Problem 1¶
Consider the following two-dimensional diffision problem. Use the discretization below to solve for the
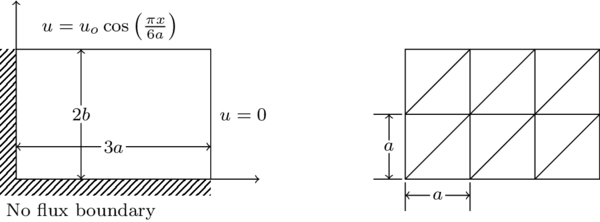
unknown diffusing concentrations at the nodes with $a = 1$ and $u_o = 100$ assuming a consistent unit system. The diffusion coefficient matrix is the identity matrix. Perform the following three tasks:
Solve this problem using a direct integration of the trianglur elements. (20 points)
Solve this problem by using a "parent" element mapping and Gauss integration on the rectangular element (i.e. ignore the triangular elements). Use a $2 \times 2$ Gauss integration rule. (20 points)
Create effective plots to visualize your results from 1 and 2. (5 points)
Note: Submit a working version of your code to Canvas. Any supplemental material explaining your answer in part (b) can be turned in to me via hard copy or scanned and submitted to Canvas with your code.
Solution
Below we will write the class and methods that will help us solve the problem.
import numpy as np
import scipy.linalg
import scipy.sparse
import scipy.sparse.linalg
class TwoDimFEM():
def __init__(self, nodes, connect, C=np.array([[1., 0, 0], [0, 1., 0], [0, 0, 0]])):
"""
Initialize the model.
input: nodes -> the nodal locations as a Numpy array of (x,y) pairs.
input: connect -> the connectivitiy array
output: the model problem object
"""
self.X = nodes[:,0]
self.Y = nodes[:,1]
#Subtract 1 to convert node numbers to be consistent with Numpy 0 indexing
self.connect = (connect - 1)
self.num_elem = len(self.connect)
self.num_dof = len(self.X)
self.Cmat = C
#Allocate global tangent and r.h.s vector
self.K = np.zeros((self.num_dof, self.num_dof), dtype=np.double)
self.F = np.zeros(self.num_dof, dtype=np.double)
def N(self, xi, eta):
"""Compute linear shape functions in natural coordinates."""
return [1 / 4. - eta / 4. - xi / 4. + (eta * xi) / 4.,
1 / 4. - eta / 4. + xi / 4. - (eta * xi) / 4.,
1 / 4. + eta / 4. + xi / 4. + (eta * xi) / 4.,
1 / 4. + eta / 4. - xi / 4. - (eta * xi) / 4.]
def dNdxi(self, eta):
"""Compute shape function derivatives with respect to xi"""
return [-1 / 4. + eta / 4.,
1 / 4. - eta / 4.,
1 / 4. + eta / 4.,
-1 / 4. - eta / 4.]
def dNdeta(self, xi):
"""Compute shape function derivatives with respect to eta"""
return [-1 / 4. + xi / 4.,
-1 / 4. - xi / 4.,
1 / 4. + xi / 4.,
1 / 4. - xi / 4.]
def compute_jacobian_matrix_and_inverse(self, xi, eta):
"""
Compute the Jacobian matrix, Det(J) and B for every element
"""
x = self.X
y = self.Y
con = self.connect
#Understand we are broadcasting the dot product to every element
J11 = np.dot(x[con], self.dNdxi(xi))
J12 = np.dot(y[con], self.dNdxi(xi))
J21 = np.dot(x[con], self.dNdeta(eta))
J22 = np.dot(y[con], self.dNdeta(eta))
#detJ is a vector containing the Jacobian determinate for every element
self.detJ = J11 * J22 - J12 * J21
self.Jinv11 = J22 / self.detJ
self.Jinv12 = -J12 / self.detJ
self.Jinv21 = -J21 / self.detJ
self.Jinv22 = J11 / self.detJ
def compute_B_matrix(self, xi, eta):
"""Computes the B matrix for a given xi and eta"""
#Returns detJ and Jinv components for this xi and eta
self.compute_jacobian_matrix_and_inverse(xi, eta)
Nmat = np.zeros((3, 4), dtype=np.double)
Nmat[0,:] = self.dNdxi(xi)
Nmat[1,:] = self.dNdeta(eta)
Nmat[2,:] = self.N(xi, eta)
zero = np.zeros(len(self.detJ))
one = np.ones(len(self.detJ))
Jmat = np.array([[self.Jinv11, self.Jinv12, zero],
[self.Jinv21, self.Jinv22, zero],
[ zero, zero, one]])
#B = J * N
return np.einsum('jk...,kl', Jmat, Nmat)
def compute_stiffness_integrand(self, xi, eta):
"""Computes the integrand of the stiffness matrix for this xi and eta"""
Cmat = self.Cmat
#Returns B
self.Bmat = self.compute_B_matrix(xi, eta)
#ke_{il} = B_{ji} C_{jk} B_{kl} \det(J)
return np.einsum('...ji,jk,...kl,...',self.Bmat,Cmat,self.Bmat,self.detJ)
def integrate_element_matrices(self):
"""Integrate stiffness matrix with Gauss integration"""
#Use 2 x 2 Gauss integration
wts = [1., 1.]
pts = [-np.sqrt(1 / 3.), np.sqrt(1 / 3.)]
