Bisulfite Sequencing - Larvae 5 day Trial¶
In [16]:
#SR edits - html embeds not working on html so generatinv nbviewer linke
!date
Fri Feb 21 07:39:49 PST 2014
In [4]:
filename='BiGo_larvae_2'
In [5]:
#ipynb viewer format:
!echo 'http://nbviewer.ipython.org/github/sr320/ipython_nb/blob/master/''{filename}''.ipynb'
http://nbviewer.ipython.org/github/sr320/ipython_nb/blob/master/BiGo_larvae_2.ipynb
In [6]:
#ipynb file:
!echo 'https://github.com/sr320/ipython_nb/blob/master/''{filename}''.ipynb'
https://github.com/sr320/ipython_nb/blob/master/BiGo_larvae_2.ipynb
TOC¶
In [ ]:
In [ ]:
Summary: 7 libraries were made. Egg, Sperm 1, Larvae1_day3, Larvae1_day5, Sperm 3, Larvae3_day3, Larvae3_day5. Egg sample has consistenly failed at sequencing, but have two sets of sequencing for everything else. The first step will be to compile libraries. MAYBE? check and see if data is different between technical reps.
In [2]:
! /Volumes/Bay3/Software/BSMAP/bsmap-2.74/bsmap -a /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/FCC3ATL/BS_CgF_ACAGTG_L004_R1_001.fastq.gz -d /Volumes/web/cnidarian/oyster.v9.fa -o /Volumes/web/cnidarian/BiGo_egg_redo.sam -p 4
BSMAP v2.74 Start at: Wed Jan 22 18:16:14 2014 Input reference file: /Volumes/web/cnidarian/oyster.v9.fa (format: FASTA) Load in 11969 db seqs, total size 558601156 bp. 26 secs passed total_kmers: 43046721 Create seed table. 97 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand: ++,-+ Single-end alignment(4 threads) Input read file: /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/FCC3ATL/BS_CgF_ACAGTG_L004_R1_001.fastq.gz (format: gzipped FASTQ) Output file: /Volumes/web/cnidarian/BiGo_egg_redo.sam (format: SAM) Thread #1: 150000 reads finished. 126 secs passed Thread #2: 100000 reads finished. 126 secs passed Thread #3: 200000 reads finished. 126 secs passed Thread #0: 50000 reads finished. 126 secs passed Thread #1: 250000 reads finished. 162 secs passed Thread #3: 350000 reads finished. 162 secs passed Thread #2: 300000 reads finished. 163 secs passed Thread #0: 400000 reads finished. 164 secs passed Thread #1: 450000 reads finished. 193 secs passed Thread #3: 500000 reads finished. 193 secs passed Thread #2: 550000 reads finished. 193 secs passed Thread #0: 600000 reads finished. 194 secs passed Thread #1: 650000 reads finished. 228 secs passed Thread #3: 700000 reads finished. 229 secs passed Thread #2: 750000 reads finished. 229 secs passed Thread #0: 800000 reads finished. 230 secs passed Thread #1: 850000 reads finished. 260 secs passed Thread #3: 900000 reads finished. 261 secs passed Thread #2: 950000 reads finished. 262 secs passed Thread #0: 1000000 reads finished. 262 secs passed Thread #1: 1050000 reads finished. 290 secs passed Thread #3: 1100000 reads finished. 291 secs passed Thread #2: 1150000 reads finished. 292 secs passed Thread #0: 1200000 reads finished. 293 secs passed Thread #1: 1250000 reads finished. 317 secs passed Thread #3: 1300000 reads finished. 319 secs passed Thread #2: 1350000 reads finished. 320 secs passed Thread #0: 1400000 reads finished. 320 secs passed Thread #1: 1450000 reads finished. 348 secs passed Thread #3: 1500000 reads finished. 349 secs passed Thread #2: 1550000 reads finished. 350 secs passed Thread #0: 1600000 reads finished. 350 secs passed Thread #1: 1650000 reads finished. 380 secs passed Thread #3: 1700000 reads finished. 382 secs passed Thread #2: 1750000 reads finished. 383 secs passed Thread #0: 1800000 reads finished. 384 secs passed Thread #1: 1850000 reads finished. 407 secs passed Thread #3: 1900000 reads finished. 408 secs passed Thread #0: 2000000 reads finished. 409 secs passed Thread #2: 1950000 reads finished. 409 secs passed Thread #2: 2151969 reads finished. 411 secs passed Thread #1: 2050000 reads finished. 431 secs passed Thread #3: 2100000 reads finished. 432 secs passed Thread #0: 2150000 reads finished. 432 secs passed Total number of aligned reads: 593117 (28%) Done. Finished at Wed Jan 22 18:23:27 2014 Total time consumed: 433 secs
