Tied Edges Using YAHMM¶
Author: Jacob Schreiber jmschreiber91@gmail.com
Tied edges are similar to tied distributions, in that they affect the training of edges. Edges which are tied change training such that a transition across one of them counts as a transition across all of them. This can be useful if you have a large repeating model, but not enough data to train each position in that model.
For example, lets say that you had a DNA profile model which looked like this:
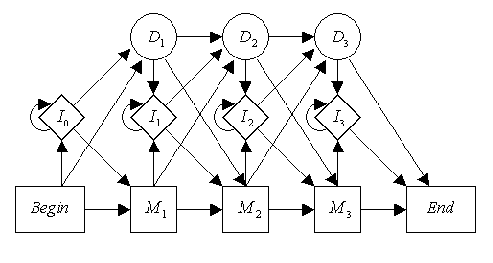
Picture from Biological Sequence Analysis: Probabilistic Methods of Proteins and Nucleic Acids by Durbin, Eddy, Krogh, and Mitchison
The code to make this model would be:
from yahmm import *
from numpy import *
model = Model( "Global Alignment")
# Define the distribution for insertions
i_d = DiscreteDistribution( { 'A': 0.25, 'C': 0.25, 'G': 0.25, 'T': 0.25 } )
# Create the insert states
i0 = State( i_d, name="I0" )
i1 = State( i_d, name="I1" )
i2 = State( i_d, name="I2" )
i3 = State( i_d, name="I3" )
# Create the match states
m1 = State( DiscreteDistribution({ "A": 0.95, 'C': 0.01, 'G': 0.01, 'T': 0.02 }) , name="M1" )
m2 = State( DiscreteDistribution({ "A": 0.003, 'C': 0.99, 'G': 0.003, 'T': 0.004 }) , name="M2" )
m3 = State( DiscreteDistribution({ "A": 0.01, 'C': 0.01, 'G': 0.01, 'T': 0.97 }) , name="M3" )
# Create the delete states
d1 = State( None, name="D1" )
d2 = State( None, name="D2" )
d3 = State( None, name="D3" )
# Add all the states to the model
model.add_states( [i0, i1, i2, i3, m1, m2, m3, d1, d2, d3 ] )
# Create transitions from match states
model.add_transition( model.start, m1, 0.9 )
model.add_transition( model.start, i0, 0.1 )
model.add_transition( m1, m2, 0.9 )
model.add_transition( m1, i1, 0.05 )
model.add_transition( m1, d2, 0.05 )
model.add_transition( m2, m3, 0.9 )
model.add_transition( m2, i2, 0.05 )
model.add_transition( m2, d3, 0.05 )
model.add_transition( m3, model.end, 0.9 )
model.add_transition( m3, i3, 0.1 )
# Create transitions from insert states
model.add_transition( i0, i0, 0.70 )
model.add_transition( i0, d1, 0.15 )
model.add_transition( i0, m1, 0.15 )
model.add_transition( i1, i1, 0.70 )
model.add_transition( i1, d2, 0.15 )
model.add_transition( i1, m2, 0.15 )
model.add_transition( i2, i2, 0.70 )
model.add_transition( i2, d3, 0.15 )
model.add_transition( i2, m3, 0.15 )
model.add_transition( i3, i3, 0.85 )
model.add_transition( i3, model.end, 0.15 )
# Create transitions from delete states
model.add_transition( d1, d2, 0.15 )
model.add_transition( d1, i1, 0.15 )
model.add_transition( d1, m2, 0.70 )
model.add_transition( d2, d3, 0.15 )
model.add_transition( d2, i2, 0.15 )
model.add_transition( d2, m3, 0.70 )
model.add_transition( d3, i3, 0.30 )
model.add_transition( d3, model.end, 0.70 )
# Call bake to finalize the structure of the model.
model.bake()
We can do somewhat trivial things which we've done before. However, we see that the model has a repeating structure of a match state, an insert state, and a delete state. With a 3-long example, it would be relatively simple to train each edge, but with even a 10-long example, there are many edges, and getting accurate measurements for each edge would be difficult.
Tied edges help to solve this problem by saying that going across certain edges is not a position-specific phenomena. For example, if a sequence aligns to i2 several times in a row, it's probably a phenomena more specific to insert states in general, than the second position in the model. All transitions from i2 to i2 now affect all self-loop insert transitions.
Lets look at some transitions first, before training.
indices = { state.name: i for i, state in enumerate( model.states ) }
transitions = model.dense_transition_matrix()
i0, i1, i2 = indices['I0'], indices['I1'], indices['I2']
d1, d2, d3 = indices['D1'], indices['D2'], indices['D3']
m1, m2, m3 = indices['M1'], indices['M2'], indices['M3']
print transitions[i0, i0], transitions[i1, i1], transitions[i2, i2]
print transitions[d1, d2], transitions[d2, d3]
print transitions[m1, m2], transitions[m2, m3]
-0.356674943939 -0.356674943939 -0.356674943939 -1.89711998489 -1.89711998489 -0.105360515658 -0.105360515658
Looks good. We didn't modify the model, and started it initiallly with same values for these.
Now lets train on some sequences.
sequences = [ list(x) for x in ( 'A', 'ACT', 'GGCA', 'TACCTGT', 'CA', 'AAAAAAAA', 'ACGTTTTTTTGG', 'ACG', 'GCACA' ) ]
model.train( sequences )
transitions = model.dense_transition_matrix()
print transitions[i0, i0], transitions[i1, i1], transitions[i2, i2]
print transitions[d1, d2], transitions[d2, d3]
print transitions[m1, m2], transitions[m2, m3]
Training improvement: -1.05389743843 Total Training Improvement: -1.05389743843 -0.671561297769 -0.363965082857 -0.367711513544 -1.06878154105 -0.425559960916 -0.524762602906 -0.466737198789
Very different now! It makes sense that the training is negative, because the sequences I randomly generated are poorly modelled by the HMM I created. I'm interested right now in looking at insertions and deletions (INDELS), which are possible but not likely under the model.
