Crassostrea gigas RNA-Seq¶
Tophat via iPlant¶
Goal: Use RNA-seq to compare expression between oysters (n=3) pre and post heat shock. __
Based on IPlant Collaborative Tutorial
 https://pods.iplantcollaborative.org/wiki/display/eot/RNA-Seq_tutorial
https://pods.iplantcollaborative.org/wiki/display/eot/RNA-Seq_tutorial
Task 1: Align read data to Crassostrea gigas genome.¶
Tophat is a specialized alignment software for RNA-seq reads that is aware of splice junctions when aligning to a reference assembly.
- Click Apps from DE workspace and select TopHat2-SE. Use search bar.
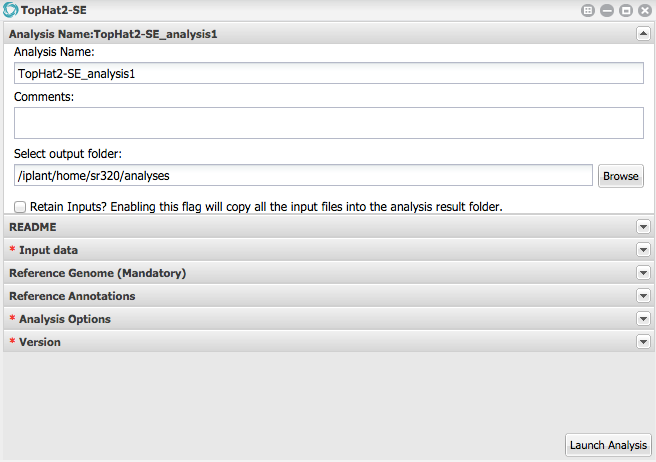
Under 'Analysis Name' leave as defaults for make any changes.
Under Input data for FASTQ files add six fastq.gz files localed in
austral-datawith prefixes 2M, 2M-HS, 4M, 4M-HS, 6M, 6M-HS.Under Reference Genome for 'Provide a reference genome file in FASTA format' select
/austral-data/Crassostrea_gigas.GCA_000297895.1.24.dna_sm.toplevel.faFor Reference Annoations add the GTF file
/austral-data/Crassostrea_gigas.GCA_000297895.1.24.gtfClick Launch Analyses and monitor the status of you job.
Once complete....
- Note this takes ~10 hours
Task 2: Assemble transcripts using Cufflinks2¶
Cufflinks assembles RNA-Seq alignments into a parsimonious set of transcripts, then estimates the relative abundances of these transcripts based on how many reads support each one, taking into account biases in library preparation protocols. A detailed manual can be found at http://cufflinks.cbcb.umd.edu/manual.html.
Open Cufflinks2
For Input Data add the six bam files from the
bamsubdirectory of the TopHat2 output.Under Reference Sequence use custom option select
/austral-data/Crassostrea_gigas.GCA_000297895.1.24.dna_sm.toplevel.faFor Reference Annoations add the GTF file
/austral-data/Crassostrea_gigas.GCA_000297895.1.24.gtfClick Launch Analyses and monitor the status of you job.
- Note this takes ~2 hours
Task 3: Merge all Cufflinks transcripts into a single transcriptome annotation file using Cuffmerge2¶
Cuffmerge merges together several Cufflinks assemblies. It handles also handles running Cuffcompare. The main purpose of this application is to make it easier to make an assembly GTF file suitable for use with Cuffdiff. A merged, empirical annotation file will be more accurate than using the standard reference annotation, as the expression of rare or novel genes and alternative splicing isoforms seen in this experiment will be better reflected in the empirical transcriptome assemblies.
Open the Cuffmerge2 app. Under 'Input Data', browse to the results of the cufflinks analyses (abovee) and add the 6 gtf files located in the
gtfsubdirectory.For Reference Annoations add the GTF file
/austral-data/Crassostrea_gigas.GCA_000297895.1.24.gtfUnder Reference Sequence use custom option select
/austral-data/Crassostrea_gigas.GCA_000297895.1.24.dna_sm.toplevel.faClick Launch Analyses and monitor the status of you job.
Task 4: Compare expression analysis using CuffDiff2¶
Cuffdiff is a program that uses the Cufflinks transcript quantification engine to calculate gene and transcript expression levels in more than one condition and test them for significant differences. http://cufflinks.cbcb.umd.edu/manual.html
- Open Cuffdiff2. For 'Input Data' Sample 1 Name enter Pre adn add three bam file from Tophat analysis..
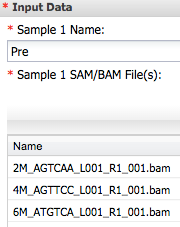
For Sample 2 enter post and add other three bam files ...
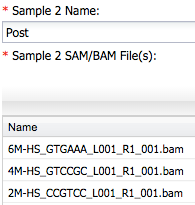
Next, open the Reference Annotations section and add a custom reference annotation file using the
merged_with_ref_ids.gtffile from the cuffmerge output folder.Click Launch Analyses and monitor the status of you job.
Cuffdiff2 Output¶
!ls ../analyses/Cuffdiff2_heat-b-2014-12-20-22-27-15.4/
cuffdiff.stderr cuffdiff_out graphs logs sorted_data
!ls ../analyses/Cuffdiff2_heat-b-2014-12-20-22-27-15.4/sorted_data/
genes.sorted_by_expression.sig.txt genes.sorted_by_expression.txt genes.sorted_by_fold.sig.txt genes.sorted_by_fold.txt transcripts.sorted_by_expression.sig.txt transcripts.sorted_by_expression.txt transcripts.sorted_by_fold.sig.txt transcripts.sorted_by_fold.txt
!head ../analyses/Cuffdiff2_heat-b-2014-12-20-22-27-15.4/sorted_data/genes.sorted_by_fold.sig.txt
gene_id gene_name sample1 sample2 fold_change direction total_fpkm q-value gene_description XLOC_003768 HSP70 Pre Post 280.15 UP 1936.97 0.00215 XLOC_032537 - Pre Post 260.76 DOWN 2614.87 0.00215 CGI_10020701 . Pre Post 239.05 DOWN 4568.52 0.00215 CGI_10002594 . Pre Post 224.07 UP 2390.25 0.00215 XLOC_003774 CGI_10010646 Pre Post 181.01 UP 1818.52 0.00215 XLOC_032785 HSP68 Pre Post 155.89 UP 373.66 0.00215 XLOC_015275 CGI_10011376 Pre Post 122.85 UP 418.73 0.0228219 CGI_10001496 . Pre Post 119.83 DOWN 1450.68 0.00215 CGI_10011376 . Pre Post 97.71 UP 528.85 0.00215
!ls ../analyses/Cuffdiff2_heat-b-2014-12-20-22-27-15.4/graphs/
Pre_Post_scatter_plot.png Pre_Post_volcano_plot.png density_plot.png


