
** DISCLAIMER:** This exercise is for illustrative purposes and only uses about 500 samples which is too small for a generalizable model. ### References
- https://developer.apple.com/library/mac/documentation/DeveloperTools/Conceptual/MachORuntime/Reference/reference.html
- https://developer.apple.com/library/mac/documentation/DeveloperTools/Conceptual/MachORuntime/Mach-O_File_Format.pdf
- http://en.wikipedia.org/wiki/Mach-O
- macholib: https://pythonhosted.org/macholib/
- Pandas: Python Data Analysis Library (http://pandas.pydata.org)
- Scikit Learn (http://scikit-learn.org) Scikit-learn: Machine Learning in Python, Pedregosa et al., JMLR 12, pp. 2825-2830, 2011.
- Matplotlib: Python 2D plotting library (http://matplotlib.org)
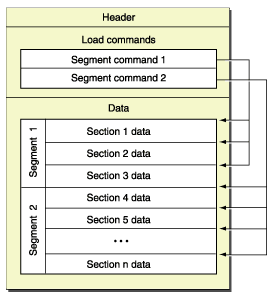
- Header Structure
- Load Commands
- Segments
- The initial layout of the file in virtual memory
- The location of the symbol table (used for dynamic linking)
- The initial execution state of the main thread of the program
- The names of shared libraries that contain definitions for the main executable’s imported symbols
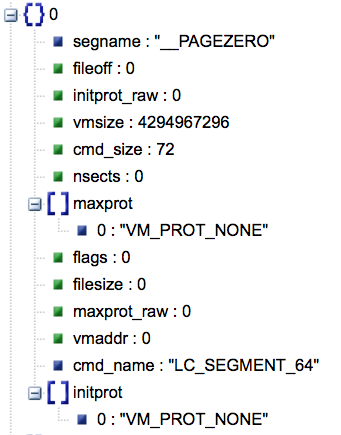
Description: macholib can be used to analyze and edit Mach-O headers, the executable format used by Mac OS X.
Wasn't the easiest to get data from each load command and other data I was interested in. So we forked repo and added that ability.
https://bitbucket.org/trogdorsey/macholib
Imports and plot defaults¶
In [1]:
import pandas as pd
print 'pandas version is', pd.__version__
import numpy as np
print 'numpy version is', np.__version__
import sklearn
print 'scikit-learn version is', sklearn.__version__
import matplotlib
print 'matplotlib version is', matplotlib.__version__
import matplotlib.pyplot as plt
pandas version is 0.13.1 numpy version is 1.8.1 scikit-learn version is 0.14.1 matplotlib version is 1.3.1
Plotting defaults and helper functions¶
In [2]:
%matplotlib inline
plt.rcParams['font.size'] = 18.0
plt.rcParams['figure.figsize'] = 16.0, 5.0
In [3]:
def plot_cm(cm, labels):
# Compute percentanges
percent = (cm*100.0)/np.array(np.matrix(cm.sum(axis=1)).T)
print 'Confusion Matrix Stats'
for i, label_i in enumerate(labels):
for j, label_j in enumerate(labels):
print "%s/%s: %.2f%% (%d/%d)" % (label_i, label_j, (percent[i][j]), cm[i][j], cm[i].sum())
# Show confusion matrix
# Thanks to kermit666 from stackoverflow
fig = plt.figure()
ax = fig.add_subplot(111)
ax.grid(b=False)
cax = ax.matshow(percent, cmap='coolwarm',vmin=0,vmax=100)
plt.title('Confusion matrix of the classifier')
fig.colorbar(cax)
ax.set_xticklabels([''] + labels)
ax.set_yticklabels([''] + labels)
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
Function to extract out the features we are interested with from the json blob¶
In [4]:
def extract_features(data):
all_features = []
if not 'error' in data['characteristics']['macho']:
for i in range(data['characteristics']['macho']['number of architectures']):
features = {}
features['entropy'] = data['metadata']['entropy']
features['file_size'] = data['metadata']['file_size']
features['number of architectures'] = data['characteristics']['macho']['number of architectures']
features['header size'] = data['characteristics']['macho']['header'][i]['size']
features['cmd count'] = data['characteristics']['macho']['header'][i]['mach header']['ncmds']
features['cmd size'] = data['characteristics']['macho']['header'][i]['mach header']['sizeofcmds']
features['cputype'] = data['characteristics']['macho']['header'][i]['mach header']['cputype_string']
for flag in data['characteristics']['macho']['header'][i]['mach header']['flags']:
features[flag['name']] = 1
features['string count'] = data['verbose']['macho']['header'][i]['const strings count']
if 'const strings' in data['verbose']['macho']['header'][i]:
features['const strings'] = data['verbose']['macho']['header'][i]['const strings']
else:
features['const strings'] = []
if 'symbol table strings' in data['verbose']['macho']['header'][i]:
features['symbol table strings'] = data['verbose']['macho']['header'][i]['symbol table strings']
else:
features['symbol table strings'] = []
features['segment names'] = []
features['section names'] = []
for command in data['verbose']['macho']['header'][i]['commands']:
if command['cmd_name'] in ['LC_SEGMENT', 'LC_SEGMENT_64']:
features['segment names'].append(command['segname'])
if command['segname'] == '__PAGEZERO':
features['pz_size'] = command['cmd_size']
features['pz_vmsize'] = command['vmsize']
features['pz_vmaddr'] = command['vmaddr']
features['pz_flags'] = command['flags']
features['pz_filesize'] = command['filesize']
features['pz_nsects'] = command['nsects']
features['pz_fileoff'] = command['fileoff']
for flag in command['initprot']:
features['pz_initprot_'+flag] = 1
for flag in command['maxprot']:
features['pz_maxprot_'+flag] = 1
if command['segname'] == '__TEXT':
features['text_size'] = command['cmd_size']
features['text_vmsize'] = command['vmsize']
features['text_vmaddr'] = command['vmaddr']
features['text_flags'] = command['flags']
features['text_filesize'] = command['filesize']
features['text_nsects'] = command['nsects']
features['text_fileoff'] = command['fileoff']
for flag in command['initprot']:
features['text_initprot_'+flag] = 1
for flag in command['maxprot']:
features['text_maxprot_'+flag] = 1
features['text_entropy'] = command['entropy']
for section in command['sections']:
if section['sectname'] == '__text':
features['text_section_reloff'] = section['reloff']
features['text_section_addr'] = section['addr']
features['text_section_align'] = section['align']
features['text_section_nreloc'] = section['nreloc']
features['text_section_offset'] = section['offset']
features['text_section_size'] = section['size']
features['text_section_reserved1'] = section['reserved1']
features['text_section_reserved2'] = section['reserved2']
features['text_section_' + section['flags']['type']] = 1
if 'attributes' in section['flags']:
for attr in section['flags']['attributes']:
features['text_section_flag_attribute_'+attr] = 1
if section['sectname'] == '__const':
features['const_section_reloff'] = section['reloff']
features['const_section_addr'] = section['addr']
features['const_section_align'] = section['align']
features['const_section_nreloc'] = section['nreloc']
features['const_section_offset'] = section['offset']
features['const_section_size'] = section['size']
features['const_section_reserved1'] = section['reserved1']
features['const_section_reserved2'] = section['reserved2']
features['const_section_' + section['flags']['type']] = 1
if 'attributes' in section['flags']:
for attr in section['flags']['attributes']:
features['const_section_flag_attribute_'+attr] = 1
if command['segname'] == '__DATA' and command['nsects'] > 0:
features['data_size'] = command['cmd_size']
features['data_vmsize'] = command['vmsize']
features['data_vmaddr'] = command['vmaddr']
features['data_flags'] = command['flags']
features['data_filesize'] = command['filesize']
features['data_nsects'] = command['nsects']
features['data_fileoff'] = command['fileoff']
for flag in command['initprot']:
features['data_initprot_'+flag] = 1
for flag in command['maxprot']:
features['data_maxprot_'+flag] = 1
features['data_entropy'] = command['entropy']
#if command['segname'] == '__IMPORT':
# features['import_size'] = command['cmd_size']
# features['import_vmsize'] = command['vmsize']
# features['import_vmaddr'] = command['vmaddr']
# features['import_flags'] = command['flags']
# features['import_filesize'] = command['filesize']
# features['import_nsects'] = command['nsects']
# features['import_fileoff'] = command['fileoff']
# for flag in command['initprot']:
# features['import_initprot_'+flag] = 1
# for flag in command['maxprot']:
# features['import_maxprot_'+flag] = 1
# features['import_entropy'] = command['entropy']
if command['segname'] == '__LINKEDIT':