More importantly, these values seem to be different depending on position. i0 to i0 is not the same as i1 to i1.
This is good if you have enough training data to get accurate values for each position. If you have enough data, the subtle differences of these edges may yield useful information. However, more than likely, you don't have enough information to train each edge accurately, and an outlier may screw with the edges. You'd want to then tie the repeating edges, so that a transition across one counted as a transition across all of them, making these phenomena position-independent. Doing this is simple using YAHMM.
model = Model( "Global Alignment")
# Define the distribution for insertions
i_d = DiscreteDistribution( { 'A': 0.25, 'C': 0.25, 'G': 0.25, 'T': 0.25 } )
# Create the insert states
i0 = State( i_d, name="I0" )
i1 = State( i_d, name="I1" )
i2 = State( i_d, name="I2" )
i3 = State( i_d, name="I3" )
# Create the match states
m1 = State( DiscreteDistribution({ "A": 0.95, 'C': 0.01, 'G': 0.01, 'T': 0.02 }) , name="M1" )
m2 = State( DiscreteDistribution({ "A": 0.003, 'C': 0.99, 'G': 0.003, 'T': 0.004 }) , name="M2" )
m3 = State( DiscreteDistribution({ "A": 0.01, 'C': 0.01, 'G': 0.01, 'T': 0.97 }) , name="M3" )
# Create the delete states
d1 = State( None, name="D1" )
d2 = State( None, name="D2" )
d3 = State( None, name="D3" )
# Add all the states to the model
model.add_states( [i0, i1, i2, i3, m1, m2, m3, d1, d2, d3 ] )
# Create transitions from match states
model.add_transition( model.start, m1, 0.9 )
model.add_transition( model.start, i0, 0.1 )
model.add_transition( m1, m2, 0.9 )
model.add_transition( m1, i1, 0.05 )
model.add_transition( m1, d2, 0.05 )
model.add_transition( m2, m3, 0.9 )
model.add_transition( m2, i2, 0.05 )
model.add_transition( m2, d3, 0.05 )
model.add_transition( m3, model.end, 0.9 )
model.add_transition( m3, i3, 0.1 )
# Create transitions from insert states
model.add_transition( i0, i0, 0.70, group="i_a" )
model.add_transition( i0, d1, 0.15, group="i_b" )
model.add_transition( i0, m1, 0.15, group="i_c" )
model.add_transition( i1, i1, 0.70, group="i_a" )
model.add_transition( i1, d2, 0.15, group="i_b" )
model.add_transition( i1, m2, 0.15, group="i_c" )
model.add_transition( i2, i2, 0.70, group="i_a" )
model.add_transition( i2, d3, 0.15, group="i_b" )
model.add_transition( i2, m3, 0.15, group="i_c" )
model.add_transition( i3, i3, 0.85, group="i_a" )
model.add_transition( i3, model.end, 0.15 )
# Create transitions from delete states
model.add_transition( d1, d2, 0.15, group="d_a" )
model.add_transition( d1, i1, 0.15, group="d_b" )
model.add_transition( d1, m2, 0.70, group="d_c" )
model.add_transition( d2, d3, 0.15, group="d_a" )
model.add_transition( d2, i2, 0.15, group="d_b" )
model.add_transition( d2, m3, 0.70, group="d_c" )
model.add_transition( d3, i3, 0.30 )
model.add_transition( d3, model.end, 0.70 )
# Call bake to finalize the structure of the model.
model.bake()
Lets run the same code as before, before training.
indices = { state.name: i for i, state in enumerate( model.states ) }
transitions = model.dense_transition_matrix()
i0, i1, i2 = indices['I0'], indices['I1'], indices['I2']
d1, d2, d3 = indices['D1'], indices['D2'], indices['D3']
m1, m2, m3 = indices['M1'], indices['M2'], indices['M3']
print transitions[i0, i0], transitions[i1, i1], transitions[i2, i2]
print transitions[d1, d2], transitions[d2, d3]
print transitions[m1, m2], transitions[m2, m3]
-0.356674943939 -0.356674943939 -0.356674943939 -1.89711998489 -1.89711998489 -0.105360515658 -0.105360515658
Looks good. Now lets see what happens after training.
sequences = [ list(x) for x in ( 'A', 'ACT', 'GGCA', 'TACCTGT', 'CA', 'AAAAAAAA', 'ACGTTTTTTTGG', 'ACG', 'GCACA' ) ]
model.train( sequences )
transitions = model.dense_transition_matrix()
print transitions[i0, i0], transitions[i1, i1], transitions[i2, i2]
print transitions[d1, d2], transitions[d2, d3]
print transitions[m1, m2], transitions[m2, m3]
Training improvement: -1.15388597427 Total Training Improvement: -1.15388597427 -0.312637409339 -0.312637409339 -0.312637409339 -0.604783721038 -0.604783721038 -0.524762602906 -0.466737198789
Looks good! The first two lines are tied edges, while the third line are edges which are not tied. As expected, the first two lines are all the same number, while the third line is position-specific.
Package Information¶
YAHMM is an open source, free to use package. Installation is as easy as pip install yahmm, or cloning from the GitHub repo.
Comments? Questions? Open an issue on the GitHub page or email me.