Back: Whirl-torial¶
In [ ]:
from __future__ import division, print_function
In [ ]:
from IPython.display import HTML, display as disp, Audio
with open("../css/css.css", "r") as f:
styles = f.read()
HTML(styles)
In [ ]:
import os
import operator
import json
import tokenize
import re
from itertools import imap, ifilter, islice, cycle
from functools import partial
import sh
import requests
import matplotlib.pyplot as plt
import numpy as np
from numpy.random import randn, randint, rand, choice
import pandas as pd
from pandas import DataFrame, Series, Index
from pandas.compat import map, StringIO
import pandas.util.testing as tm
In [ ]:
pd.options.display.max_rows = 10
pd.options.display.max_columns = 7
try:
from mpltools import style
style.use('ggplot')
except ImportError:
pass
# because of our bg color
plt.rc('text', color='white')
plt.rc('axes', labelcolor='white')
plt.rc('xtick', color='white')
plt.rc('ytick', color='white')
In [ ]:
%matplotlib inline
In [ ]:
def insert_page(url):
"""Embed a webpage in the notebook"""
disp(HTML('<iframe src=%r width=700 height=350></iframe>' % url))
def read_text(*args, **kwargs):
"""Simple text reader because I don't like typing ``with`` every time"""
with open(*args, **kwargs) as f:
return f.read()
def highlight(filename, style='fruity'):
"""Syntax highlight a file based on its extension"""
from pygments import highlight as h
from pygments.lexers import guess_lexer_for_filename
from pygments.formatters import HtmlFormatter
code = read_text(filename, mode='rt')
formatter = HtmlFormatter(style=style)
lexer = guess_lexer_for_filename(filename, code)
disp(HTML('<style type="text/css">{0}</style>{1}'.format(
formatter.get_style_defs('.highlight'),
h(code, lexer, formatter))))
def gen_frames(n, size, f=randn):
"""Generate `n` frames of size `size` using the function `f`."""
return (DataFrame(f(*sz)) for sz in [size] * n)
New Features since v0.11.0¶
(or Interactive Release Notes)¶
numexpr speedups¶
More apparent for long(ish) expressions with large(ish) arrays¶
In [ ]:
x, y, z, w = gen_frames(4, size=(1e6, 20))
In [ ]:
def show_faster(num, denom):
ratio = num / denom
disp(HTML('numexpr is <b>%.2g</b>× as fast' % ratio))
def biggish():
disp(HTML('<b>biggish</b>'))
with tm.use_numexpr(True):
Y = %timeit -r 1 -n 1 -o x + y + z + w ** 3
with tm.use_numexpr(False):
N = %timeit -r 1 -n 1 -o x + y + z + w ** 3
show_faster(N.best, Y.best)
def smallish():
disp(HTML('<b>smallish</b>'))
with tm.use_numexpr(False):
Y = %timeit -r 1 -n 1 -o x + y
with tm.use_numexpr(False):
N = %timeit -r 1 -n 1 -o x + y
show_faster(N.best, Y.best)
biggish()
smallish()
In [ ]:
insert_page("http://www.fdic.gov/bank/individual/failed/banklist.html")
In [ ]:
url = '../data/banklist.html'
dfs = pd.read_html(url) # returns a list of all tables found on the page
In [ ]:
assert len(dfs) == 1, "you're wrong about me"
df = dfs.pop()
In [ ]:
# not sure where those extra columns are from ...
df
Select tables based on class¶
In [ ]:
dat_url = 'tmp.html'
with open(dat_url, 'w') as f:
DataFrame(randn(2, 2)).to_html(f, classes=['first'])
f.write('\n\n')
DataFrame(randn(2, 2)).to_html(f, classes=['second'])
In [ ]:
highlight(dat_url)
In [ ]:
df, = pd.read_html(dat_url, attrs={'class': 'first'}, index_col=0)
df
In [ ]:
dfs = pd.read_html(dat_url, index_col=0)
for df in dfs:
disp(df)
# not really a way to tell which table is which; ordered by appearance in HTML
In [ ]:
top_url = 'http://www.tylervigen.com'
url = 'http://www.tylervigen.com/view_correlation?id=1703'
In [ ]:
insert_page(top_url)
In [ ]:
insert_page(url)
In [ ]:
raw = requests.get(url).text
match = r'Divorce rate in Maine'
dfs = pd.read_html(raw, match=match, header=0, index_col=0)
In [ ]:
dfs[-1]
In [ ]:
# get rid of junk columns
df = dfs[-1].dropna(how='all', axis=(0, 1)).T
# better names
df.columns = ['mn_divorce_rate', 'per_capita_marg']
# rename generic index name to year
df = df.reset_index().rename(columns={'index': 'year'})
# make years integers
df = df.convert_objects(convert_numeric=True)
df
In [ ]:
def blacken_legend_text(leg):
for t in leg.get_texts():
t.set_color('k')
fig, (ax, ax2) = plt.subplots(2, 1, figsize=(8, 6))
# maine divorces
ln = ax.plot(df.mn_divorce_rate.values, r'ro-', label='Divorce Rate / 1000 People')
ax.set_xticklabels(df.year)
ax.set_xlabel('Year')
ax.set_ylabel(ln[0].get_label())
# butter eating
axt = ax.twinx()
lt = axt.plot(df.per_capita_marg.values, r'bo-', label='Per Capita Lbs of Margarine')
axt.set_ylabel(lt[0].get_label())
# scatter plot
ax2.scatter(df.mn_divorce_rate.values, df.per_capita_marg.values, s=100)
ax2.set_xlabel('MN Divorce Rate')
ax2.set_ylabel('Margarine')
ax2.set_title(r'Divorce vs. Margarine, $r = %.2g$' % df.mn_divorce_rate.corr(df.per_capita_marg))
ax2.axis('tight')
# legend madness
lns = ln + lt
leg = ax.legend(lns, [l.get_label() for l in lns], loc=0)
blacken_legend_text(leg)
fig.tight_layout()
DataFrame.replace() with regular expressions¶
In [ ]:
tips = pd.read_csv('s3://nyqpug/tips.csv')
In [ ]:
# add some random lower cased versions of yes and no
nrows = len(tips)
tips.loc[(rand(nrows) > 0.5) & (tips.smoker == 'Yes'), 'smoker'] = 'yes'
tips.loc[(rand(nrows) > 0.5) & (tips.smoker == 'No'), 'smoker'] = 'no'
In [ ]:
tips.smoker.value_counts().plot(kind='bar')
In [ ]:
# sanity check
tips.smoker.value_counts()
In [ ]:
repd = tips.replace(regex={'smoker': {'[yY]es': True, '[nN]o': False}})
repd
In [ ]:
repd_all = tips.replace(regex={'[yY]es': True, '[nN]o': False})
repd_all
In [ ]:
jsfile = 'data.json'
In [ ]:
%%writefile $jsfile
{
"name": ["Bob Jones", "Karen Smith"],
"age": [28, 26],
"gender": ["M", "F"]
}
In [ ]:
pd.read_json(jsfile) # no problemo
In [ ]:
# can also use keys as the rows instead of columns
pd.read_json(jsfile, orient='index')
In [ ]:
%%writefile $jsfile
{
"region": {
"Canada": {
"name": "Bob Jones",
"age": 28,
"gender": "M"
},
"USA": {
"name": "Karen Smith",
"age": 26,
"gender": "F"
}
}
}
In [ ]:
disp(pd.read_json(jsfile, orient='records'))
disp(Audio(os.path.join(os.pardir, 'mp3', 'w.mp3'), autoplay=True))
In [ ]:
# disp(Audio(os.path.join(os.pardir, 'mp3', 'c.mp3'), autoplay=True))
read_json (not so simple)¶
pandas plays nicely with other libraries¶
In [ ]:
data = read_text(jsfile)
In [ ]:
# avoid read_json entirely :)
# get transposed
df = DataFrame(json.loads(data)["region"])
df = df.T.convert_objects(convert_numeric=True)
df
In [ ]:
df.dtypes
In [ ]:
jq = sh.jq.bake('-M') # -M disables colorizing
In [ ]:
rule = "(.region)" # this rule is essentially data["region"]
out = jq(rule, _in=data).stdout
res = pd.read_json(out, orient='index')
res
In [ ]:
res.dtypes
Let's try something a bit hairier...¶
adapted from this StackOverflow question¶
In [ ]:
%%writefile $jsfile
{
"intervals": [
{
"pivots": "Jane Smith",
"series": [
{
"interval_id": 0,
"p_value": 1
},
{
"interval_id": 1,
"p_value": 1.1162791357932633e-8
},
{
"interval_id": 2,
"p_value": 0.0000028675012051504467
}
]
},
{
"pivots": "Bob Smith",
"series": [
{
"interval_id": 0,
"p_value": 1
},
{
"interval_id": 1,
"p_value": 1.1162791357932633e-8
},
{
"interval_id": 2,
"p_value": 0.0000028675012051504467
}
]
}
]
}
In [ ]:
%%writefile rule.txt
[{pivots: .intervals[].pivots,
interval_id: .intervals[].series[].interval_id,
p_value: .intervals[].series[].p_value}] | unique
In [ ]:
data = read_text(jsfile)
# check out http://stedolan.github.io/jq/manual for more details on these rules
rule = read_text('rule.txt')
out = jq(rule, _in=data).stdout
js = json.loads(out)
In [ ]:
js[:2]
In [ ]:
res = pd.read_json(out)
res
In [ ]:
res.dtypes
v0.13¶
DataFrame.isin()str.extract()- Experimental Features
query/evalmsgpackIOGoogle BigQueryIO
In [ ]:
names = list(filter(None, read_text('names.txt').split('\n')))
names
In [ ]:
df = DataFrame(dict(zip(['math', 'physics'],
[names[:5], names[-5:]])))
df
In [ ]:
df.isin(['Brook', 'Bradley', 'Richie', 'Sarah'])
str.extract()¶
In [ ]:
!grep -P '^[a-zA-Z_]\w*$' /usr/share/dict/cracklib-small | head -10