In [2]:
#FEB 13 redux looking at R1 AND R2
! /Volumes/Bay3/Software/BSMAP/bsmap-2.74/bsmap -a /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC3ATL/BS_CgF_ACAGTG_L004_R1_001.fastq.gz -b /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC3ATL/BS_CgF_ACAGTG_L004_R2_001.fastq.gz -d /Volumes/web/cnidarian/oyster.v9.fa -o /Volumes/web/cnidarian/BiGo_egg_redo4.sam -p 4
BSMAP v2.74 Start at: Thu Feb 13 17:26:33 2014 Input reference file: /Volumes/web/cnidarian/oyster.v9.fa (format: FASTA) Load in 11969 db seqs, total size 558601156 bp. 17 secs passed total_kmers: 43046721 Create seed table. 91 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC3ATL/BS_CgF_ACAGTG_L004_R1_001.fastq.gz (format: gzipped FASTQ) Input read file #2: /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC3ATL/BS_CgF_ACAGTG_L004_R2_001.fastq.gz (format: gzipped FASTQ) Output file: /Volumes/web/cnidarian/BiGo_egg_redo4.sam (format: SAM) Thread #3: 50000 read pairs finished. 229 secs passed Thread #1: 100000 read pairs finished. 232 secs passed Thread #2: 150000 read pairs finished. 235 secs passed Thread #0: 200000 read pairs finished. 239 secs passed Thread #3: 250000 read pairs finished. 354 secs passed Thread #0: 400000 read pairs finished. 356 secs passed Thread #1: 300000 read pairs finished. 359 secs passed Thread #2: 350000 read pairs finished. 364 secs passed Thread #3: 450000 read pairs finished. 473 secs passed Thread #0: 500000 read pairs finished. 477 secs passed Thread #1: 550000 read pairs finished. 481 secs passed Thread #2: 600000 read pairs finished. 489 secs passed Thread #1: 750000 read pairs finished. 582 secs passed Thread #3: 650000 read pairs finished. 583 secs passed Thread #0: 700000 read pairs finished. 590 secs passed Thread #2: 800000 read pairs finished. 590 secs passed Thread #1: 850000 read pairs finished. 682 secs passed Thread #3: 900000 read pairs finished. 688 secs passed Thread #0: 950000 read pairs finished. 697 secs passed Thread #2: 1000000 read pairs finished. 700 secs passed Thread #3: 1100000 read pairs finished. 791 secs passed Thread #1: 1050000 read pairs finished. 795 secs passed Thread #0: 1150000 read pairs finished. 795 secs passed Thread #2: 1200000 read pairs finished. 801 secs passed Thread #3: 1250000 read pairs finished. 896 secs passed Thread #1: 1300000 read pairs finished. 903 secs passed Thread #0: 1350000 read pairs finished. 907 secs passed Thread #2: 1400000 read pairs finished. 914 secs passed Thread #1: 1500000 read pairs finished. 1004 secs passed Thread #3: 1450000 read pairs finished. 1007 secs passed Thread #0: 1550000 read pairs finished. 1008 secs passed Thread #2: 1600000 read pairs finished. 1018 secs passed Thread #1: 1650000 read pairs finished. 1109 secs passed Thread #3: 1700000 read pairs finished. 1114 secs passed Thread #0: 1750000 read pairs finished. 1117 secs passed Thread #2: 1800000 read pairs finished. 1126 secs passed Thread #1: 1850000 read pairs finished. 1205 secs passed Thread #3: 1900000 read pairs finished. 