
processing chunk line 0 to 1157485 (1.06684994698 s elapsed) pushing filt3_T3D3j.txt... parsing 927E672B... finished filt3_T3D3
In [141]:
smp="T3D5"
In [142]:
#cg only
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_{smp}.txt> methratio_out_{smp}_CG.txt
!sed '/NA/d' methratio_out_{smp}_CG.txt > methratio_out_{smp}_CGna.txt
#make join column, 3x etc
!awk '{if ($8 >= 3) print $1"_"$2,$1,$2,$2+1,"CpG",$5,$8}' <methratio_out_T3D5_CGna.txt> filt_methratio_out_T3D5_CGj.txt
#print header
!echo "{smp}ID {smp}scaffold {smp}start {smp}end {smp}feat {smp}ratio {smp}coverage" >> {smp}head
#add header
!cat {smp}head filt_methratio_out_{smp}_CGj.txt > hdfilt_methratio_out_{smp}_CGj.txt
!tr ' ' "\t" <hdfilt_methratio_out_{smp}_CGj.txt> filt3_{smp}j.txt
#upload to sqlshare
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d filt3_{smp} filt3_{smp}j.txt
processing chunk line 0 to 1183434 (1.03248810768 s elapsed) pushing filt3_T3D5j.txt... parsing 0165A793... finished filt3_T3D5
Using SQLShare to bin CpG in M1 / M3 comparison¶
In [ ]:
SELECT * FROM [sr320@washington.edu].[filt3_M1]m1
join
[sr320@washington.edu].[filt3_M3]m3
on
m1.M1ID=m3.M3ID
left join
[sr320@washington.edu].[CG_Ensembl_featurejoin]cg
on
m1.M1ID=cg.ChrStart
In [ ]:
#confirmed will work in theory? meaning joined to feature
In [ ]:
SELECT * FROM [sr320@washington.edu].[filt3M1_M3_CGfeature]
where M1ratio > 0.5
and
M3ratio > 0.5
In [ ]:
!python /Users/sr320/sqlshare-pythonclient/tools/fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[filt3M1_M3_CGfeature] where M1ratio > 0.5 and M3ratio > 0.5" -f tsv -o /Volumes/web/cnidarian/M1_M3_05.txt
In [2]:
!head /Volumes/web/cnidarian/M1_M3_05.txt
In [3]:
!wc /Volumes/web/cnidarian/M1_M3_05.txt
0 0 0 /Volumes/web/cnidarian/M1_M3_05.txt
In [ ]:
#above was done with join on feature tracks
#lets do with just simple Joins on samples
In [5]:
!head /Volumes/web/cnidarian/M1_M3_05_simple.txt
In [6]:
#lets do heatmaps
#need to get Methylation levels for all 6 samples (common loci)
Towards Hierarchical Clustering and Heatmaps¶
Step1: Getting table with information from all 6 samples
SELECT * FROM [sr320@washington.edu].[filt3_M1]m1
join
[sr320@washington.edu].[filt3_M3]m3
on
m1.M1ID=m3.M3ID
join
[sr320@washington.edu].[filt3_T1D3]t1d3
on
m1.M1ID=t1d3.T1D3ID
join
[sr320@washington.edu].[filt3_T1D5]t1d5
on
m1.M1ID=t1d5.T1D5ID
join
[sr320@washington.edu].[filt3_T3D3]t3d3
on
m1.M1ID=t3d3.T3D3ID
join
[sr320@washington.edu].[filt3_T3D5]t3d5
on
m1.M1ID=t3d5.T3D5ID
SELECT M1ID,M1ratio,M3ratio,T1D3ratio,T1D5ratio,T3D3ratio,T3D5ratio
FROM [sr320@washington.edu].[filt3_M1]m1
join
[sr320@washington.edu].[filt3_M3]m3
on
m1.M1ID=m3.M3ID
join
[sr320@washington.edu].[filt3_T1D3]t1d3
on
m1.M1ID=t1d3.T1D3ID