ke = np.zeros((self.num_elem, 4, 4))
for i in range(2):
for j in range(2):
ke += wts[i] * wts[j] * self.compute_stiffness_integrand(pts[i], pts[j])
return ke
def assemble(self):
"""Assemble element stiffness into global"""
#Check if using natural coordinate quads, if so we will integrate using Gauss integration
if len(connect[0]) == 4:
ke = self.integrate_element_matrices()
for i in range(self.num_elem):
idx_grid = np.ix_(self.connect[i], self.connect[i])
self.K[idx_grid] += ke[i]
#Check if using triangles, if so we will use the analytic stiffness matrix
elif len(connect[0]) == 3:
c00 = self.Cmat[2,2]
c11 = self.Cmat[0,0]
c12 = self.Cmat[0,1]
c21 = self.Cmat[1,0]
c22 = self.Cmat[1,1]
ke = np.array([[1 / 12. * (c00 + 6 * (c11 + c12 + c21 + c22)),
1 / 24. * (c00 - 12 * (c11 + c21)),
1 / 24. * (c00 - 12 * (c12 + c22))],
[1 / 24. * (c00 - 12 * (c11 + c12)),
1 / 12. * (c00 + 6 * c11),
1 / 24. * (c00 + 12 * c12)],
[1 / 24. * (c00 - 12 * (c21 + c22)),
1 / 24. * (c00 + 12 * c21),
1 / 12. * (c00 + 6 * c22)]])
for i in range(self.num_elem):
idx_grid = np.ix_(self.connect[i], self.connect[i])
self.K[idx_grid] += ke
def apply_essential_bc(self, nodes, values):
"""
Modifies stiffness matrix and r.h.s. vector for essential b.c.'s
at location of nodes with corresponding values.
"""
#Subtract 1 to bring the node numbers into alignment with Python indexing
node_idx = nodes - 1
row_replace = np.zeros(self.num_dof)
for value_idx, node in enumerate(node_idx):
self.K[node] = row_replace
self.K[node,node] = 1
self.F[node] = values[value_idx]
def solve(self):
"""Solve the global system of equations"""
self.K = scipy.sparse.csr_matrix(self.K)
return scipy.sparse.linalg.spsolve(self.K,self.F)
Now we can solve the problem. I will answer part (2) first using the square elements with linear isoparametric interpolation.
#The node locations
nodes = np.array([[0,0],[1,0],[2,0],[3,0],
[0,1],[1,1],[2,1],[3,1],
[0,2],[1,2],[2,2],[3,2]], dtype=np.double)
#The connectivity array for the quadratic elements using standard node numbering scheme
connect = np.array([[1, 2, 6, 5],
[2, 3, 7, 6],
[3, 4, 8, 7],
[5, 6, 10, 9],
[6, 7, 11, 10],
[7, 8, 12, 11]], dtype=np.int64)
#Create a boundary condition node set and vaules for the top side
ns1 = np.array([9, 10, 11, 12], dtype=np.int64)
val1 = np.cos(np.pi * np.array([0., 1., 2., 3.]) / 6.) * 100
#Create a boundary condition node set for the right side
ns2 = np.array([4, 8, 12], dtype=np.int64)
val2 = np.zeros(len(ns2))
#Instatiate the problem
problem = TwoDimFEM(nodes, connect)
#Assemble
problem.assemble()
#Apply boundary conditions
problem.apply_essential_bc(ns1,val1)
problem.apply_essential_bc(ns2,val2)
#Solve
u = problem.solve()
u
array([ 61.28417886, 53.07365574, 30.64208943, 0. ,
70.29995898, 60.88155036, 35.14997949, 0. ,
100. , 86.60254038, 50. , 0. ])
%matplotlib inline
import matplotlib.pyplot as plt
#Plot the results
plot = plt.contourf(nodes[:,0].reshape(3,4), nodes[:,1].reshape(3,4), u.reshape(3,4),cmap="coolwarm")
plt.colorbar(plot, orientation='horizontal', shrink=0.6);
plt.clim(0,100)
plt.axes().set_aspect('equal')
/Users/john/miniconda3/lib/python3.5/site-packages/matplotlib/collections.py:590: FutureWarning: elementwise comparison failed; returning scalar instead, but in the future will perform elementwise comparison
if self._edgecolors == str('face'):
Now we will solve part (1) using the triangles and an analytic stiffness matrix
#The node locations are the same, so we will updated the connectivities
connect = np.array([[5, 1, 6],
[2, 6, 1],
[6, 2, 7],
[3, 7, 2],
[7, 3, 8],
[4, 8, 3],
[9, 5, 10],
[6, 10, 5],
[10, 6, 11],
[7, 11, 6],
[11, 7, 12],
[8, 12, 7]], dtype=np.int64)
#Instantiate the problem
problem = TwoDimFEM(nodes, connect)
#Assemble
problem.assemble()
#Apply boundary conditions
problem.apply_essential_bc(ns1,val1)
problem.apply_essential_bc(ns2,val2)
#Solve
u = problem.solve()
#Plot the results
plot = plt.contourf(nodes[:,0].reshape(3,4), nodes[:,1].reshape(3,4), u.reshape(3,4),cmap="coolwarm")
plt.colorbar(plot, orientation='horizontal', shrink=0.6);
plt.clim(0,100)
plt.axes().set_aspect('equal')
/Users/john/miniconda3/lib/python3.5/site-packages/matplotlib/collections.py:590: FutureWarning: elementwise comparison failed; returning scalar instead, but in the future will perform elementwise comparison
if self._edgecolors == str('face'):