1210 secs passed Thread #0: 1950000 read pairs finished. 1215 secs passed Thread #2: 2000000 read pairs finished. 1228 secs passed Thread #2: 2151969 read pairs finished. 1232 secs passed Thread #1: 2050000 read pairs finished. 1280 secs passed Thread #3: 2100000 read pairs finished. 1283 secs passed Thread #0: 2150000 read pairs finished. 1285 secs passed Total number of aligned reads: pairs: 455489 (21%) single a: 189879 (8.8%) single b: 483377 (22%) Done. Finished at Thu Feb 13 17:47:59 2014 Total time consumed: 1286 secs
In [1]:
#FEB 13 redux looking at filtered R1
! /Volumes/Bay3/Software/BSMAP/bsmap-2.74/bsmap -a /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC3ATL/filtered_BS_CgF_ACAGTG_L004_R1.fastq.gz -d /Volumes/web/cnidarian/oyster.v9.fa -o /Volumes/web/cnidarian/BiGo_egg_redo3.sam -p 4
BSMAP v2.74 Start at: Thu Feb 13 17:20:34 2014 Input reference file: /Volumes/web/cnidarian/oyster.v9.fa (format: FASTA) Load in 11969 db seqs, total size 558601156 bp. 30 secs passed total_kmers: 43046721 Create seed table. 120 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand: ++,-+ Single-end alignment(4 threads) Input read file: /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC3ATL/filtered_BS_CgF_ACAGTG_L004_R1.fastq.gz (format: gzipped FASTQ) Output file: /Volumes/web/cnidarian/BiGo_egg_redo3.sam (format: SAM) Thread #1: 50000 reads finished. 145 secs passed Thread #0: 100000 reads finished. 145 secs passed Thread #3: 150000 reads finished. 147 secs passed Thread #2: 200000 reads finished. 147 secs passed Thread #1: 250000 reads finished. 169 secs passed Thread #0: 300000 reads finished. 169 secs passed Thread #3: 350000 reads finished. 170 secs passed Thread #2: 400000 reads finished. 171 secs passed Thread #1: 450000 reads finished. 191 secs passed Thread #0: 500000 reads finished. 191 secs passed Thread #3: 550000 reads finished. 192 secs passed Thread #2: 600000 reads finished. 193 secs passed Thread #1: 650000 reads finished. 215 secs passed Thread #0: 700000 reads finished. 215 secs passed Thread #3: 750000 reads finished. 216 secs passed Thread #2: 800000 reads finished. 217 secs passed Thread #1: 850000 reads finished. 240 secs passed Thread #0: 900000 reads finished. 241 secs passed Thread #3: 950000 reads finished. 242 secs passed Thread #2: 1000000 reads finished. 243 secs passed Thread #1: 1050000 reads finished. 267 secs passed Thread #0: 1100000 reads finished. 267 secs passed Thread #3: 1150000 reads finished. 269 secs passed Thread #2: 1200000 reads finished. 270 secs passed Thread #1: 1250000 reads finished. 296 secs passed Thread #0: 1300000 reads finished. 298 secs passed Thread #3: 1350000 reads finished. 300 secs passed Thread #2: 1400000 reads finished. 302 secs passed Thread #1: 1450000 reads finished. 326 secs passed Thread #0: 1500000 reads finished. 327 secs passed Thread #3: 1550000 reads finished. 329 secs passed Thread #2: 1600000 reads finished. 330 secs passed Thread #2: 1786493 reads finished. 353 secs passed Thread #1: 1650000 reads finished. 356 secs passed Thread #0: 1700000 reads finished. 356 secs passed Thread #3: 1750000 reads finished. 358 secs passed Total number of aligned reads: 573217 (32%) Done. Finished at Thu Feb 13 17:26:32 2014 Total time consumed: 358 secs
In [15]:
from IPython.display import Image
Image(url= 'http://eagle.fish.washington.edu/cnidarian/skitch/BiGo_larvae_1892C389.png')
Out[15]:
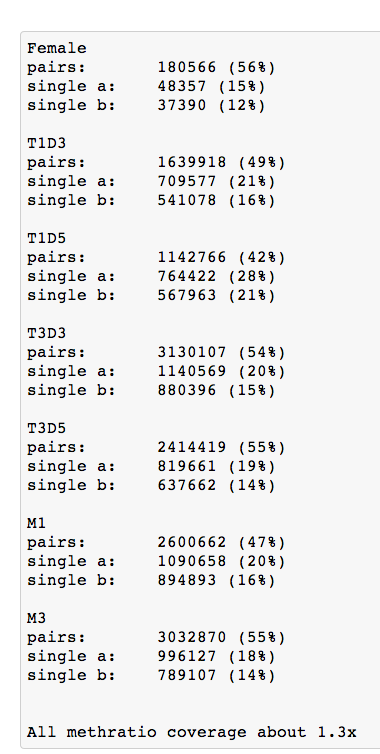
Coverage is different but other samples were paired and longer?