join
[sr320@washington.edu].[filt3_T1D5]t1d5
on
m1.M1ID=t1d5.T1D5ID
join
[sr320@washington.edu].[filt3_T3D3]t3d3
on
m1.M1ID=t3d3.T3D3ID
join
[sr320@washington.edu].[filt3_T3D5]t3d5
on
m1.M1ID=t3d5.T3D5ID
In [9]:
!python /Users/sr320/sqlshare-pythonclient/tools/fetchdata.py -s "SELECT M1ID,M1ratio,M3ratio,T1D3ratio,T1D5ratio,T3D3ratio,T3D5ratio FROM [sr320@washington.edu].[filt3_M1]m1 join [sr320@washington.edu].[filt3_M3]m3 on m1.M1ID=m3.M3ID join [sr320@washington.edu].[filt3_T1D3]t1d3 on m1.M1ID=t1d3.T1D3ID join [sr320@washington.edu].[filt3_T1D5]t1d5 on m1.M1ID=t1d5.T1D5ID join [sr320@washington.edu].[filt3_T3D3]t3d3 on m1.M1ID=t3d3.T3D3ID join [sr320@washington.edu].[filt3_T3D5]t3d5 on m1.M1ID=t3d5.T3D5ID" -f tsv -o /Volumes/web/cnidarian/filt3_BiGoLarvae_common.txt
In [33]:
!python /Users/sr320/sqlshare-pythonclient/tools/fetchdata.py -s "SELECT M1ratio,M3ratio,T1D3ratio,T1D5ratio,T3D3ratio,T3D5ratio FROM [sr320@washington.edu].[filt3_M1]m1 join [sr320@washington.edu].[filt3_M3]m3 on m1.M1ID=m3.M3ID join [sr320@washington.edu].[filt3_T1D3]t1d3 on m1.M1ID=t1d3.T1D3ID join [sr320@washington.edu].[filt3_T1D5]t1d5 on m1.M1ID=t1d5.T1D5ID join [sr320@washington.edu].[filt3_T3D3]t3d3 on m1.M1ID=t3d3.T3D3ID join [sr320@washington.edu].[filt3_T3D5]t3d5 on m1.M1ID=t3d5.T3D5ID" -f tsv -o /Volumes/web/cnidarian/filt3_BiGoLarvae_common2.txt
In [34]:
!head /Volumes/web/cnidarian/filt3_BiGoLarvae_common2.txt
In [1]:
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import scipy.spatial.distance as distance
import scipy.cluster.hierarchy as sch
test data¶
In [6]:
testDF.shape
--------------------------------------------------------------------------- NameError Traceback (most recent call last) <ipython-input-6-cddb38ed45ab> in <module>() ----> 1 testDF.shape NameError: name 'testDF' is not defined
In [7]:
print(testDF)
--------------------------------------------------------------------------- NameError Traceback (most recent call last) <ipython-input-7-38028a8e5e97> in <module>() ----> 1 print(testDF) NameError: name 'testDF' is not defined
In [2]:
from pandas import *
# read data from data file into a pandas DataFrame
larDB = read_table("http://eagle.fish.washington.edu/cnidarian/filt3_BiGoLarvae_common3.txt", # name of the data file
#sep=",", # what character separates each column?
na_values=["", " "]) # what values should be considered "blank" values?
In [3]:
print(larDB)
<class 'pandas.core.frame.DataFrame'> Int64Index: 5129 entries, 0 to 5128 Data columns (total 6 columns): M1ratio 5128 non-null values M3ratio 5128 non-null values T1D3ratio 5128 non-null values T1D5ratio 5128 non-null values T3D3ratio 5128 non-null values T3D5ratio 5128 non-null values dtypes: float64(6)
In [4]:
