Model Evaluation¶
Review of last class¶
- Goal was to predict the species of an unknown iris
- Made predictions using KNN models with different values of K
- Need a way to choose the "best" model: the one that "generalizes" to "out-of-sample" data
Solution: Create a procedure that estimates how well a model is likely to perform on out-of-sample data and use that to choose between models.
Evaluation procedure #1: Train and test on the entire dataset¶
- Train the model on the entire dataset.
- Test the model on the same dataset, and evaluate how well we did by comparing the predicted response values with the true response values.
# read the iris data into a DataFrame
import pandas as pd
url = 'http://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data'
col_names = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species']
iris = pd.read_csv(url, header=None, names=col_names)
# map each iris species to a number
iris['species_num'] = iris.species.map({'Iris-setosa':0, 'Iris-versicolor':1, 'Iris-virginica':2})
# store feature matrix in "X"
feature_cols = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width']
X = iris[feature_cols]
# store response vector in "y"
y = iris.species_num
KNN (K=50)¶
# import the class
from sklearn.neighbors import KNeighborsClassifier
# instantiate the model
knn = KNeighborsClassifier(n_neighbors=50)
# train the model on the entire dataset
knn.fit(X, y)
# predict the response values for the observations in X ("test the model")
knn.predict(X)
array([0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 1, 1, 2, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 2, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 1, 2, 2, 2, 2, 2, 2, 2, 2,
2, 2, 2, 2, 1, 2, 1, 2, 2, 2, 2, 1, 1, 2, 2, 2, 2, 2, 1, 2, 2, 2, 2,
1, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2], dtype=int64)
# store the predicted response values
y_pred = knn.predict(X)
To evaluate a model, we also need an evaluation metric:
- Numeric calculation used to quantify the performance of a model
- Appropriate metric depends on the goals of your problem
Most common choices for classification problems:
- Classification accuracy: percentage of correct predictions (reward function)
- Classification error: percentage of incorrect predictions (loss function)
In this case, we'll use classification accuracy.
# compute classification accuracy
from sklearn import metrics
print metrics.accuracy_score(y, y_pred)
0.94
This is known as training accuracy because we are testing the model on the same data we used to train the model.
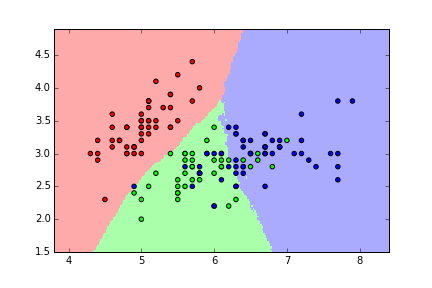
KNN (K=1)¶
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X, y)
y_pred = knn.predict(X)
print metrics.accuracy_score(y, y_pred)
1.0
Does that mean that K=1 is the best value for K?
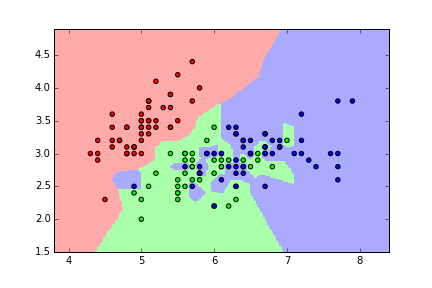
Problems with training and testing on the same data¶
- Goal is to estimate likely performance of a model on out-of-sample data
- But, maximizing training accuracy rewards overly complex models that won't necessarily generalize
- Unnecessarily complex models overfit the training data:
- Will do well when tested using the in-sample data
- May do poorly on out-of-sample data
- Learns the "noise" in the data rather than the "signal"
- From Quora: What is an intuitive explanation of overfitting?
Thus, training accuracy is not a good estimate of out-of-sample accuracy.
Evaluation procedure #2: Train/test split¶
- Split the dataset into two pieces: a training set and a testing set.
- Train the model on the training set.
- Test the model on the testing set, and evaluate how well we did.
What does this accomplish?
- Model can be trained and tested on different data (we treat testing data like out-of-sample data).
- Response values are known for the testing set, and thus predictions can be evaluated.
- Testing accuracy is a better estimate than training accuracy of out-of-sample performance.