In [16]:
#Lets run good sperm sample with only one end...
#M3
! /Volumes/Bay3/Software/BSMAP/bsmap-2.74/bsmap -a /Volumes/NGS\ Drive/NGS\ Raw\ Data/Cg_larvae_BSseq/filtered_BS_CgM3_GATCAG_L007_R1.fastq.gz -d /Volumes/web/cnidarian/oyster.v9.fa -o /Volumes/web/cnidarian/BiGo_lar_M3_1side.sam -p 3
BSMAP v2.74 Start at: Fri Jan 24 07:54:19 2014 Input reference file: /Volumes/web/cnidarian/oyster.v9.fa (format: FASTA) Load in 11969 db seqs, total size 558601156 bp. 19 secs passed total_kmers: 43046721 Create seed table. 76 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand: ++,-+ Single-end alignment(3 threads) Input read file: /Volumes/NGS Drive/NGS Raw Data/Cg_larvae_BSseq/filtered_BS_CgM3_GATCAG_L007_R1.fastq.gz (format: gzipped FASTQ) Output file: /Volumes/web/cnidarian/BiGo_lar_M3_1side.sam (format: SAM) Thread #0: 50000 reads finished. 102 secs passed Thread #2: 100000 reads finished. 103 secs passed Thread #1: 150000 reads finished. 103 secs passed Thread #0: 200000 reads finished. 128 secs passed Thread #2: 250000 reads finished. 129 secs passed Thread #1: 300000 reads finished. 129 secs passed Thread #0: 350000 reads finished. 149 secs passed Thread #2: 400000 reads finished. 150 secs passed Thread #1: 450000 reads finished. 151 secs passed Thread #0: 500000 reads finished. 172 secs passed Thread #2: 550000 reads finished. 173 secs passed Thread #1: 600000 reads finished. 173 secs passed Thread #0: 650000 reads finished. 194 secs passed Thread #2: 700000 reads finished. 196 secs passed Thread #1: 750000 reads finished. 197 secs passed Thread #0: 800000 reads finished. 224 secs passed Thread #2: 850000 reads finished. 226 secs passed Thread #1: 900000 reads finished. 227 secs passed Thread #0: 950000 reads finished. 261 secs passed Thread #2: 1000000 reads finished. 264 secs passed Thread #1: 1050000 reads finished. 265 secs passed Thread #0: 1100000 reads finished. 288 secs passed Thread #2: 1150000 reads finished. 290 secs passed Thread #1: 1200000 reads finished. 291 secs passed Thread #0: 1250000 reads finished. 309 secs passed Thread #2: 1300000 reads finished. 311 secs passed Thread #1: 1350000 reads finished. 312 secs passed Thread #0: 1400000 reads finished. 329 secs passed Thread #2: 1450000 reads finished. 330 secs passed Thread #1: 1500000 reads finished. 330 secs passed Thread #0: 1550000 reads finished. 349 secs passed Thread #2: 1600000 reads finished. 349 secs passed Thread #1: 1650000 reads finished. 350 secs passed Thread #0: 1700000 reads finished. 375 secs passed Thread #2: 1750000 reads finished. 376 secs passed Thread #1: 1800000 reads finished. 377 secs passed Thread #0: 1850000 reads finished. 403 secs passed Thread #1: 1950000 reads finished. 405 secs passed Thread #2: 1900000 reads finished. 405 secs passed Thread #0: 2000000 reads finished. 429 secs passed Thread #1: 2050000 reads finished. 430 secs passed Thread #2: 2100000 reads finished. 430 secs passed Thread #0: 2150000 reads finished. 448 secs passed Thread #1: 2200000 reads finished. 449 secs passed Thread #2: 2250000 reads finished. 450 secs passed Thread #0: 2300000 reads finished. 470 secs passed Thread #1: 2350000 reads finished. 470 secs passed Thread #2: 2400000 reads finished. 471 secs passed Thread #0: 2450000 reads finished. 495 secs passed Thread #1: 2500000 reads finished. 496 secs passed Thread #2: 2550000 reads finished. 497 secs passed Thread #0: 2600000 reads finished. 522 secs passed Thread #1: 2650000 reads finished. 524 secs passed Thread #2: 2700000 reads finished. 524 secs passed Thread #0: 2750000 reads finished. 549 secs passed Thread #1: 2800000 reads finished. 551 secs passed Thread #2: 2850000 reads finished. 552 secs passed Thread #0: 2900000 reads finished. 572 secs passed Thread #1: 2950000 reads finished. 573 secs passed Thread #2: 3000000 reads finished. 574 secs passed Thread #0: 3050000 reads finished. 589 secs passed Thread #1: 3100000 reads finished. 591 secs passed Thread #2: 3150000 reads finished. 592 secs passed Thread #0: 3200000 reads finished. 609 secs passed Thread #1: 3250000 reads finished. 612 secs passed Thread #2: 3300000 reads finished. 614 secs passed Thread #0: 3350000 reads finished. 635 secs passed Thread #1: 3400000 reads finished. 638 secs passed Thread #2: 3450000 reads finished. 639 secs passed Thread #0: 3500000 reads finished. 661 secs passed Thread #1: 3550000 reads finished. 663 secs passed Thread #2: 3600000 reads finished. 664 secs passed Thread #0: 3650000 reads finished. 681 secs passed Thread #1: 3700000 reads finished. 684 secs passed Thread #2: 3750000 reads finished. 685 secs passed Thread #0: 3800000 reads finished. 701 secs passed Thread #1: 3850000 reads finished. 704 secs passed Thread #2: 3900000 reads finished. 705 secs passed Thread #0: 3950000 reads finished. 722 secs passed Thread #1: 4000000 reads finished. 724 secs passed Thread #2: 4050000 reads finished. 725 secs passed Thread #0: 4100000 reads finished. 743 secs passed Thread #1: 4150000 reads finished. 746 secs passed Thread #2: 4200000 reads finished. 747 secs passed Thread #0: 4250000 reads finished. 765 secs passed Thread #1: 4300000 reads finished. 768 secs passed Thread #2: 4350000 reads finished. 769 secs passed Thread #0: 4400000 reads finished. 785 secs passed Thread #1: 4450000 reads finished. 788 secs passed Thread #2: 4500000 reads finished. 789 secs passed Thread #0: 4550000 reads finished. 805 secs passed Thread #1: 4600000 reads finished. 808 secs passed Thread #2: 4650000 reads finished. 810 secs passed Thread #0: 4700000 reads finished. 825 secs passed Thread #1: 4750000 reads finished. 830 secs passed Thread #2: 4800000 reads finished. 832 secs passed Thread #0: 4850000 reads finished. 847 secs passed Thread #1: 4900000 reads finished. 852 secs passed Thread #2: 4950000 reads finished. 853 secs passed Thread #0: 5000000 reads finished. 869 secs passed Thread #1: 5050000 reads finished. 874 secs passed Thread #2: 5100000 reads finished. 875 secs passed Thread #0: 5150000 reads finished. 888 secs passed Thread #1: 5200000 reads finished. 894 secs passed Thread #2: 5250000 reads finished. 895 secs passed Thread #0: 5300000 reads finished. 909 secs passed Thread #1: 5350000 reads finished. 914 secs passed Thread #2: 5400000 reads finished. 915 secs passed Thread #2: 5526741 reads finished. 926 secs passed Thread #0: 5450000 reads finished. 929 secs passed Thread #1: 5500000 reads finished. 933 secs passed Total number of aligned reads: 3707307 (67%) Done. Finished at Fri Jan 24 08:09:52 2014 Total time consumed: 933 secs
Going Forward with Sperm and Larvae - Combining¶
In [5]:
#stop
#issue with files
from IPython.display import Image
Image(url= 'http://eagle.fish.washington.edu/cnidarian/skitch/trilobite_and_BiGo_Larvae_and_Mackenzie_Gavery_1892E4D6.png')
Out[5]:
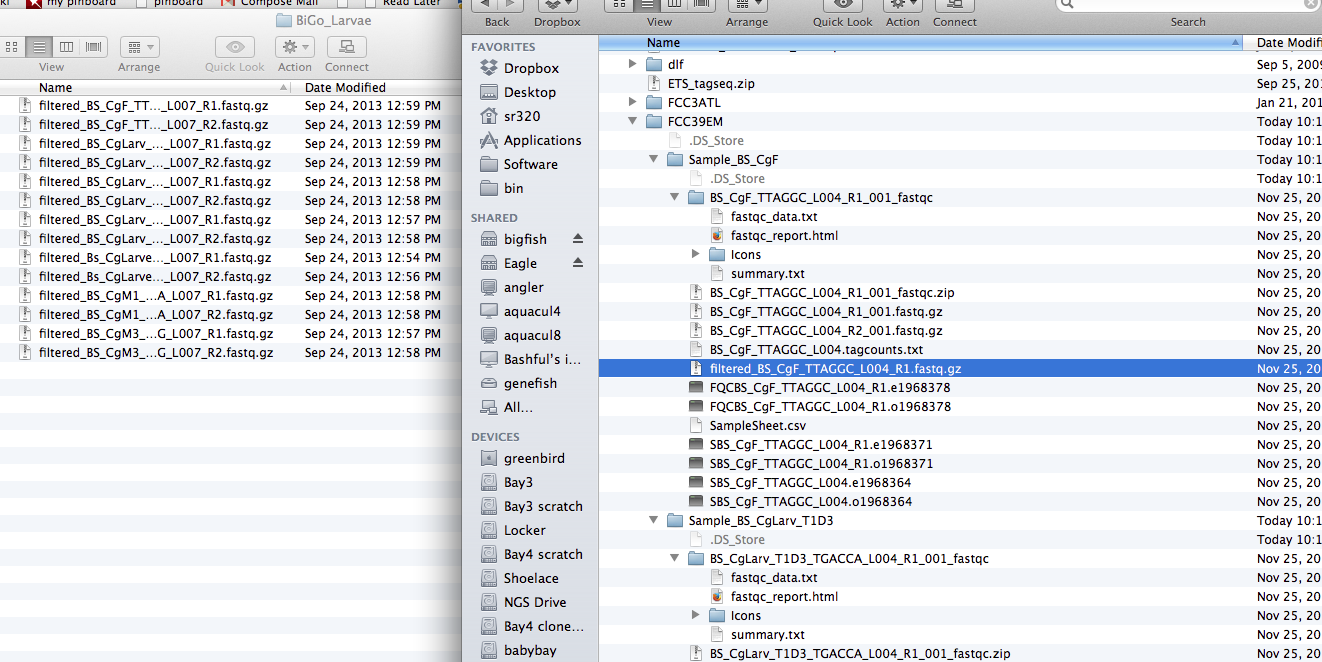
In [ ]:
Only 1 filtered fasta
September Run¶
In [ ]:
Lets QC in IPlant DE
In [10]:
!ils
/iplant/home/sr320: filtered_400_NoIndex_L006_R1.fastq.gz Galaxy10-[Cut_on_data_7].tabular Gil_1.fq.gz Gil_2.fq.gz oyster.v9.fa oyster.v9.glean.final.rename.CDS.gff solid0078_20091105_BB3.fastq solid0078_20091105_DH3.fastq SPID_and_GO_number.tabular Transcriptome_module1_blastxout_CE_modified.txt Unknown_Transcriptome_31545_contigs.fa C- /iplant/home/sr320/AltSplice C- /iplant/home/sr320/analyses C- /iplant/home/sr320/bamout C- /iplant/home/sr320/bisque_data C- /iplant/home/sr320/Black_Abalone C- /iplant/home/sr320/cacheServiceTempDir C- /iplant/home/sr320/ce C- /iplant/home/sr320/Cgigas_v9 C- /iplant/home/sr320/coge_data C- /iplant/home/sr320/fish546 C- /iplant/home/sr320/iplant_ws C- /iplant/home/sr320/OlympiaOyster C- /iplant/home/sr320/OlyO_PacBio C- /iplant/home/sr320/OlyO_SNPhunt C- /iplant/home/sr320/qdod C- /iplant/home/sr320/RNAseq_co_BS
In [11]:
!imkdir BiGo_larvae
In [12]:
!ils
/iplant/home/sr320: filtered_400_NoIndex_L006_R1.fastq.gz Galaxy10-[Cut_on_data_7].tabular Gil_1.fq.gz Gil_2.fq.gz oyster.v9.fa oyster.v9.glean.final.rename.CDS.gff solid0078_20091105_BB3.fastq solid0078_20091105_DH3.fastq SPID_and_GO_number.tabular Transcriptome_module1_blastxout_CE_modified.txt Unknown_Transcriptome_31545_contigs.fa C- /iplant/home/sr320/AltSplice C- /iplant/home/sr320/analyses C- /iplant/home/sr320/bamout C- /iplant/home/sr320/BiGo_larvae C- /iplant/home/sr320/bisque_data C- /iplant/home/sr320/Black_Abalone C- /iplant/home/sr320/cacheServiceTempDir C- /iplant/home/sr320/ce C- /iplant/home/sr320/Cgigas_v9 C- /iplant/home/sr320/coge_data C- /iplant/home/sr320/fish546 C- /iplant/home/sr320/iplant_ws C- /iplant/home/sr320/OlympiaOyster C- /iplant/home/sr320/OlyO_PacBio C- /iplant/home/sr320/OlyO_SNPhunt C- /iplant/home/sr320/qdod C- /iplant/home/sr320/RNAseq_co_BS
In [15]:
!icd BiGo_larvae
In [16]:
!iput -f -V -r /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/BiGo_Larvae