# helper for cleaning up axes by removing ticks, tick labels, frame, etc.
def clean_axis(ax):
"""Remove ticks, tick labels, and frame from axis"""
ax.get_xaxis().set_ticks([])
ax.get_yaxis().set_ticks([])
for sp in ax.spines.values():
sp.set_visible(False)
In [5]:
# look at raw data
axi = imshow(larDB,interpolation='nearest',cmap=cm.RdBu)
ax = axi.get_axes()
clean_axis(ax)
In [6]:
# calculate pairwise distances for rows
pairwise_dists = distance.squareform(distance.pdist(larDB))
print 'Number of rows: {0}'.format(larDB.shape[0])
print 'Size of distance matrix: {0}'.format(pairwise_dists.shape)
Number of rows: 5129 Size of distance matrix: (5129, 5129)
In [7]:
# cluster
clusters = sch.linkage(pairwise_dists,method='complete')
In [8]:
# dendrogram
den = sch.dendrogram(clusters)
In [11]:
axi = imshow(larDB.ix[den['leaves']],interpolation='nearest',cmap=cm.RdBu)
ax = axi.get_axes()
clean_axis(ax)
Moving up to 5x coverage - CLC¶
In [ ]:
!python /Users/sr320/sqlshare-pythonclient/tools/fetchdata.py -s "SELECT M1ID,M1ratio,M3ratio,T1D3ratio,T1D5ratio,T3D3ratio,T3D5ratio FROM [sr320@washington.edu].[filt3_M1]m1 join [sr320@washington.edu].[filt3_M3]m3 on m1.M1ID=m3.M3ID join [sr320@washington.edu].[filt3_T1D3]t1d3 on m1.M1ID=t1d3.T1D3ID join [sr320@washington.edu].[filt3_T1D5]t1d5 on m1.M1ID=t1d5.T1D5ID join [sr320@washington.edu].[filt3_T3D3]t3d3 on m1.M1ID=t3d3.T3D3ID join [sr320@washington.edu].[filt3_T3D5]t3d5 on m1.M1ID=t3d5.T3D5ID where [M1coverage] >= '5' and [M3coverage] >= '5' and [T1D3coverage] >= '5' and [T1D5coverage] >= '5' and [T3D3coverage] >= '5' and [T3D5coverage] >= '5'" -f tsv -o /Volumes/web/cnidarian/filt5_BiGoLarvae_common.txt
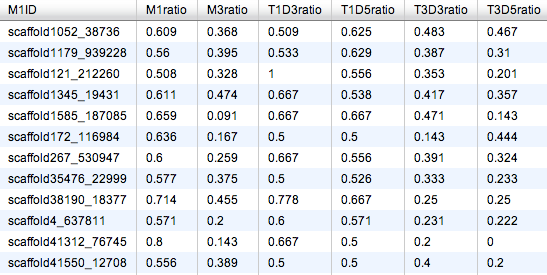
Revisiting specific queries using SQLShare¶
SELECT M1ID,M1ratio,M3ratio,T1D3ratio,T1D5ratio,T3D3ratio,T3D5ratio
FROM [sr320@washington.edu].[filt3_M1]m1
join
[sr320@washington.edu].[filt3_M3]m3
on
m1.M1ID=m3.M3ID
join
[sr320@washington.edu].[filt3_T1D3]t1d3
on
m1.M1ID=t1d3.T1D3ID
join
[sr320@washington.edu].[filt3_T1D5]t1d5
on
m1.M1ID=t1d5.T1D5ID
join
[sr320@washington.edu].[filt3_T3D3]t3d3
on
m1.M1ID=t3d3.T3D3ID
join
[sr320@washington.edu].[filt3_T3D5]t3d5
on
m1.M1ID=t3d5.T3D5ID
where
[M1coverage] >= '5'
and
[M3coverage] >= '5'
and
[T1D3coverage] >= '5'
and
[T1D5coverage] >= '5'
and
[T3D3coverage] >= '5'
and
[T3D5coverage] >= '5'
and
M1ratio >= 0.5
and
T1D3ratio >= 0.5
and
T1D5ratio >= 0.5
and
M3ratio < 0.5
and
T3D3ratio < 0.5
and
T3D5ratio < 0.5
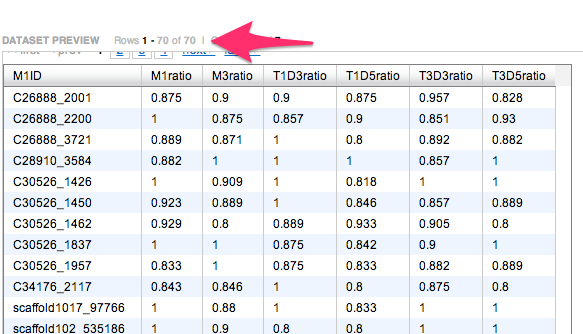
SELECT M1ID,M1ratio,M3ratio,T1D3ratio,T1D5ratio,T3D3ratio,T3D5ratio
FROM [sr320@washington.edu].[filt3_M1]m1
join
[sr320@washington.edu].[filt3_M3]m3
on
m1.M1ID=m3.M3ID
join
[sr320@washington.edu].[filt3_T1D3]t1d3
on
m1.M1ID=t1d3.T1D3ID
join
[sr320@washington.edu].[filt3_T1D5]t1d5
on
m1.M1ID=t1d5.T1D5ID
join
[sr320@washington.edu].[filt3_T3D3]t3d3
on
m1.M1ID=t3d3.T3D3ID