features['linkedit_size'] = command['cmd_size']
features['linkedit_vmsize'] = command['vmsize']
features['linkedit_vmaddr'] = command['vmaddr']
features['linkedit_flags'] = command['flags']
features['linkedit_filesize'] = command['filesize']
features['linkedit_nsects'] = command['nsects']
features['linkedit_fileoff'] = command['fileoff']
for flag in command['initprot']:
features['linkedit_initprot_'+flag] = 1
for flag in command['maxprot']:
features['linkedit_maxprot_'+flag] = 1
if 'sections' in command:
for section in command['sections']:
features['section names'].append(section['sectname'])
if command['cmd_name'] == 'LC_SYMTAB':
features['strsize'] = command['strsize']
features['stroff'] = command['stroff']
features['symoff'] = command['symoff']
features['nsyms'] = command['nsyms']
if command['cmd_name'] in ['LC_DYLD_INFO_ONLY', 'LC_DYLD_INFO']:
features['lazy_bind_size'] = command['lazy_bind_size']
features['rebase_size'] = command['rebase_size']
features['weak_bind_size'] = command['weak_bind_size']
features['lazy_bind_off'] = command['lazy_bind_off']
features['export_off'] = command['export_off']
features['export_size'] = command['export_size']
features['bind_off'] = command['bind_off']
features['rebase_off'] = command['rebase_off']
features['bind_size'] = command['bind_size']
features['weak_bind_off'] = command['weak_bind_off']
if command['cmd_name'] == 'LC_DYSYMTAB':
features['nextdefsym'] = command['nextdefsym']
features['extreloff'] = command['extreloff']
features['nlocrel'] = command['nlocrel']
features['indirectsymoff'] = command['indirectsymoff']
features['modtaboff'] = command['modtaboff']
features['iundefsym'] = command['iundefsym']
features['ntoc'] = command['ntoc']
features['ilocalsym'] = command['ilocalsym']
features['nundefsym'] = command['nundefsym']
features['nextrefsyms'] = command['nextrefsyms']
features['locreloff'] = command['locreloff']
features['nmodtab'] = command['nmodtab']
features['nlocalsym'] = command['nlocalsym']
features['tocoff'] = command['tocoff']
features['extrefsymoff'] = command['extrefsymoff']
features['nindirectsyms'] = command['nindirectsyms']
features['iextdefsym'] = command['iextdefsym']
features['nextrel'] = command['nextrel']
features.update(data['verbose']['macho']['header'][i]['command type count'])
if 'LC_SEGMENT' in features:
features['number of segments'] = features['LC_SEGMENT']
else:
features['number of segments'] = features['LC_SEGMENT_64']
if 'entry point' in data['verbose']['macho']['header'][i]:
for key, value in data['verbose']['macho']['header'][i]['entry point'].iteritems():
features['entry point ' + key] = value
all_features.append(features)
return all_features
In [5]:
def load_files(file_list):
import json
features_list = []
for filename in file_list:
with open(filename,'rb') as f:
features = extract_features(json.loads(f.read()))
features_list.extend(features)
return features_list
The set of good files came from my MacBook Pro running Mavericks.¶
In [6]:
# Good files
import glob
good_list = glob.glob('data/good/*.results')
good_features = load_files(good_list)
print "Files:", len(good_list)
print "Number of feature vectors:", len(good_features)
Files: 266 Number of feature vectors: 337
The set of bad files came from Contagio and VirusTotal¶
In [7]:
# Bad files
bad_list = glob.glob('data/bad/*.results')
bad_features = load_files(bad_list)
print "Files:", len(bad_list)
print "Number of feature vectors:", len(bad_features)
Files: 261 Number of feature vectors: 302

There are 3 (or 6 when you take into consideration endianness) different magic values for MachO binaries. One (two) for 32-bit binaries, one (two) for 64-bit binaries, and one (two) for fat binaries.
The magic value for fat binaries is cafebabe, which you might recognize as the magic value for java class files.
In [8]:
df_good_orig = pd.DataFrame.from_records(good_features)
df_good_orig['label'] = 'good'
df_good_orig.head()
Out[8]:
| LC_CODE_SEGMENT_SPLIT_INFO | LC_CODE_SIGNATURE | LC_DATA_IN_CODE | LC_DYLD_INFO | LC_DYLD_INFO_ONLY | LC_DYLIB_CODE_SIGN_DRS | LC_DYSYMTAB | LC_FUNCTION_STARTS | LC_ID_DYLIB | LC_LOAD_DYLIB | LC_LOAD_DYLINKER | LC_LOAD_WEAK_DYLIB | LC_MAIN | LC_REEXPORT_DYLIB | LC_RPATH | LC_SEGMENT | LC_SEGMENT_64 | LC_SOURCE_VERSION | LC_SYMTAB | LC_UNIXTHREAD | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | NaN | 1 | 1 | NaN | 1 | 1 | 1 | 1 | NaN | 1 | 1 | NaN | 1 | NaN | NaN | NaN | 4 | 1 | 1 | NaN | ... |
| 1 | NaN | 1 | 1 | NaN | 1 | 1 | 1 | 1 | NaN | 9 | 1 | NaN | 1 | NaN | NaN | NaN | 4 | 1 | 1 | NaN | ... |
| 2 | NaN | 1 | 1 | NaN | 1 | 1 | 1 | 1 | NaN | 9 | 1 | NaN | 1 | NaN | NaN | 4 | NaN | 1 | 1 | NaN | ... |
| 3 | NaN | 1 | 1 | NaN | 1 | 1 | 1 | 1 | NaN | 5 | 1 | NaN | 1 | NaN | NaN | NaN | 4 | 1 | 1 | NaN | ... |
| 4 | NaN | 1 | 1 | NaN | 1 | 1 | 1 | 1 | NaN | 1 | 1 | NaN | 1 | NaN | NaN | NaN | 4 | 1 | 1 | NaN | ... |
5 rows × 148 columns
In [9]:
df_good = df_good_orig
for col in df_good.columns:
if col[0:3] in ['LC_', 'MH_']:
#print col
df_good[col].fillna(0, inplace=True)
df_good.fillna(-1, inplace=True)
df_good_orig.head()
Out[9]:
| LC_CODE_SEGMENT_SPLIT_INFO | LC_CODE_SIGNATURE | LC_DATA_IN_CODE | LC_DYLD_INFO | LC_DYLD_INFO_ONLY | LC_DYLIB_CODE_SIGN_DRS | LC_DYSYMTAB | LC_FUNCTION_STARTS | LC_ID_DYLIB | LC_LOAD_DYLIB | LC_LOAD_DYLINKER | LC_LOAD_WEAK_DYLIB | LC_MAIN | LC_REEXPORT_DYLIB | LC_RPATH | LC_SEGMENT | LC_SEGMENT_64 | LC_SOURCE_VERSION | LC_SYMTAB | LC_UNIXTHREAD | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 0 | 0 | 0 | 4 | 1 | 1 | 0 | ... |
| 1 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 9 | 1 | 0 | 1 | 0 | 0 | 0 | 4 | 1 | 1 | 0 | ... |
| 2 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 9 | 1 | 0 | 1 | 0 | 0 | 4 | 0 | 1 | 1 | 0 | ... |
| 3 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 5 | 1 | 0 | 1 | 0 | 0 | 0 | 4 | 1 | 1 | 0 | ... |
| 4 | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 0 | 0 | 0 | 4 | 1 | 1 | 0 | ... |
5 rows × 148 columns
In [10]:
df_bad_orig = pd.DataFrame.from_records(bad_features)
df_bad_orig['label'] = 'bad'
df_bad = df_bad_orig
for col in df_bad.columns:
if col[0:3] in ['LC_', 'MH_']:
#print col
df_bad[col].fillna(0, inplace=True)
df_bad.fillna(-1, inplace=True)
df_bad.head()
Out[10]:
| LC_CODE_SIGNATURE | LC_DATA_IN_CODE | LC_DYLD_INFO | LC_DYLD_INFO_ONLY | LC_DYSYMTAB | LC_ENCRYPTION_INFO | LC_FUNCTION_STARTS | LC_ID_DYLIB | LC_LAZY_LOAD_DYLIB | LC_LOAD_DYLIB | LC_LOAD_DYLINKER | LC_SEGMENT | LC_SEGMENT_64 | LC_SYMTAB | LC_TWOLEVEL_HINTS | LC_UNIXTHREAD | LC_UUID | LC_VERSION_MIN_IPHONEOS | LC_VERSION_MIN_MACOSX | MH_BINDATLOAD | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 8 | 1 | 0 | 4 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | ... |
| 1 | 0 | 0 | 1 | 0 | 1 | 0 | 0 | 1 | 0 | 8 | 0 | 0 | 3 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | ... |
| 2 | 1 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 2 | 1 | 0 | 4 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | ... |
| 3 | 1 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 2 | 1 | 4 | 0 | 1 | 0 | 1 | 1 | 0 | 0 | 0 | ... |
| 4 | 1 | 1 | 0 | 1 | 1 | 0 | 1 | 0 | 0 | 11 | 0 | 4 | 0 | 1 | 0 | 0 | 1 | 0 | 1 | 0 | ... |
5 rows × 153 columns
Let's explore the data.¶
In [11]:
df_good['cputype'].value_counts().plot(kind='bar')
Out[11]:
<matplotlib.axes.AxesSubplot at 0x112d9ec90>
In [12]:
df_bad['cputype'].value_counts().plot(kind='bar')
Out[12]:
<matplotlib.axes.AxesSubplot at 0x112db1cd0>
Let's get rid of the PowerPC cputype because that switch happened 9 years ago. I think we can all agree that malware authors won't be targeting that CPU in the future. And we probably want to examine malware for iOS devices separately.