NOTICE: irodsHost=data.iplantcollaborative.org NOTICE: irodsPort=1247 NOTICE: irodsUserName=sr320 NOTICE: irodsZone=iplant NOTICE: created irodsHome=/iplant/home/sr320 NOTICE: created irodsCwd=/iplant/home/sr320 NOTICE: irodsCwd=/iplant/home/sr320/BiGo_larvae C- /iplant/home/sr320/BiGo_larvae/BiGo_Larvae: From server: NumThreads=11, addr:128.196.172.248, port:20049, cookie=1784423902 filtered_BS_CgM3_GATCAG_L 329.845 MB | 19.206 sec | 11 thr | 17.174 MB/s From server: NumThreads=10, addr:128.196.172.248, port:20240, cookie=874741332 filtered_BS_CgM3_GATCAG_L 290.549 MB | 17.154 sec | 10 thr | 16.937 MB/s From server: NumThreads=11, addr:128.196.172.248, port:20001, cookie=1488887683 filtered_BS_CgM1_ACTTGA_L 330.421 MB | 16.623 sec | 11 thr | 19.878 MB/s From server: NumThreads=10, addr:128.196.172.248, port:20362, cookie=696179826 filtered_BS_CgM1_ACTTGA_L 290.698 MB | 18.305 sec | 10 thr | 15.881 MB/s From server: NumThreads=11, addr:128.196.172.248, port:20022, cookie=2050863989 filtered_Bs_CgLarve_T3D3_ 348.038 MB | 14.489 sec | 11 thr | 24.020 MB/s From server: NumThreads=10, addr:128.196.172.248, port:20148, cookie=551333819 filtered_Bs_CgLarve_T3D3_ 306.693 MB | 13.414 sec | 10 thr | 22.864 MB/s From server: NumThreads=9, addr:128.196.172.248, port:20086, cookie=2112982237 filtered_BS_CgLarv_T3D5_C 264.221 MB | 13.685 sec | 9 thr | 19.308 MB/s From server: NumThreads=8, addr:128.196.172.248, port:20253, cookie=295873469 filtered_BS_CgLarv_T3D5_C 232.154 MB | 10.373 sec | 8 thr | 22.380 MB/s From server: NumThreads=6, addr:128.196.172.248, port:20259, cookie=1552130862 filtered_BS_CgLarv_T1D5_A 162.282 MB | 10.573 sec | 6 thr | 15.349 MB/s From server: NumThreads=5, addr:128.196.172.248, port:20268, cookie=102187342 filtered_BS_CgLarv_T1D5_A 142.537 MB | 11.286 sec | 5 thr | 12.630 MB/s From server: NumThreads=7, addr:128.196.172.248, port:20240, cookie=1903399127 filtered_BS_CgLarv_T1D3_T 200.030 MB | 68.625 sec | 7 thr | 2.915 MB/s From server: NumThreads=6, addr:128.196.172.248, port:20128, cookie=648041722 filtered_BS_CgLarv_T1D3_T 175.973 MB | 63.189 sec | 6 thr | 2.785 MB/s filtered_BS_CgF_TTAGGC_L0 19.369 MB | 13.399 sec | 0 thr | 1.446 MB/s filtered_BS_CgF_TTAGGC_L0 17.302 MB | 2.144 sec | 0 thr | 8.070 MB/s From server: NumThreads=2, addr:128.196.172.248, port:20085, cookie=258361285 filtered_BS_CgF_TTAGGC_L0 62.435 MB | 13.086 sec | 2 thr | 4.771 MB/s .DS_Store 0.006 MB | 0.184 sec | 0 thr | 0.032 MB/s
No QC by Core - using iPlant DE
In [51]:
!ils
/iplant/home/sr320/BiGo_larvae: filtered_BS_CgF_TTAGGC_L007_R1.fastq filtered_BS_CgF_TTAGGC_L007_R2.fastq filtered_Bs_CgLarve_T3D3_GCCAAT_L007_R1.fastq filtered_Bs_CgLarve_T3D3_GCCAAT_L007_R2.fastq filtered_BS_CgLarv_T1D3_TGACCA_L007_R1.fastq filtered_BS_CgLarv_T1D3_TGACCA_L007_R2.fastq filtered_BS_CgLarv_T1D5_ACAGTG_L007_R1.fastq filtered_BS_CgLarv_T1D5_ACAGTG_L007_R2.fastq filtered_BS_CgLarv_T3D5_CAGATC_L007_R1.fastq filtered_BS_CgLarv_T3D5_CAGATC_L007_R2.fastq filtered_BS_CgM1_ACTTGA_L007_R1.fastq filtered_BS_CgM1_ACTTGA_L007_R2.fastq filtered_BS_CgM3_GATCAG_L007_R1.fastq filtered_BS_CgM3_GATCAG_L007_R2.fastq C- /iplant/home/sr320/BiGo_larvae/FastQC_0.10.1__multi-file__analysis1-2014-02-03-09-29-30.603