join
[sr320@washington.edu].[filt3_T3D5]t3d5
on
m1.M1ID=t3d5.T3D5ID
where
[M1coverage] >= '5'
and
[M3coverage] >= '5'
and
[T1D3coverage] >= '5'
and
[T1D5coverage] >= '5'
and
[T3D3coverage] >= '5'
and
[T3D5coverage] >= '5'
and
M1ratio >= 0.8
and
T1D3ratio >= 0.8
and
T1D5ratio >= 0.8
and
M3ratio >= 0.8
and
T3D3ratio >= 0.8
and
T3D5ratio >= 0.8
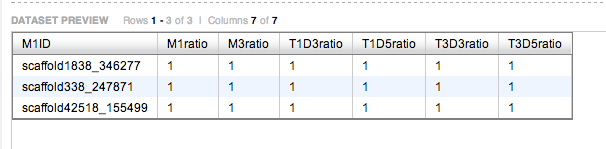
SELECT M1ID,M1ratio,M3ratio,T1D3ratio,T1D5ratio,T3D3ratio,T3D5ratio
FROM [sr320@washington.edu].[filt3_M1]m1
join
[sr320@washington.edu].[filt3_M3]m3
on
m1.M1ID=m3.M3ID
join
[sr320@washington.edu].[filt3_T1D3]t1d3
on
m1.M1ID=t1d3.T1D3ID
join
[sr320@washington.edu].[filt3_T1D5]t1d5
on
m1.M1ID=t1d5.T1D5ID
join
[sr320@washington.edu].[filt3_T3D3]t3d3
on
m1.M1ID=t3d3.T3D3ID
join
[sr320@washington.edu].[filt3_T3D5]t3d5
on
m1.M1ID=t3d5.T3D5ID
where
[M1coverage] >= '5'
and
[M3coverage] >= '5'
and
[T1D3coverage] >= '5'
and
[T1D5coverage] >= '5'
and
[T3D3coverage] >= '5'
and
[T3D5coverage] >= '5'
and
M1ratio = 0
and
T1D3ratio = 0
and
T1D5ratio = 0
and
M3ratio = 0
and
T3D3ratio = 0
and
T3D5ratio = 0
In [2]:
!python /Users/sr320/sqlshare-pythonclient/tools/fetchdata.py -s "SELECT M1ID,M1ratio,M3ratio,T1D3ratio,T1D5ratio,T3D3ratio,T3D5ratio FROM [sr320@washington.edu].[filt3_M1]m1 join [sr320@washington.edu].[filt3_M3]m3 on m1.M1ID=m3.M3ID join [sr320@washington.edu].[filt3_T1D3]t1d3 on m1.M1ID=t1d3.T1D3ID join [sr320@washington.edu].[filt3_T1D5]t1d5 on m1.M1ID=t1d5.T1D5ID join [sr320@washington.edu].[filt3_T3D3]t3d3 on m1.M1ID=t3d3.T3D3ID join [sr320@washington.edu].[filt3_T3D5]t3d5 on m1.M1ID=t3d5.T3D5ID where [M1coverage] >= '5' and [M3coverage] >= '5' and [T1D3coverage] >= '5' and [T1D5coverage] >= '5' and [T3D3coverage] >= '5' and [T3D5coverage] >= '5' and M1ratio = 0 and T1D3ratio = 0 and T1D5ratio = 0 and M3ratio = 0 and T3D3ratio = 0 and T3D5ratio = 0" -f tsv -o /Volumes/web/cnidarian/filt5_BiGoLarvae_allzero.txt
In [3]:
!head /Volumes/web/cnidarian/filt5_BiGoLarvae_allzero.txt
In [4]:
!wc /Volumes/web/cnidarian/filt5_BiGoLarvae_allzero.txt
5053 35371 158327 /Volumes/web/cnidarian/filt5_BiGoLarvae_allzero.txt
In [5]:
!wc /Volumes/web/cnidarian/filt5_BiGoLarvae_common.txt
11210 78470 427112 /Volumes/web/cnidarian/filt5_BiGoLarvae_common.txt
SELECT M1ID,M1ratio,T1D3ratio,T1D5ratio,((m1ratio + T1D3ratio + T1D5ratio)/3) as mean_1lin,
(Select stdev(v) from (values (M1ratio), (T1D3ratio), (T1D5ratio)) as value(v)) as [stdev],
(Select stdev(v) from (values (M1ratio), (T1D3ratio), (T1D5ratio)) as value(v)) / ((m1ratio + T1D3ratio + T1D5ratio)/3) as CI
FROM [sr320@washington.edu].[filt3_M1]m1
join
[sr320@washington.edu].[filt3_M3]m3
on
m1.M1ID=m3.M3ID
join
[sr320@washington.edu].[filt3_T1D3]t1d3
on
m1.M1ID=t1d3.T1D3ID
join
[sr320@washington.edu].[filt3_T1D5]t1d5
on
m1.M1ID=t1d5.T1D5ID
join
[sr320@washington.edu].[filt3_T3D3]t3d3
on
m1.M1ID=t3d3.T3D3ID