In [13]:
df_all_orig = pd.concat([df_bad, df_good], ignore_index=True)
df_all = df_all_orig
for col in df_all.columns:
if col[0:3] in ['LC_', 'MH_']:
#print col
df_all[col].fillna(0, inplace=True)
df_all.fillna(-1, inplace=True)
# Break out by cpu type
cond = df_all['cputype'] == 'x86_64'
df_x64 = df_all[cond]
cond = df_all['cputype'] == 'i386'
df_x86 = df_all[cond]
df_all = pd.concat([df_x64, df_x86], ignore_index=True)
#print df['symbol table strings']
df = df_all.drop(['const strings', 'symbol table strings', 'segment names',
'section names', 'entry point section name', 'entry point segment name',
'entry point instruction type'], axis=1)
df['cputype'].value_counts().plot(kind='bar')
Out[13]:
<matplotlib.axes.AxesSubplot at 0x112fe7b50>
Let's start to blindly explore the data¶
In [14]:
cond = df['label'] == 'good'
good = df[cond]
bad = df[~cond]
bad['cmd count'].hist(alpha=.5,label='bad',bins=40)
good['cmd count'].hist(alpha=.5,label='good',bins=40)
plt.legend()
Out[14]:
<matplotlib.legend.Legend at 0x11338d1d0>
In [15]:
df.boxplot('cmd count', 'label')
plt.xlabel('bad vs. good files')
plt.ylabel('Number of Commands')
plt.title('Comparision of Number of Commands')
plt.suptitle('')
Out[15]:
<matplotlib.text.Text at 0x1135dca90>
In [16]:
df.boxplot(column='cmd size', by='label')
plt.xlabel('bad vs. good files')
plt.ylabel('Command Size')
plt.title('Comparision of Command Sizes')
plt.suptitle("")
Out[16]:
<matplotlib.text.Text at 0x11374e810>
In [17]:
# Split the classes up so we can set colors, size, labels
cond = df['label'] == 'good'
good = df[cond]
bad = df[~cond]
plt.scatter(good['cmd count'], good['cmd size'],
s=140, c='#aaaaff', label='Good', alpha=.4)
plt.scatter(bad['cmd count'], bad['cmd size'],
s=40, c='r', label='Bad', alpha=.5)
plt.legend()
plt.xlabel('Command Count')
plt.ylabel('Command Size')
plt.title('Command Size Vs Command Count')
Out[17]:
<matplotlib.text.Text at 0x1138d4690>
In [18]:
df.boxplot('entropy', 'label')
plt.xlabel('bad vs. good files')
plt.ylabel('Number of Commands')
plt.title('Comparision of Entropy')
plt.suptitle('')
Out[18]:
<matplotlib.text.Text at 0x113519b90>
In [19]:
df.boxplot('number of segments', 'label')
plt.xlabel('bad vs. good files')
plt.ylabel('Number of Segments')
plt.title('Comparision of Number of Segments')
plt.suptitle('')
Out[19]:
<matplotlib.text.Text at 0x11412ce50>
In [20]:
cond = df['label'] == 'good'
good = df[cond]
bad = df[~cond]
plt.scatter(good['entropy'], good['number of segments'],
s=140, c='#aaaaff', label='Good', alpha=.4)
plt.scatter(bad['entropy'], bad['number of segments'],
s=40, c='r', label='Bad', alpha=.5)
plt.legend()
plt.xlabel('Entropy')
plt.ylabel('Number of Segments')
plt.title('Number of Segments Vs Entropy')
Out[20]:
<matplotlib.text.Text at 0x1142b6790>
First we try classifying binaries just based on cmd count and cmd size¶
In [21]:
# List of feature vectors (scikit learn uses 'X' for the matrix of feature vectors)
X_cmd = df.as_matrix(['cmd count', 'cmd size'])
# Labels (scikit learn uses 'y' for classification labels)
y = np.array(df['label'].tolist())
In [22]:
# Random Forest is a popular ensemble machine learning classifier.
# http://scikit-learn.org/dev/modules/generated/sklearn.ensemble.RandomForestClassifier.html
#
import sklearn.ensemble
clf_cmd = sklearn.ensemble.RandomForestClassifier(n_estimators=50)
In [23]:
# Now we can use scikit learn's cross validation to assess predictive performance.
scores = sklearn.cross_validation.cross_val_score(clf_cmd, X_cmd, y, cv=10)
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
Accuracy: 0.89 (+/- 0.07)
In [31]:
my_seed = 1022
my_tsize = .2
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_cmd, y, test_size=my_tsize, random_state=my_seed)
clf_cmd.fit(X_train, y_train)
clf_cmd_scores = clf_cmd.score(X_test, y_test)
print("Accuracy: %0.2f" % clf_cmd_scores)
y_pred = clf_cmd.predict(X_test)
from sklearn.metrics import confusion_matrix
labels = ['good', 'bad']
cm = confusion_matrix(y_test, y_pred, labels)
plot_cm(cm, labels)
Accuracy: 0.91 Confusion Matrix Stats good/good: 92.75% (64/69) good/bad: 7.25% (5/69) bad/good: 11.63% (5/43) bad/bad: 88.37% (38/43)
Now try number of segments and entropy. This shouldn't work all that great.¶
In [32]:
clf_2 = sklearn.ensemble.RandomForestClassifier(n_estimators=50)
X_2 = df.as_matrix(['number of segments', 'entropy'])
X_train, X_test, y_train, y_test = train_test_split(X_2, y, test_size=my_tsize, random_state=my_seed)
clf_2.fit(X_train, y_train)
y_pred = clf_2.predict(X_test)
from sklearn.metrics import confusion_matrix
labels = ['good', 'bad']
cm = confusion_matrix(y_test, y_pred, labels)
plot_cm(cm, labels)
Confusion Matrix Stats good/good: 73.91% (51/69) good/bad: 26.09% (18/69) bad/good: 18.60% (8/43) bad/bad: 81.40% (35/43)
Okay now try putting in ALL the features... except the label, which would be cheating. And the cputype, which is a string.¶
In [33]:
clf_all = sklearn.ensemble.RandomForestClassifier(n_estimators=50)
no_label = list(df.columns.values)
no_label.remove('label')
no_label.remove('cputype')
X = df.as_matrix(no_label)
# 80/20 Split for predictive test
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=my_tsize, random_state=my_seed)
clf_all.fit(X_train, y_train)
y_pred = clf_all.predict(X_test)
cm = confusion_matrix(y_test, y_pred, labels)
plot_cm(cm, labels)
Confusion Matrix Stats good/good: 92.75% (64/69) good/bad: 7.25% (5/69) bad/good: 6.98% (3/43) bad/bad: 93.02% (40/43)
In [34]:
# Feature Selection
# Which features best deferentiated the two classes?
# Here we're going to grab the feature_importances from the classifier itself,
importances = zip(no_label, clf_all.feature_importances_)
importances.sort(key=lambda k:k[1], reverse=True)
for im in importances[0:20]:
print im[0].ljust(30), im[1]
LC_UNIXTHREAD 0.11111604246 LC_CODE_SIGNATURE 0.0500117721965 rebase_off 0.0407851853834 LC_MAIN 0.0385643285089 text_section_align 0.0339347105743 LC_DYLIB_CODE_SIGN_DRS 0.0335313105805 LC_DYLD_INFO_ONLY 0.0304556902729 linkedit_filesize 0.029905256752 MH_PIE 0.0226196128034 LC_SOURCE_VERSION 0.0198478375291 pz_size 0.0178616171746 rebase_size 0.017668781554 text_section_addr 0.0159335856278 data_entropy 0.0156907971405 LC_FUNCTION_STARTS 0.0150161007743 data_size 0.0149447133839 linkedit_vmsize 0.014562058373 const_section_size 0.0135111773331 extreloff 0.0132383705557 LC_VERSION_MIN_MACOSX 0.0127062868088
Closer look at important features¶
The LC_UNIXTHREAD command defines the entry point into the executable. In 10.8 and above, this was replaced by LC_MAIN. As stated above, I grabbed my clean binaries from my laptop, which is running Mavericks (10.9). The malware was grabbed from Contagio and VirusTotal, with no thought as to what version it was compiled on.In [35]:
df_good['LC_UNIXTHREAD'].value_counts().plot(kind='bar', label='good')
plt.legend()
Out[35]:
<matplotlib.legend.Legend at 0x11410c9d0>
In [36]:
df_bad['LC_UNIXTHREAD'].value_counts().plot(kind='bar')
Out[36]:
<matplotlib.axes.AxesSubplot at 0x1155239d0>
My clean set clearly has a sample bias towards LC_MAIN. We do not want to overfit our model and since we would like our classifier to be agnostic to OS version, we will remove those labels.
Over fitting problems: https://www.youtube.com/watch?v=DQWI1kvmwRg
Over fitting problems: https://www.youtube.com/watch?v=DQWI1kvmwRg
Now try all the same features as before, but without LC_UNIXTHREAD and LC_MAIN¶
In [42]:
clf_some = sklearn.ensemble.RandomForestClassifier(n_estimators=50)
some = list(df.columns.values)
some.remove('label')
some.remove('cputype')
some.remove('LC_UNIXTHREAD')
some.remove('LC_MAIN')
X_some = df.as_matrix(some)
# 80/20 Split for predictive test
X_train_some, X_test_some, y_train_some, y_test_some = train_test_split(X_some, y, test_size=my_tsize, random_state=my_seed)
clf_some.fit(X_train_some, y_train_some)
y_pred_some = clf_some.predict(X_test_some)
cm = confusion_matrix(y_test_some, y_pred_some, labels)
plot_cm(cm, labels)
Confusion Matrix Stats good/good: 95.65% (66/69) good/bad: 4.35% (3/69) bad/good: 6.98% (3/43) bad/bad: 93.02% (40/43)
Hey, it's the same(ish). That's kinda nice.
In [43]:
importances = zip(some, clf_some.feature_importances_)
importances.sort(key=lambda k:k[1], reverse=True)
for im in importances[0:20]:
print im[0].ljust(30), im[1]
rebase_off 0.0833325758996 text_section_align 0.0428642619275 LC_SOURCE_VERSION 0.0414863657063 LC_DYLIB_CODE_SIGN_DRS 0.0390404405528 linkedit_filesize 0.0343569215656 rebase_size 0.0268211203307 LC_FUNCTION_STARTS 0.0258331878885 LC_DATA_IN_CODE 0.0257389166901 LC_VERSION_MIN_MACOSX 0.0248798919781 LC_CODE_SIGNATURE 0.0231596346234 pz_size 0.0230647740647 LC_DYLD_INFO_ONLY 0.0227836821431 MH_PIE 0.0197788472089 data_nsects 0.0193926037336 pz_vmaddr 0.0180289427176 lazy_bind_size 0.0156826479795 linkedit_vmsize 0.0142413695731 cmd count 0.0137541943859 nlocrel 0.0130961872487 bind_off 0.012773950971
What about the strings?¶
In [44]:
symbol_strings = []
for strings, label in zip(df_all['symbol table strings'], df_all['label']):
for symbol in strings:
symbol_strings.append({'symbol string': symbol, 'label': label})
pd_symbols = pd.DataFrame.from_records(symbol_strings)
pd_symbols.head()
Out[44]:
| label | symbol string | |
|---|---|---|
| 0 | bad | __mh_execute_header |
| 1 | bad | _AuthorizationCreate |
| 2 | bad | _AuthorizationExecuteWithPrivileges |
| 3 | bad | _BIO_ctrl |
| 4 | bad | _BIO_f_base64 |
5 rows × 2 columns
In [45]:
import sys
sys.path.insert(0,'..')
import data_hacking.simple_stats as ss
In [46]:
# Spin up our g_test class
g_test = ss.GTest()
# Here we'd like to see how strongly various strings are assoicated with being clean or malware.
df_ct, df_cd, df_symbol_stats = g_test.highest_gtest_scores(pd_symbols['symbol string'],
pd_symbols['label'], N=0, matches=0, min_volume=5)
In [47]:
df_symbol_stats.dropna(inplace=True)
df_symbol_stats.sort('bad_g', ascending=0).head(25)
Out[47]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| H1\x8d5A`\x02 | 4836 | 0 | 1.000000 | 0.000000 | 4836 | 3570.141890 | 2935.283635 | 1265.858110 | 0.000000 |
| L\x89\xe7\xff\x158`\x02 | 4836 | 0 | 1.000000 | 0.000000 | 4836 | 3570.141890 | 2935.283635 | 1265.858110 | 0.000000 |
| __ZStL8__ioinit | 160 | 0 | 1.000000 | 0.000000 | 160 | 118.118838 | 97.114430 | 41.881162 | 0.000000 |
| ___tcf_0 | 113 | 6 | 0.949580 | 0.050420 | 119 | 87.850886 | 56.894804 | 31.149114 | -19.764316 |
| __ZN5boost6systemL14posix_categoryE | 92 | 0 | 1.000000 | 0.000000 | 92 | 67.918332 | 55.840797 | 24.081668 | 0.000000 |
| __ZN5boost6systemL11native_ecatE | 92 | 0 | 1.000000 | 0.000000 | 92 | 67.918332 | 55.840797 | 24.081668 | 0.000000 |
| __ZN5boost6systemL10errno_ecatE | 92 | 0 | 1.000000 | 0.000000 | 92 | 67.918332 | 55.840797 | 24.081668 | 0.000000 |
| __GLOBAL__I_a | 94 | 1 | 0.989474 | 0.010526 | 95 | 70.133060 | 55.065291 | 24.866940 | -6.427078 |
| ___cxx_global_var_init | 90 | 0 | 1.000000 | 0.000000 | 90 | 66.441847 | 54.626867 | 23.558153 | 0.000000 |
| __Z41__static_initialization_and_destruction_0ii | 111 | 7 | 0.940678 | 0.059322 | 118 | 87.112643 | 53.796854 | 30.887357 | -20.782115 |
| ___cxx_global_var_init2 | 88 | 0 | 1.000000 | 0.000000 | 88 | 64.965361 | 53.412936 | 23.034639 | 0.000000 |
| ___cxx_global_var_init3 | 88 | 0 | 1.000000 | 0.000000 | 88 | 64.965361 | 53.412936 | 23.034639 | 0.000000 |
| ___cxx_global_var_init4 | 88 | 0 | 1.000000 | 0.000000 | 88 | 64.965361 | 53.412936 | 23.034639 | 0.000000 |
| ___cxx_global_var_init1 | 88 | 0 | 1.000000 | 0.000000 | 88 | 64.965361 | 53.412936 | 23.034639 | 0.000000 |
| __ZN12_GLOBAL__N_12_1E | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| __ZN12_GLOBAL__N_12_5E | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| ___cxx_global_var_init8 | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| __ZN12_GLOBAL__N_12_9E | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| __ZN12_GLOBAL__N_12_2E | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| ___cxx_global_var_init9 | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| __ZN12_GLOBAL__N_12_8E | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| ___cxx_global_var_init11 | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| __ZN12_GLOBAL__N_12_7E | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| ___cxx_global_var_init6 | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
| ___cxx_global_var_init5 | 86 | 0 | 1.000000 | 0.000000 | 86 | 63.488876 | 52.199006 | 22.511124 | 0.000000 |
25 rows × 9 columns
In [48]:
df_symbol_stats.sort('good_g', ascending=0).head(25)