In [ ]:
def gen_filenames(n, pattern='%d_%s', dict_file='/usr/share/dict/words'):
matches_id = partial(re.match, '^%s$' % tokenize.Name)
interpolator = partial(operator.mod, pattern)
with open(dict_file, 'rt') as f:
only_valid_names = ifilter(matches_id, cycle(f))
n_matches = islice(only_valid_names, 0, n)
for el in imap(interpolator, enumerate(imap(str.strip, n_matches))):
yield el
In [ ]:
vids = Series(list(gen_filenames(30, pattern='%d_%s.mp4')))
vids
In [ ]:
ext = vids.str.extract('(?P<num>\d+)_(?P<name>.+)')
ext
In [ ]:
ext = ext.convert_objects(convert_numeric=True)
disp(ext.dtypes)
ext
v0.13 Experimental Features¶
query/eval- msgpack IO
- Google BigQuery IO
In [ ]:
n = 1e6
df = DataFrame({'a': randint(10, size=n),
'b': rand(n),
'c': rand(n)})
df.head()
In [ ]:
sub = df.query('1 <= a <= 5 and 0.1 < b < 0.4 and 0.5 <= c <= 0.95')
sub
In [ ]:
qtime = %timeit -o df.query('1 <= a <= 5 and 0.1 < b < 0.4 and 0.5 <= c <= 0.95')
pytime = %timeit -o df.loc[(1 <= df.a) & (df.a <= 5) & (0.1 <= df.b) & (df.b <= 0.4) & (0.5 <= df.c) & (df.c <= 0.9)]
print('query is %.2gx faster than pure Python' % (pytime.best / qtime.best))
In [ ]:
A, B, C, D = (DataFrame(randn(n, 40)) for _ in range(4))
In [ ]:
qtime = %timeit -r 1 -n 1 -o pd.eval('A + B * 2 + C / D ** 3 * B / C + A ** 10 < A ** 5')
In [ ]:
pytime = %timeit -r 1 -n 1 -o A + B * 2 + C / D ** 3 * B / C + A ** 10 < A ** 5
print('query is %.2gx faster than pure Python' % (pytime.best / qtime.best))
Local variables¶
In [ ]:
a = rand()
df.query('a <= @a <= b')
MessagePack IO (to_msgpack/read_msgpack)¶
In [ ]:
df.head(2).to_msgpack()
In [ ]:
s = pd.to_msgpack(None, # we want the raw bytes output so pass None
Series(randn(2)),
['yep', 'a', 'list'],
randn(2),
{'a': 2, 'b': 3})
sio = StringIO(s)
pd.read_msgpack(sio)
Google BigQuery IO¶
In [ ]:
highlight('query.sql')
In [ ]:
query = read_text('query.sql')
In [ ]:
df = pd.read_gbq(query, project_id='metal-lantern-572')
Notice the NaTs and NaNs. Those are where other repositories have valid pull request dates
In [ ]:
df = df.rename(columns=lambda x: x.replace('payload_pull_request_', ''))
In [ ]:
df.dtypes
df
In [ ]:
df['created_at'] = pd.to_datetime(df.created_at)
In [ ]:
df
In [ ]:
# set the index to the datetime column just created
df = df.set_index('created_at').sort_index()
df
In [ ]:
s = df.additions
In [ ]:
def remove_time(ax):
replacer = lambda x: x.get_text().replace(' 00:00:00', '')
ax.set_xticklabels(list(map(replacer, ax.get_xticklabels())))
In [ ]:
r = s.resample('B', how='sum')
r.index.name = 'Pull Request Day'
ax = r.plot(kind='bar', figsize=(18, 5))
remove_time(ax)
ax.set_ylabel('Pull Request Additions per Business Day')
ax.get_figure().autofmt_xdate()
Non user facing but worth mentioning:¶
Jeff Reback's refactor of Series to use composition instead of inheriting from numpy.ndarray. Bravo!¶
v0.14 (soon to be released)¶
MultiIndexslicingnlargest/nsmallest- hexbin, pie, and table plotting
Prelude to MultiIndex slicing¶
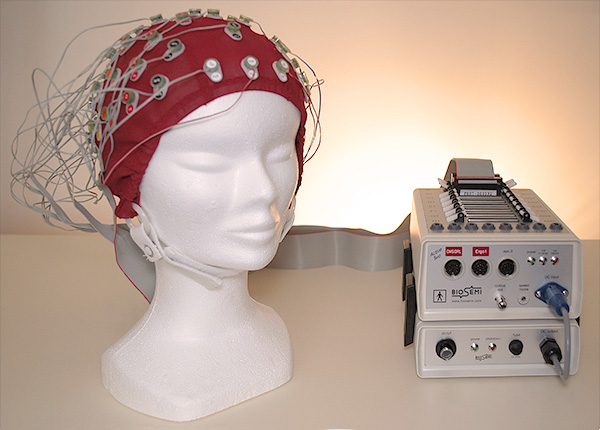

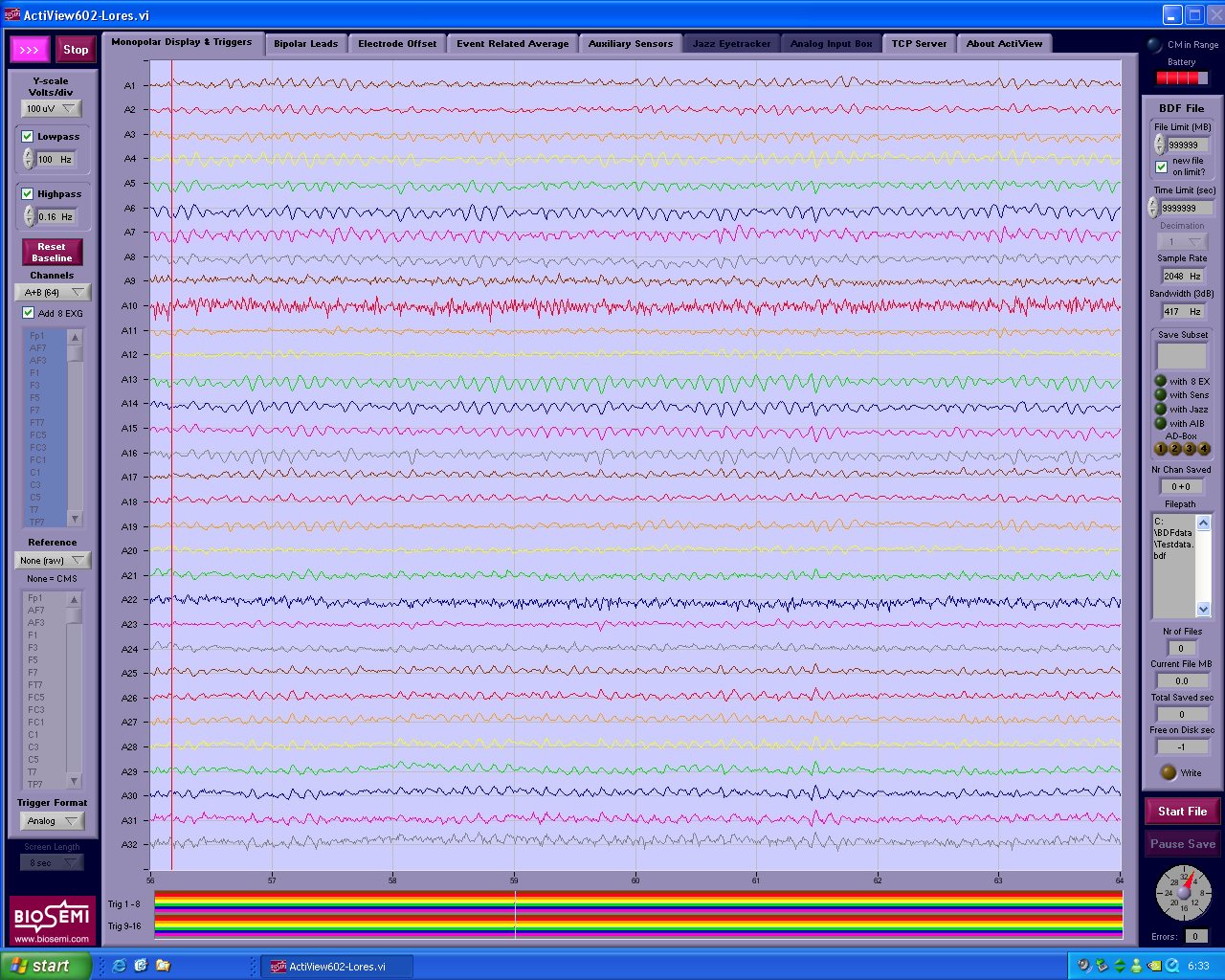
MultiIndex Slicing¶
In [ ]:
def channel_index(name, nchannels):
return list(zip(np.repeat(name, nchannels), range(nchannels)))
In [ ]:
# simulate our EEG data set
fs = 256 # sampling rate
neeg = 8 # number of EEG channels
nex = 4 # number of auxiliary channels
nsensors = 2 # number of gsr channels
eeg_chan = channel_index('eeg', neeg)
ex_chan = channel_index('ex', nex)
sens_chan = channel_index('gsr', nsensors)
disp(eeg_chan)
disp(ex_chan)
disp(sens_chan)
In [ ]:
columns = pd.MultiIndex.from_tuples(eeg_chan + ex_chan + sens_chan,
names=['signal', 'channel'])
# 10 seconds of fake data
df = pd.DataFrame(np.random.randn(fs * 10, columns.labels[0].size), columns=columns)
# add in some nans (e.g., a person moved around during these samples)
df.loc[rand(len(df)) < 0.20, 'eeg'] = np.nan
df.head()
In [ ]:
# simulate a stimulus marker
df['stim'] = np.sort(randint(10, size=len(df)))
df
In [ ]:
df.loc[:, np.s_[('ex', 'stim'), :]].head()
In [ ]:
# the EX and STIM channels where EEG channels 0 and 1 are not null
row_idx = df.eeg[[0, 1]].notnull().all(axis=1)
col_idx = np.s_[('ex', 'stim'), :]
In [ ]:
col_idx
In [ ]:
res = df.loc[row_idx, col_idx]
res
In [ ]:
# use np.s_ to construct slices
assert slice(None, 1000, 2) == np.s_[:1000:2] # Which would you prefer?
assert slice(900, None, -1) == np.s_[900::-1]
The expression
np.s_[('ex', 'stim'), :]
says select 'ex' and 'stim' from the first level of the MultiIndex and the : as saying give me all columns from the second level
nlargest/nsmallest¶
If $n << $ len(s) you'll see a performance improvement.¶
Idea: Don't sort the whole array when you only need 5 values because¶
In [ ]:
s = Series(randn(1000000), name='a')
In [ ]:
a = %timeit -o s.nlargest(5) # 5 << 1,000,000
b = %timeit -o s.order(ascending=False).head()
print('nlargest is %.2gx faster than order + head' % (b.best / a.best))
In [ ]:
a = %timeit -o s.nsmallest(5)
b = %timeit -o s.order().head()
print('nsmallest is %.2gx faster than order + head' % (b.best / a.best))