Side note on "unpacking"¶
def min_max(nums):
smallest = min(nums)
largest = max(nums)
return [smallest, largest]
min_and_max = min_max([1, 2, 3])
print min_and_max
print type(min_and_max)
[1, 3] <type 'list'>
the_min, the_max = min_max([1, 2, 3])
print the_min
print type(the_min)
print the_max
print type(the_max)
1 <type 'int'> 3 <type 'int'>
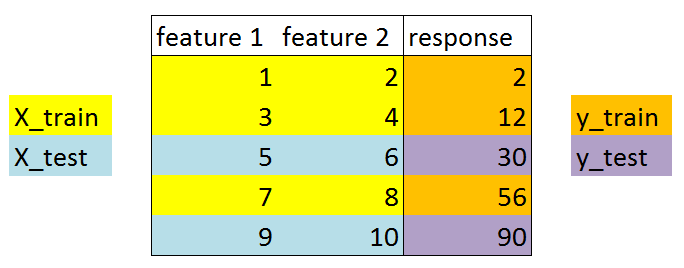
Understanding the train_test_split function¶
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4)
# before splitting
print X.shape
# after splitting
print X_train.shape
print X_test.shape
(150, 4) (90L, 4L) (60L, 4L)
# before splitting
print y.shape
# after splitting
print y_train.shape
print y_test.shape
(150L,) (90L,) (60L,)

Understanding the random_state parameter¶
# WITHOUT a random_state parameter
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4)
# print the first element of each object
print X_train[0]
print X_test[0]
print y_train[0]
print y_test[0]
[ 6.1 3. 4.6 1.4] [ 6.2 2.9 4.3 1.3] 1 1
# WITH a random_state parameter
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=4)
# print the first element of each object
print X_train[0]
print X_test[0]
print y_train[0]
print y_test[0]
[ 4.6 3.4 1.4 0.3] [ 6.4 2.8 5.6 2.1] 0 2
Using the train/test split procedure (K=1)¶
# STEP 1: split X and y into training and testing sets (using random_state for reproducibility)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=4)
# STEP 2: train the model on the training set (using K=1)
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train, y_train)
KNeighborsClassifier(algorithm='auto', leaf_size=30, metric='minkowski',
metric_params=None, n_neighbors=1, p=2, weights='uniform')
# STEP 3: test the model on the testing set, and check the accuracy
y_pred = knn.predict(X_test)
print metrics.accuracy_score(y_test, y_pred)
0.95
Repeat for K=50¶
knn = KNeighborsClassifier(n_neighbors=50)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
print metrics.accuracy_score(y_test, y_pred)
0.933333333333
Search for the "best" value of K¶
# calculate TRAINING ERROR and TESTING ERROR for K=1 through 50
k_range = range(1, 51)
training_error = []
testing_error = []
for k in k_range:
knn = KNeighborsClassifier(n_neighbors=k)
# training error
knn.fit(X, y)
y_pred = knn.predict(X)
training_error.append(1 - metrics.accuracy_score(y, y_pred))
# testing error
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
testing_error.append(1 - metrics.accuracy_score(y_test, y_pred))
%matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# plot the relationship between K (HIGH TO LOW) and TESTING ERROR
plt.plot(k_range, testing_error)
plt.gca().invert_xaxis()
plt.xlabel('Value of K for KNN')
plt.ylabel('Testing Error')
<matplotlib.text.Text at 0x176cb860>
What can we conclude?
- A value of K around 11 is likely the best value for K when using KNN on the iris dataset.
- When given the measurements of an unknown iris, we estimate that we would be able to correctly predict its species 98% of the time.
Training error versus testing error¶
# create a DataFrame of K, training error, and testing error
df = pd.DataFrame({'K': k_range, 'train':training_error, 'test':testing_error}).set_index('K').sort_index(ascending=False)
df.head()
| test | train | |
|---|---|---|
| K | ||
| 50 | 0.066667 | 0.060000 |
| 49 | 0.083333 | 0.040000 |
| 48 | 0.066667 | 0.053333 |
| 47 | 0.050000 | 0.046667 |
| 46 | 0.050000 | 0.053333 |
# plot the relationship between K (HIGH TO LOW) and both TRAINING ERROR and TESTING ERROR
df.plot()
<matplotlib.axes._subplots.AxesSubplot at 0x176f04e0>
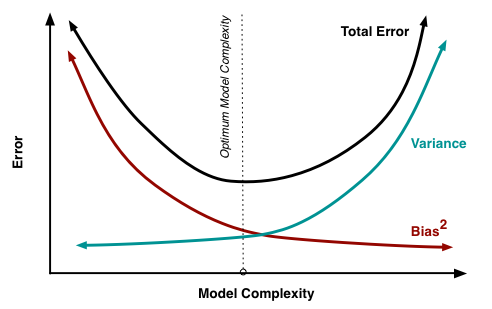
Roughly speaking:
- Training error decreases as model complexity increases (lower value of K)
- Testing error is minimized at the optimum model complexity
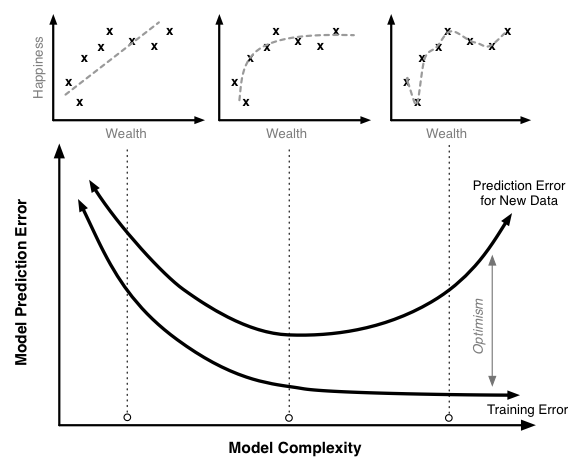
Making predictions on out-of-sample data¶
Given the measurements of a (truly) unknown iris, how do we predict its species?
# instantiate the model with the best known parameters
knn = KNeighborsClassifier(n_neighbors=11)
# re-train the model with X and y (not X_train and y_train) - why?
knn.fit(X, y)
# make a prediction for an out-of-sample observation
knn.predict([3, 5, 4, 2])
array([1], dtype=int64)
Disadvantages of train/test split?¶
What would happen if the train_test_split function had split the data differently? Would we get the same exact results as before?
# try different values for random_state
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=4)
knn = KNeighborsClassifier(n_neighbors=50)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
print metrics.accuracy_score(y_test, y_pred)
0.933333333333
- Testing accuracy is a high-variance estimate of out-of-sample accuracy
- K-fold cross-validation overcomes this limitation and provides more reliable estimates
- But, train/test split is still useful because of its flexibility and speed