Out[48]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| dyld_stub_binder | 173 | 276 | 0.385301 | 0.614699 | 449 | 331.470990 | -224.986050 | 117.529010 | 471.251050 |
| radr://5614542 | 107 | 236 | 0.311953 | 0.688047 | 343 | 253.217260 | -184.343670 | 89.782740 | 456.159232 |
| ___stderrp | 35 | 187 | 0.157658 | 0.842342 | 222 | 163.889888 | -108.069271 | 58.110112 | 437.119579 |
| _strcmp | 51 | 169 | 0.231818 | 0.768182 | 220 | 162.413403 | -118.148571 | 57.586597 | 363.893795 |
| ___stack_chk_fail | 92 | 190 | 0.326241 | 0.673759 | 282 | 208.184453 | -150.261006 | 73.815547 | 359.272784 |
| ___stack_chk_guard | 92 | 190 | 0.326241 | 0.673759 | 282 | 208.184453 | -150.261006 | 73.815547 | 359.272784 |
| _fprintf | 39 | 156 | 0.200000 | 0.800000 | 195 | 143.957334 | -101.864515 | 51.042666 | 348.564572 |
| ___stdoutp | 8 | 120 | 0.062500 | 0.937500 | 128 | 94.495071 | -39.505698 | 33.504929 | 306.191801 |
| __mh_execute_header | 218 | 225 | 0.492099 | 0.507901 | 443 | 327.041534 | -176.838162 | 115.958466 | 298.290746 |
| _printf | 46 | 123 | 0.272189 | 0.727811 | 169 | 124.763023 | -91.795275 | 44.236977 | 251.565340 |
| ___stdinp | 0 | 93 | 0.000000 | 1.000000 | 93 | 68.656575 | 0.000000 | 24.343425 | 249.302811 |
| _fwrite | 55 | 116 | 0.321637 | 0.678363 | 171 | 126.239509 | -91.393256 | 44.760491 | 220.925324 |
| _fputc | 32 | 101 | 0.240602 | 0.759398 | 133 | 98.186284 | -71.752361 | 34.813716 | 215.152034 |
| ___error | 85 | 127 | 0.400943 | 0.599057 | 212 | 156.507461 | -103.776912 | 55.492539 | 210.296380 |
| ___snprintf_chk | 0 | 78 | 0.000000 | 1.000000 | 78 | 57.582934 | 0.000000 | 20.417066 | 209.092680 |
| _exit | 230 | 189 | 0.548926 | 0.451074 | 419 | 309.323708 | -136.302148 | 109.676292 | 205.712812 |
| _puts | 27 | 89 | 0.232759 | 0.767241 | 116 | 85.636158 | -62.330620 | 30.363842 | 191.418329 |
| _optind | 5 | 74 | 0.063291 | 0.936709 | 79 | 58.321176 | -24.565273 | 20.678824 | 188.693330 |
| _strncmp | 28 | 88 | 0.241379 | 0.758621 | 116 | 85.636158 | -62.602573 | 30.363842 | 187.278840 |
| _putchar | 6 | 74 | 0.075000 | 0.925000 | 80 | 59.059419 | -27.441415 | 20.940581 | 186.831670 |
| _optarg | 1 | 69 | 0.014286 | 0.985714 | 70 | 51.676992 | -7.890025 | 18.323008 | 182.980956 |
| _strtol | 8 | 73 | 0.098765 | 0.901235 | 81 | 59.797662 | -32.184400 | 21.202338 | 180.506805 |
| _stat$INODE64 | 21 | 80 | 0.207921 | 0.792079 | 101 | 74.562517 | -53.218850 | 26.437483 | 177.159010 |
| _strlen | 171 | 150 | 0.532710 | 0.467290 | 321 | 236.975920 | -111.592881 | 84.024080 | 173.859559 |
| ___maskrune | 15 | 73 | 0.170455 | 0.829545 | 88 | 64.965361 | -43.974121 | 23.034639 | 168.405207 |
25 rows × 9 columns
In [49]:
def tokenize_string(string):
if '___cxx_global_var_init' in string:
return '___cxx_global_var_init_TOKEN'
elif '__ZN12_GLOBAL' in string:
return '__ZN12_GLOBAL_TOKEN'
else:
return string
In [50]:
pd_symbols['tokenized string'] = pd_symbols['symbol string'].map(lambda x: tokenize_string(x))
In [51]:
df_ct, df_cd, df_symbol_stats = g_test.highest_gtest_scores(pd_symbols['tokenized string'],
pd_symbols['label'], N=10, matches=5, min_volume=5)
In [52]:
df_symbol_stats.dropna(inplace=True)
df_symbol_stats.sort('bad_g', ascending=0).head(10)
Out[52]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| L\x89\xe7\xff\x158`\x02 | 4836 | 0 | 1.000000 | 0.000000 | 4836 | 4428.513498 | 851.367726 | 407.486502 | 0.000000 |
| H1\x8d5A`\x02 | 4836 | 0 | 1.000000 | 0.000000 | 4836 | 4428.513498 | 851.367726 | 407.486502 | 0.000000 |
| ___cxx_global_var_init_TOKEN | 1560 | 0 | 1.000000 | 0.000000 | 1560 | 1428.552741 | 274.634750 | 131.447259 | 0.000000 |
| __ZN12_GLOBAL_TOKEN | 870 | 5 | 0.994286 | 0.005714 | 875 | 801.271570 | 143.190314 | 73.728430 | -26.909506 |
| _malloc | 160 | 130 | 0.551724 | 0.448276 | 290 | 265.564292 | -162.138608 | 24.435708 | 434.587122 |
| _strlen | 171 | 150 | 0.532710 | 0.467290 | 321 | 293.952199 | -185.279734 | 27.047801 | 513.908873 |
| radr://5614542 | 107 | 236 | 0.311953 | 0.688047 | 343 | 314.098455 | -230.451818 | 28.901545 | 991.170152 |
| _exit | 230 | 189 | 0.548926 | 0.451074 | 419 | 383.694615 | -235.413120 | 35.305385 | 634.174947 |
| __mh_execute_header | 218 | 225 | 0.492099 | 0.507901 | 443 | 405.672349 | -270.778127 | 37.327651 | 808.364717 |
| dyld_stub_binder | 173 | 276 | 0.385301 | 0.614699 | 449 | 411.166783 | -299.534738 | 37.833217 | 1096.941787 |
10 rows × 9 columns
In [53]:
def g_aggregate(df_stats, sequence, name):
try:
g_scores = [df_stats.ix[tokenize_string(item)][name] for item in sequence]
except KeyError:
return 0
return sum(g_scores)/len(g_scores) if g_scores else 0 # Average
In [54]:
df_all['symbol table strings malicious_g'] = \
df_all['symbol table strings'].map(lambda x: g_aggregate(df_symbol_stats, x, 'bad_g'))
df_all['symbol table strings clean_g'] = \
df_all['symbol table strings'].map(lambda x: g_aggregate(df_symbol_stats, x, 'good_g'))
In [55]:
segment_names = []
for strings, label in zip(df_all['segment names'], df_all['label']):
for name in strings:
segment_names.append({'segment names': name, 'label': label})
pd_segment_names = pd.DataFrame.from_records(segment_names)
pd_segment_names.head()
Out[55]:
| label | segment names | |
|---|---|---|
| 0 | bad | __PAGEZERO |
| 1 | bad | __TEXT |
| 2 | bad | __DATA |
| 3 | bad | __LINKEDIT |
| 4 | bad | __TEXT |
5 rows × 2 columns
In [56]:
#Spin up our g_test class
g_test = ss.GTest()
df_ct, df_cd, df_segment_stats = g_test.highest_gtest_scores(pd_segment_names['segment names'],
pd_segment_names['label'], N=0, matches=0, min_volume=0)
In [57]:
df_segment_stats.dropna(inplace=True)
df_segment_stats.sort('bad_g', ascending=0).head(10)
Out[57]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| __OBJC | 55 | 17 | 0.763889 | 0.236111 | 72 | 34.263158 | 52.058877 | 37.736842 | -27.112400 |
| __IMPORT | 48 | 28 | 0.631579 | 0.368421 | 76 | 36.166667 | 27.174060 | 39.833333 | -19.739976 |
| __PAGEZERO | 218 | 225 | 0.492099 | 0.507901 | 443 | 210.813596 | 14.615046 | 232.186404 | -14.148050 |
| 3 | 0 | 1.000000 | 0.000000 | 3 | 1.427632 | 4.455573 | 1.572368 | 0.000000 | |
| __INIT_STUB | 2 | 0 | 1.000000 | 0.000000 | 2 | 0.951754 | 2.970382 | 1.048246 | 0.000000 |
| .ida | 0 | 4 | 0.000000 | 1.000000 | 4 | 1.903509 | 0.000000 | 2.096491 | 5.168234 |
| __LINKEDIT | 253 | 305 | 0.453405 | 0.546595 | 558 | 265.539474 | -24.477253 | 292.460526 | 25.609064 |
| __TEXT | 253 | 305 | 0.453405 | 0.546595 | 558 | 265.539474 | -24.477253 | 292.460526 | 25.609064 |
| __DATA | 253 | 311 | 0.448582 | 0.551418 | 564 | 268.394737 | -29.889069 | 295.605263 | 31.577645 |
9 rows × 9 columns
In [58]:
s = []
for strings, label in zip(df_all['section names'], df_all['label']):
for name in strings:
s.append({'section names': name, 'label': label})
pd_section_names = pd.DataFrame.from_records(s)
print pd_section_names.shape
pd_section_names.head()
(8289, 2)
Out[58]:
| label | section names | |
|---|---|---|
| 0 | bad | __text |
| 1 | bad | __symbol_stub1 |
| 2 | bad | __stub_helper |
| 3 | bad | __cstring |
| 4 | bad | __unwind_info |
5 rows × 2 columns
In [59]:
#Spin up our g_test class
g_test = ss.GTest()
# Here we'd like to see how various exploits (description) are related to
# the ASN (Autonomous System Number) associated with the ip/domain.
df_ct, df_cd, df_section_stats = g_test.highest_gtest_scores(pd_section_names['section names'],
pd_section_names['label'], N=0, matches=0, min_volume=5)
In [60]:
df_section_stats.dropna(inplace=True)
df_section_stats.sort('bad_g', ascending=0).head(10)
Out[60]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| __dyld | 192 | 53 | 0.783673 | 0.216327 | 245 | 125.706961 | 162.640082 | 119.293039 | -85.996853 |
| __symbol_stub1 | 55 | 1 | 0.982143 | 0.017857 | 56 | 28.733020 | 71.421483 | 27.266980 | -6.611353 |
| __symbol_stub | 124 | 62 | 0.666667 | 0.333333 | 186 | 95.434672 | 64.936223 | 90.565328 | -46.988195 |
| __common | 212 | 156 | 0.576087 | 0.423913 | 368 | 188.816986 | 49.102615 | 179.183014 | -43.228130 |
| __instance_vars | 43 | 6 | 0.877551 | 0.122449 | 49 | 25.141392 | 46.154870 | 23.858608 | -16.564627 |
| __cls_refs | 50 | 12 | 0.806452 | 0.193548 | 62 | 31.811557 | 45.219334 | 30.188443 | -22.141260 |
| __message_refs | 50 | 12 | 0.806452 | 0.193548 | 62 | 31.811557 | 45.219334 | 30.188443 | -22.141260 |
| __module_info | 51 | 13 | 0.796875 | 0.203125 | 64 | 32.837737 | 44.905221 | 31.162263 | -22.730721 |
| __meta_class | 44 | 8 | 0.846154 | 0.153846 | 52 | 26.680661 | 44.022056 | 25.319339 | -18.434031 |
| __symbols | 44 | 8 | 0.846154 | 0.153846 | 52 | 26.680661 | 44.022056 | 25.319339 | -18.434031 |
10 rows × 9 columns
In [61]:
df_section_stats.sort('good_g', ascending=0).head(10)
Out[61]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| __stubs | 28 | 210 | 0.117647 | 0.882353 | 238 | 122.115334 | -82.474641 | 115.884666 | 249.695077 |
| __got | 29 | 171 | 0.145000 | 0.855000 | 200 | 102.617927 | -73.295575 | 97.382073 | 192.553326 |
| __unwind_info | 192 | 301 | 0.389452 | 0.610548 | 493 | 252.953191 | -105.872288 | 240.046809 | 136.218346 |
| __stub_helper | 192 | 273 | 0.412903 | 0.587097 | 465 | 238.586681 | -83.419129 | 226.413319 | 102.161853 |
| __nl_symbol_ptr | 198 | 274 | 0.419492 | 0.580508 | 472 | 242.178309 | -79.757266 | 229.821691 | 96.351745 |
| __const | 272 | 341 | 0.443719 | 0.556281 | 613 | 314.523947 | -79.020434 | 298.476053 | 90.837472 |
| __la_symbol_ptr | 204 | 273 | 0.427673 | 0.572327 | 477 | 244.743757 | -74.293446 | 232.256243 | 88.250273 |
| __cstring | 242 | 302 | 0.444853 | 0.555147 | 544 | 279.120762 | -69.070092 | 264.879238 | 79.216414 |
| __eh_frame | 126 | 186 | 0.403846 | 0.596154 | 312 | 160.083967 | -60.332974 | 151.916033 | 75.299763 |
| __text | 255 | 304 | 0.456172 | 0.543828 | 559 | 286.817107 | -59.966417 | 272.182893 | 67.216504 |
10 rows × 9 columns
In [62]:
df_all['segment names malicious_g'] = df_all['segment names'].map(lambda x: g_aggregate(df_segment_stats, x, 'bad_g'))
df_all['segment names clean_g'] = df_all['segment names'].map(lambda x: g_aggregate(df_segment_stats, x, 'good_g'))
df_all['section names malicious_g'] = df_all['section names'].map(lambda x: g_aggregate(df_section_stats, x, 'bad_g'))
df_all['section names clean_g'] = df_all['section names'].map(lambda x: g_aggregate(df_section_stats, x, 'good_g'))
In [63]:
g_test = ss.GTest()
df_entrypoint = pd.DataFrame(df_all, columns=['entry point section name', 'entry point segment name',
'entry point instruction type', 'label'])
In [64]:
df_entrypoint = df_entrypoint.replace(0, np.nan)
df_entrypoint.dropna(inplace=True)
df_entrypoint.head()
Out[64]:
| entry point section name | entry point segment name | entry point instruction type | label | |
|---|---|---|---|---|
| 0 | __text | __TEXT | push | bad |
| 1 | -1 | -1 | -1 | bad |
| 2 | __text | __TEXT | pop | bad |
| 3 | __text | __TEXT | push | bad |
| 4 | __text | __TEXT | add | bad |
5 rows × 4 columns
In [65]:
df_ct, df_cd, df_ep_section_stats = g_test.highest_gtest_scores(df_entrypoint['entry point section name'],
df_entrypoint['label'], N=0, matches=0, min_volume=5)
df_ct, df_cd, df_ep_segment_stats = g_test.highest_gtest_scores(df_entrypoint['entry point segment name'],
df_entrypoint['label'], N=0, matches=0, min_volume=5)
df_ct, df_cd, df_ep_instruction_stats = g_test.highest_gtest_scores(df_entrypoint['entry point instruction type'],
df_entrypoint['label'], N=0, matches=0, min_volume=5)
In [66]:
df_ep_section_stats.head()
Out[66]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| __text | 214 | 225 | 0.487472 | 0.512528 | 439 | 199.901786 | 29.168132 | 239.098214 | -27.348302 |
| -1 | 34 | 80 | 0.298246 | 0.701754 | 114 | 51.910714 | -28.775199 | 62.089286 | 40.552511 |
| COULD NOT FIND | 7 | 0 | 1.000000 | 0.000000 | 7 | 3.187500 | 11.013425 | 3.812500 | 0.000000 |
3 rows × 9 columns
In [67]:
df_ep_segment_stats.head()
Out[67]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| __TEXT | 214 | 225 | 0.487472 | 0.512528 | 439 | 199.901786 | 29.168132 | 239.098214 | -27.348302 |
| -1 | 34 | 80 | 0.298246 | 0.701754 | 114 | 51.910714 | -28.775199 | 62.089286 | 40.552511 |
| COULD NOT FIND | 7 | 0 | 1.000000 | 0.000000 | 7 | 3.187500 | 11.013425 | 3.812500 | 0.000000 |
3 rows × 9 columns
In [68]:
df_ep_instruction_stats.dropna(inplace=True)
df_ep_instruction_stats.sort('bad_g', ascending=0).head(20)
Out[68]:
| bad | good | bad_cd | good_cd | total_cd | bad_exp | bad_g | good_exp | good_g | |
|---|---|---|---|---|---|---|---|---|---|
| push | 155 | 153 | 0.503247 | 0.496753 | 308 | 140.250000 | 30.999557 | 167.750000 | -28.163277 |
| COULD NOT FIND | 7 | 0 | 1.000000 | 0.000000 | 7 | 3.187500 | 11.013425 | 3.812500 | 0.000000 |
| pop | 5 | 1 | 0.833333 | 0.166667 | 6 | 2.732143 | 6.043517 | 3.267857 | -2.368269 |
| das | 0 | 8 | 0.000000 | 1.000000 | 8 | 3.642857 | 0.000000 | 4.357143 | 9.722000 |
| add | 42 | 54 | 0.437500 | 0.562500 | 96 | 43.714286 | -3.360448 | 52.285714 | 3.484173 |
| -1 | 34 | 80 | 0.298246 | 0.701754 | 114 | 51.910714 | -28.775199 | 62.089286 | 40.552511 |
6 rows × 9 columns
In [69]:
df_entrypoint['entry point instruction type'].value_counts().plot(kind='bar')
Out[69]:
<matplotlib.axes.AxesSubplot at 0x116998210>
In [70]:
clf_final = sklearn.ensemble.RandomForestClassifier(n_estimators=50)
final_labels = list(df_all.columns.values)
final_labels.remove('label')
final_labels.remove('cputype')
final_labels.remove('LC_UNIXTHREAD')
final_labels.remove('LC_MAIN')
final_labels.remove('const strings')
final_labels.remove('symbol table strings')
final_labels.remove('segment names')
final_labels.remove('section names')
final_labels.remove('entry point section name')
final_labels.remove('entry point segment name')
final_labels.remove('entry point instruction type')
X_final = df_all.as_matrix(final_labels)
y_final = np.array(df_all['label'].tolist())
In [71]:
scores = sklearn.cross_validation.cross_val_score(clf_final, X_final, y_final, cv=10, n_jobs=5)
print("Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std() * 2))
Accuracy: 0.96 (+/- 0.06)
In [74]:
# 80/20 Split for predictive test
X_train_final, X_test_final, y_train_final, y_test_final = train_test_split(X_final, y_final,
test_size=my_tsize,
random_state=my_seed)
clf_final.fit(X_train_final, y_train_final)
y_pred_final = clf_final.predict(X_test_final)
cm_final = confusion_matrix(y_test_final, y_pred_final, labels)
plot_cm(cm_final, labels)
Confusion Matrix Stats good/good: 94.20% (65/69) good/bad: 5.80% (4/69) bad/good: 2.33% (1/43) bad/bad: 97.67% (42/43)
You can set threshold values to your priorities.¶
In [75]:
y_probs_final = clf_final.predict_proba(X_test_final)[:,0]
thres = .8 # This can be set to whatever you'd like
y_pred_final[y_probs_final<thres] = 'good'
y_pred_final[y_probs_final>=thres] = 'bad'
cm = confusion_matrix(y_test_final, y_pred_final, labels)
plot_cm(cm, labels)
Confusion Matrix Stats good/good: 98.55% (68/69) good/bad: 1.45% (1/69) bad/good: 16.28% (7/43) bad/bad: 83.72% (36/43)
Most important features¶
Now that we added in the g-scores from the strings of section names, segment names, and symbol table, now what are the most important features?
In [76]:
importances = zip(final_labels, clf_final.feature_importances_)
importances.sort(key=lambda k:k[1], reverse=True)
for im in importances[0:20]:
print im[0].ljust(35), im[1]
section names malicious_g 0.0657407559464 LC_SOURCE_VERSION 0.0553265015965 section names clean_g 0.0535257311096 LC_DYLIB_CODE_SIGN_DRS 0.048614836495 LC_DATA_IN_CODE 0.0464634185482 rebase_size 0.043812886794 LC_CODE_SIGNATURE 0.0334035056079 rebase_off 0.0311568256564 text_section_align 0.0232383559178 data_nsects 0.021464002581 linkedit_vmsize 0.0198348290427 lazy_bind_size 0.0173961188418 pz_vmaddr 0.0165259506433 pz_vmsize 0.0156923760629 nextdefsym 0.0152103851287 pz_fileoff 0.0141570209658 entry point offset 0.0140948505669 nlocrel 0.0131386093896 bind_size 0.0121543445893 pz_flags 0.0120473460984
Try a new classifier with only the 20 most important features¶
In [77]:
clf_most = sklearn.ensemble.RandomForestClassifier(n_estimators=50)
X = df_all.as_matrix([item[0] for item in importances[:20]])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=my_tsize, random_state=my_seed)
clf_most.fit(X_train, y_train)
y_pred = clf_most.predict(X_test)
cm = confusion_matrix(y_test, y_pred, labels)
plot_cm(cm, labels)
Confusion Matrix Stats good/good: 92.75% (64/69) good/bad: 7.25% (5/69) bad/good: 2.33% (1/43) bad/bad: 97.67% (42/43)
In [78]:
y_probs = clf_most.predict_proba(X_test)[:,0]
thres = .92 # This can be set to whatever you'd like
y_pred[y_probs<thres] = 'good'
y_pred[y_probs>=thres] = 'bad'
cm = confusion_matrix(y_test, y_pred, labels)
plot_cm(cm, labels)
Confusion Matrix Stats good/good: 98.55% (68/69) good/bad: 1.45% (1/69) bad/good: 20.93% (9/43) bad/bad: 79.07% (34/43)
Using a different classifier¶
Let's try a different classifier: Extra Trees Classifier (like RandomForest, but even more random).http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.ExtraTreesClassifier.html
In [88]:
clf_most = sklearn.ensemble.ExtraTreesClassifier(n_estimators=50)
X = df_all.as_matrix([item[0] for item in importances[:20]])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=my_tsize, random_state=my_seed)
clf_most.fit(X_train, y_train)
y_pred = clf_most.predict(X_test)
cm = confusion_matrix(y_test, y_pred, labels)
plot_cm(cm, labels)
Confusion Matrix Stats good/good: 94.20% (65/69) good/bad: 5.80% (4/69) bad/good: 2.33% (1/43) bad/bad: 97.67% (42/43)
Going one step further¶
It's nice to give a prediction on whether the binary is good or bad, but it would be nicer to include why that prediction is given. Let's explore the rebase_size and rebase_off features from the LC_DYLD_INFO command. The dyld_info_command contains the file offsets and sizes of the new compressed form of the information dyld needs to load the image.Dyld rebases an image whenever dyld loads it at an address different from its preferred address.
- rebase_off is the file offset to rebase info
- rebase_size is the size of rebase info
In [89]:
df_all.boxplot(column='rebase_off', by='label')
plt.xlabel('bad vs. good files')
plt.ylabel('rebase_off')
plt.title('Comparision of rebase offset')
plt.suptitle("")
Out[89]:
<matplotlib.text.Text at 0x115e8f8d0>
Not the most useful plot, let's try again, but not show the outliers.
In [90]:
df_all.boxplot(column='rebase_off', by='label', sym='')
plt.xlabel('bad vs. good files')
plt.ylabel('rebase_off')
plt.title('Comparision of rebase offset')
plt.suptitle("")
Out[90]:
<matplotlib.text.Text at 0x115ea0b50>
In [91]:
cond = df_all['label'] == 'good'
good_all = df_all[cond]
bad_all = df_all[~cond]
bvc = pd.DataFrame(bad_all['rebase_off'].value_counts(), columns=['count'])
bvc['perctange'] = bvc/bad_all.shape[0]
gvc = pd.DataFrame(good_all['rebase_off'].value_counts(), columns=['count'])
gvc['perctange'] = gvc/good_all.shape[0]
print 'Most common rebase_off values from malicious set'
print bvc.head(5)
print ''
print 'Most common rebase_off values from clean set'
print gvc.head()
Most common rebase_off values from malicious set
count perctange
0 125 0.490196
-1 81 0.317647
20480 10 0.039216
8192 10 0.039216
319488 4 0.015686
[5 rows x 2 columns]
Most common rebase_off values from clean set
count perctange
8192 53 0.173770
12288 49 0.160656
-1 29 0.095082
16384 27 0.088525
0 21 0.068852
[5 rows x 2 columns]
In [92]:
bad_all['rebase_off'].hist(alpha=1,label='bad',bins=1000)
good_all['rebase_off'].hist(alpha=1,label='good',bins=1000)
plt.title('Values for rebase offset')
plt.legend()
plt.xlim(-50,3000000)
Out[92]:
(-50, 3000000)
Well, that wasn't very helpful, let's not show the long tail on the plot¶
In [93]:
bad_all['rebase_off'].hist(alpha=1,label='bad',bins=2000)
good_all['rebase_off'].hist(alpha=1,label='good',bins=2000)
plt.legend()
plt.title('Values for rebase offset')
plt.xlim(-50,80000)
Out[93]:
(-50, 80000)
In [94]:
df_all.boxplot(column='rebase_size', by='label')
plt.xlabel('bad vs. good files')
plt.ylabel('rebase_size')
plt.title('Comparision of rebase size')
plt.suptitle("")
Out[94]:
<matplotlib.text.Text at 0x1169b0250>
In [95]:
df_all.boxplot(column='rebase_size', by='label', sym='')
plt.xlabel('bad vs. good files')
plt.ylabel('rebase_size')
plt.title('Comparision of rebase size')
plt.suptitle("")
Out[95]:
<matplotlib.text.Text at 0x1169bbbd0>
In [96]:
bvc = pd.DataFrame(bad_all['rebase_size'].value_counts(), columns=['count'])
bvc['percentange'] = bvc/bad_all.shape[0]
gvc = pd.DataFrame(good_all['rebase_size'].value_counts(), columns=['count'])
gvc['percentange'] = gvc/good_all.shape[0]
print 'Most common rebase_size values from malicious set'
print bvc.head(5)
print ''
print 'Most common rebase_size values from clean set'
print gvc.head(5)
Most common rebase_size values from malicious set
count percentange
0 125 0.490196
-1 81 0.317647
48 6 0.023529
16 6 0.023529
28 5 0.019608
[5 rows x 2 columns]
Most common rebase_size values from clean set
count percentange
8 80 0.262295
16 44 0.144262
-1 29 0.095082
0 21 0.068852
24 17 0.055738
[5 rows x 2 columns]
In [97]:
plt.scatter(good_all['rebase_off'], good_all['rebase_size'],
s=140, c='#aaaaff', label='Good', alpha=.4)
plt.scatter(bad_all['rebase_off'], bad['rebase_size'],
s=40, c='r', label='Bad', alpha=.5)
plt.title('rebase_off vs rebase_size')
plt.xlabel('rebase_off')
plt.ylabel('rebase_size')
Out[97]:
<matplotlib.text.Text at 0x119130550>
Let's zoom in around the origin¶
In [98]:
plt.scatter(good_all['rebase_off'], good_all['rebase_size'],
s=140, c='#aaaaff', label='Good', alpha=.4)
plt.scatter(bad_all['rebase_off'], bad['rebase_size'],
s=40, c='r', label='Bad', alpha=.5)
plt.xlim(-1000,150000)
plt.ylim(-100,1100)
plt.title('rebase_off vs rebase_size')
plt.xlabel('rebase_off')
plt.ylabel('rebase_size')
Out[98]:
<matplotlib.text.Text at 0x118632e10>
In [ ]:
plt.scatter(good_all['rebase_off'], good_all['rebase_size'],
s=140, c='#aaaaff', label='Good', alpha=.4)
plt.scatter(bad_all['rebase_off'], bad['rebase_size'],
s=40, c='r', label='Bad', alpha=.5)
plt.xlim(-10,10)
plt.ylim(-10,10)
plt.title('rebase_off vs rebase_size')
plt.xlabel('rebase_off')
plt.ylabel('rebase_size')
How does current AV do on these files?¶
Scan data is as of 2014/04/28In [99]:
def load_vt_data(vt_list):
import json
import collections
results = {}
for filename in vt_list:
with open(filename,'rb') as f:
vt_data = f.read()
if len(vt_data) == 0:
break
vt = json.loads(vt_data)
for engine in vt['scans'].keys():
if engine not in results:
results[engine] = 0
if vt['scans'][engine]['detected']:
results[engine] += 1
r = []
for key, value in results.iteritems():
r.append({'Engine': key, 'Count': value})
return r
In [100]:
vt_list = glob.glob('data/bad/vt_data/*.vtdata')
num_files = len(vt_list)
vt_results = load_vt_data(vt_list)
vt_df = pd.DataFrame.from_records(vt_results, columns=['Engine', 'Count'])
vt_df['Files Scanned'] = num_files
vt_df['Percentage'] = vt_df['Count']/num_files
vt_df.sort('Count', ascending=0).head(52)
Out[100]:
| Engine | Count | Files Scanned | Percentage | |
|---|---|---|---|---|
| 18 | Avast | 261 | 261 | 1.000000 |
| 40 | GData | 261 | 261 | 1.000000 |
| 28 | F-Secure | 261 | 261 | 1.000000 |
| 25 | Ad-Aware | 260 | 261 | 0.996169 |
| 1 | MicroWorld-eScan | 260 | 261 | 0.996169 |
| 21 | BitDefender | 260 | 261 | 0.996169 |
| 20 | Kaspersky | 255 | 261 | 0.977011 |
| 34 | Emsisoft | 254 | 261 | 0.973180 |
| 2 | nProtect | 254 | 261 | 0.973180 |
| 45 | ESET-NOD32 | 254 | 261 | 0.973180 |
| 31 | AntiVir | 245 | 261 | 0.938697 |
| 27 | Comodo | 242 | 261 | 0.927203 |
| 26 | Sophos | 240 | 261 | 0.919540 |
| 29 | DrWeb | 240 | 261 | 0.919540 |
| 47 | Ikarus | 238 | 261 | 0.911877 |
| 19 | ClamAV | 231 | 261 | 0.885057 |
| 51 | Qihoo-360 | 230 | 261 | 0.881226 |
| 49 | AVG | 228 | 261 | 0.873563 |
| 4 | CAT-QuickHeal | 226 | 261 | 0.865900 |
| 12 | NANO-Antivirus | 224 | 261 | 0.858238 |
| 17 | TrendMicro-HouseCall | 203 | 261 | 0.777778 |
| 14 | Symantec | 190 | 261 | 0.727969 |
| 38 | Microsoft | 175 | 261 | 0.670498 |
| 42 | AhnLab-V3 | 170 | 261 | 0.651341 |
| 0 | Bkav | 149 | 261 | 0.570881 |
| 30 | VIPRE | 128 | 261 | 0.490421 |
| 32 | TrendMicro | 102 | 261 | 0.390805 |
| 5 | McAfee | 90 | 261 | 0.344828 |
| 33 | McAfee-GW-Edition | 90 | 261 | 0.344828 |
| 23 | ViRobot | 85 | 261 | 0.325670 |
| 15 | Norman | 55 | 261 | 0.210728 |
| 46 | Rising | 49 | 261 | 0.187739 |
| 13 | F-Prot | 42 | 261 | 0.160920 |
| 41 | Commtouch | 41 | 261 | 0.157088 |
| 35 | Jiangmin | 40 | 261 | 0.153257 |
| 9 | K7AntiVirus | 36 | 261 | 0.137931 |
| 10 | K7GW | 35 | 261 | 0.134100 |
| 16 | TotalDefense | 34 | 261 | 0.130268 |
| 48 | Fortinet | 27 | 261 | 0.103448 |
| 22 | Agnitum | 26 | 261 | 0.099617 |
| 43 | VBA32 | 25 | 261 | 0.095785 |
| 7 | Zillya | 21 | 261 | 0.080460 |
| 50 | Panda | 11 | 261 | 0.042146 |
| 3 | CMC | 5 | 261 | 0.019157 |
| 36 | Antiy-AVL | 1 | 261 | 0.003831 |
| 24 | ByteHero | 0 | 261 | 0.000000 |
| 44 | Baidu-International | 0 | 261 | 0.000000 |
| 6 | Malwarebytes | 0 | 261 | 0.000000 |
| 11 | TheHacker | 0 | 261 | 0.000000 |
| 39 | AegisLab | 0 | 261 | 0.000000 |
| 37 | Kingsoft | 0 | 261 | 0.000000 |
| 8 | SUPERAntiSpyware | 0 | 261 | 0.000000 |
52 rows × 4 columns
In [101]:
vt_list = glob.glob('data/good/vt_data/*.vtdata')
num_files = len(vt_list)
vt_results = load_vt_data(vt_list)
vt_df = pd.DataFrame.from_records(vt_results, columns=['Engine', 'Count'])
vt_df['Files Scanned'] = num_files
vt_df['Percentage'] = vt_df['Count']/num_files
vt_df.sort('Count', ascending=0).head(15)
Out[101]:
| Engine | Count | Files Scanned | Percentage | |
|---|---|---|---|---|
| 51 | Qihoo-360 | 0 | 266 | 0 |
| 50 | Panda | 0 | 266 | 0 |
| 23 | SUPERAntiSpyware | 0 | 266 | 0 |
| 22 | NANO-Antivirus | 0 | 266 | 0 |
| 21 | BitDefender | 0 | 266 | 0 |
| 20 | Kaspersky | 0 | 266 | 0 |
| 19 | ClamAV | 0 | 266 | 0 |
| 18 | Avast | 0 | 266 | 0 |
| 17 | TrendMicro-HouseCall | 0 | 266 | 0 |
| 16 | TotalDefense | 0 | 266 | 0 |
| 15 | Norman | 0 | 266 | 0 |
| 14 | Symantec | 0 | 266 | 0 |
| 13 | F-Prot | 0 | 266 | 0 |
| 12 | Agnitum | 0 | 266 | 0 |
| 11 | K7AntiVirus | 0 | 266 | 0 |
15 rows × 4 columns