In [53]:
!iget -r FastQC_0.10.1__multi-file__analysis1-2014-02-03-09-29-30.603
In [38]:
cd /Volumes/web/cnidarian
/Volumes/web/cnidarian
September Run FastQC Reports¶
In [3]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgF_TTAGGC_L007_R1_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[3]:
In [60]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgF_TTAGGC_L007_R2_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[60]:
In [61]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_Bs_CgLarve_T3D3_GCCAAT_L007_R1_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[61]:
In [62]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_Bs_CgLarve_T3D3_GCCAAT_L007_R2_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[62]:
Check "non-filtered" T3D3 files¶
In [74]:
pwd
Out[74]:
u'/Volumes/web/cnidarian'
In [75]:
!iget -r FastQC_0.10.1__multi-file__analysis1-2014-02-03-11-29-22.343
In [79]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314b/Bs_CgLarve_T3D3_GCCAAT_L007_R1_001_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[79]:
In [80]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314b/Bs_CgLarve_T3D3_GCCAAT_L007_R2_001_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[80]:
In [63]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgLarv_T1D3_TGACCA_L007_R1_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[63]:
In [64]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgLarv_T1D3_TGACCA_L007_R2_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[64]:
In [65]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgLarv_T1D5_ACAGTG_L007_R1_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[65]:
In [66]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgLarv_T1D5_ACAGTG_L007_R2_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[66]:
In [67]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgLarv_T3D5_CAGATC_L007_R1_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[67]:
In [68]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgLarv_T3D5_CAGATC_L007_R2_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[68]:
In [69]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgM1_ACTTGA_L007_R1_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[69]:
In [70]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgM1_ACTTGA_L007_R2_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[70]:
In [71]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgM3_GATCAG_L007_R1_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[71]:
In [72]:
from IPython.display import HTML
HTML('<iframe src=http://eagle.fish.washington.edu/cnidarian/FastQC_0.10.1_020314/filtered_BS_CgM3_GATCAG_L007_R2_fastqc/fastqc_report.html width=100% height=550></iframe>')
Out[72]:
November Run¶
Joining / Merging¶
Lets have a look at a Pair of files
In [ ]:
first need to de compress