join
[sr320@washington.edu].[filt3_T3D5]t3d5
on
m1.M1ID=t3d5.T3D5ID
where
[M1coverage] >= '5'
and
[M3coverage] >= '5'
and
[T1D3coverage] >= '5'
and
[T1D5coverage] >= '5'
and
[T3D3coverage] >= '5'
and
[T3D5coverage] >= '5'
and
M1ratio > 0
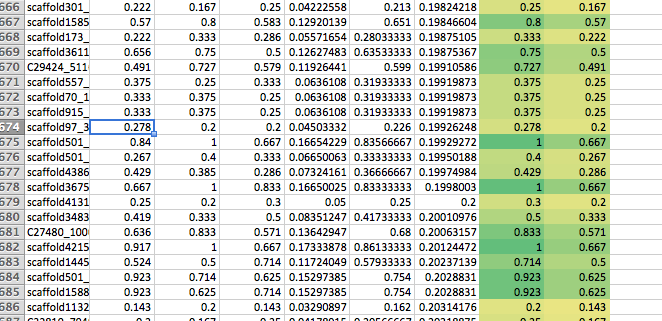
In [ ]:
#lets consider taking 674 1 lineaage with cv <0.2 and get info. this is on 11,000 that match across 6
Creating IGV files¶
In [1]:
cd /Volumes/web/cnidarian/BiGo_larvae_3/
/Volumes/web/cnidarian/BiGo_larvae_3
In [2]:
mkdir igv
In [3]:
cd igv
/Volumes/web/cnidarian/BiGo_larvae_3/igv
In [16]:
sp="M3"
In [14]:
#command for only obtaining the context '__CG_'
#!grep "[A-Z][A-Z]CG[A-Z]" </Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_{sp}.txt> {sp}_methratio_CG.txt
#5x coverage
!awk '{if ($8 >= 5) print $1,$2-1,$2+1,"CpG",$5}' <M1_methratio_CG.txt> M1_methratio_CG5x.igv
!tr ' ' "\t" <M1_methratio_CG5x.igv> M1_CG5x.igv
In [15]:
!head M1_CG5x.igv
C12768 102 104 CpG 0.167 C12806 75 77 CpG 0.200 C12806 77 79 CpG 0.200 C12806 141 143 CpG 0.375 C12924 29 31 CpG 0.000 C12924 37 39 CpG 0.000 C12924 51 53 CpG 0.000 C12924 59 61 CpG 0.000 C12924 126 128 CpG 0.000 C12924 135 137 CpG 0.000
In [17]:
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" </Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_{sp}.txt> {sp}_methratio_CG.txt
In [18]:
sp="T1D3"
In [19]:
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" </Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_{sp}.txt> {sp}_methratio_CG.txt
In [20]:
sp="T1D5"
In [21]:
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" </Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_{sp}.txt> {sp}_methratio_CG.txt
grep: (standard input): Input/output error
In [22]:
sp="M3"
In [23]:
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" </Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_{sp}.txt> {sp}_methratio_CG.txt
shell-init: error retrieving current directory: getcwd: cannot access parent directories: No such file or directory /bin/sh: /Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_M3.txt: No such file or directory
In [24]:
sp="T3D3"
In [25]:
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" </Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_{sp}.txt> {sp}_methratio_CG.txt
shell-init: error retrieving current directory: getcwd: cannot access parent directories: No such file or directory /bin/sh: /Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_T3D3.txt: No such file or directory
In [26]:
sp="T3D5"
In [27]:
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" </Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_{sp}.txt> {sp}_methratio_CG.txt
shell-init: error retrieving current directory: getcwd: cannot access parent directories: No such file or directory /bin/sh: /Volumes/web/cnidarian/BiGo_larvae_3/methratio_out_T3D5.txt: No such file or directory
In [ ]: