Goal: Scalable & Reproducible Analysis¶
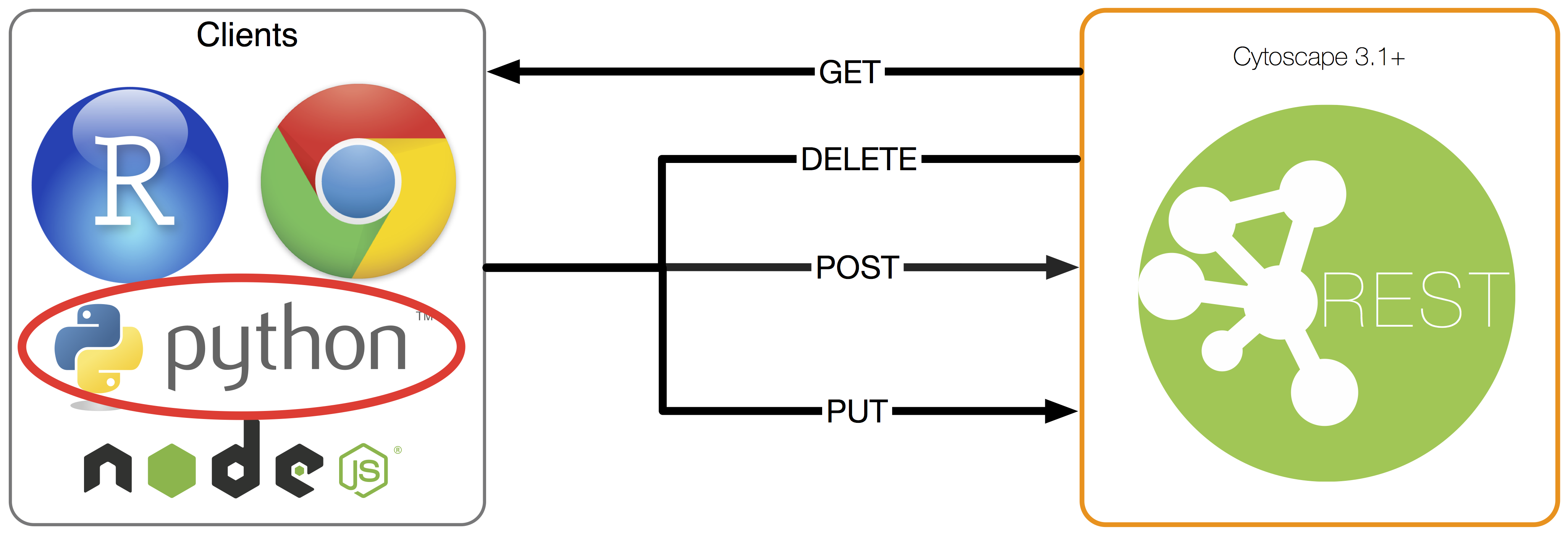
Make Cytoscape external tool friendly
Cytoscape Session File = Result¶
IPython Notebook + cyREST = Process¶
New to IPython Notebook?¶
There is a great Nature article about this technology:
Interactive notebooks: Sharing the code¶
The free IPython notebook makes data analysis easier to record, understand and reproduce.
by Helen Shen 05 November 2014
In [1]:
import requests # Best REST client for Python!
import json # Standard toolkit to do something with JSON in Python
import networkx as nx # Popular python library for network analysis
from IPython.display import Image # Library to display images in IPython Notebook
import pandas as pd # Pandas - de-facto standard tool for data analysis in Python
from py2cytoscape import util # Small utility to convert Python objects to Cytoscape-friendly ones.
# Default Port Number for cyREST
PORT_NUMBER = 1234
# Header for posting data to the server as JSON
HEADERS = {'Content-Type': 'application/json'}
# The base URL to talking to local instance of Cytoscape
BASE_URL = 'http://localhost:' + str(PORT_NUMBER) + '/v1/'
styles_url = BASE_URL + 'styles'
print('\n\n\nCytoscape service is located at: ' + BASE_URL + '\n\n\n')
print('Available Styles are: ' + styles_url + '\n\n\n')
Cytoscape service is located at: http://localhost:1234/v1/ Available Styles are: http://localhost:1234/v1/styles
In [2]:
def create_scale_free_graph(size, index):
scale_free_graph = nx.scale_free_graph(size)
scale_free_graph.graph['name'] = 'Scale-Free Graph ' + str(index)
return scale_free_graph
# Give a fresh start... Delete all networks
requests.delete(BASE_URL + 'networks')
# Create 10 scale-free networks on-the-fly
networks = []
suids = []
for i in range(5):
network = create_scale_free_graph(100, i)
networks.append(network)
# Send generated network to Cytoscape
res1 = requests.post(BASE_URL + 'networks', data=json.dumps(util.from_networkx(network)), headers=HEADERS)
res1_dict = json.loads(res1.content)
new_suid = res1_dict['networkSUID']
suids.append(new_suid)
# Apply layout!
requests.get(BASE_URL + 'apply/layouts/force-directed/' + str(new_suid))
requests.get(BASE_URL + 'apply/styles/Directed/' + str(new_suid))
In [3]:
network_image_url = BASE_URL+ 'networks/' + str(suids[0]) + '/views/first.png'
print('\n\n\nThe network Image is available here: ' + network_image_url + '\n\n\n')
Image(network_image_url)
The network Image is available here: http://localhost:1234/v1/networks/52/views/first.png
Out[3]:
Use External Services¶
What is this (messy) code doing?¶
- Search KEGG Disease database to get list of cancer types
- Get list of pathways related to those cancers
- Feed the list of KEGG pathways to Cytoscape and visualize them
Agents working in this workflow¶
- Cytoscape - Visualizer & KEGG data reader
- IPython Notebook - Conductor
- KEGG Pathway Database - External Service
- KEGG Disease Database - External Service
- KEGG API - External Service
In [4]:
import io
# KEGG API
KEGG_API_URL = 'http://rest.kegg.jp/'
# Find information about 'cancer' from KEGG disease database.
query = 'cancer'
res = requests.get(KEGG_API_URL + '/find/disease/' + query)
pathway_list = res.content.decode('utf8')
disease_df = pd.read_csv(io.StringIO(pathway_list), delimiter='\t', header=None, names=['id', 'name'])
disease_df.head(10)
Out[4]:
| id | name | |
|---|---|---|
| 0 | ds:H00013 | Small cell lung cancer |
| 1 | ds:H00014 | Non-small cell lung cancer |
| 2 | ds:H00016 | Oral cancer |
| 3 | ds:H00017 | Esophageal cancer |
| 4 | ds:H00018 | Gastric cancer |
| 5 | ds:H00019 | Pancreatic cancer |
| 6 | ds:H00020 | Colorectal cancer |
| 7 | ds:H00022 | Bladder cancer |
| 8 | ds:H00023 | Testicular cancer |
| 9 | ds:H00024 | Prostate cancer |
In [5]:
disease_ids = disease_df['id']
disease_urls = disease_ids.apply(lambda x: KEGG_API_URL + 'get/' + x)
def disease_parser(entry):
lines = entry.split('\n')
data = {}
last_key = None
for line in lines:
if '///' in line:
return data
parts = line.split(' ')
if parts[0] is not None and len(parts[0]) != 0:
last_key = parts[0]
data[parts[0]] = line.replace(parts[0], '').strip()
else:
last_val = data[last_key]
data[last_key] = last_val + '|' + line.strip()
return data
result = []
for url in disease_urls:
res = requests.get(url)
rows = disease_parser(res.content)
result.append(rows)
disease_df = pd.DataFrame(result)
pathways = disease_df['PATHWAY'].dropna().unique()
p_urls = []
for pathway in pathways:
entries = pathway.split('|')
for en in entries:
url = KEGG_API_URL + 'get/' + en.split(' ')[0].split('(')[0] + '/kgml'
p_urls.append(url)
In [ ]:
disease_df = pd.DataFrame(result)
pathways = disease_df['PATHWAY'].dropna().unique()
p_urls = []
for pathway in pathways:
entries = pathway.split('|')
for en in entries:
url = KEGG_API_URL + 'get/' + en.split(' ')[0].split('(')[0] + '/kgml'
p_urls.append(url)
Import all cancer pathways into Cytoscape from KEGG Pathway DB¶
In [ ]:
def create_from_list(network_list):
server_res = requests.post(BASE_URL + 'networks?source=url&collection=' + query, data=json.dumps(network_list), headers=HEADERS)
return json.loads(server_res.content)
requests.delete(BASE_URL + 'networks')
url_list = list(set(p_urls))
pathway_suids = create_from_list(url_list)
Embedded Visualization with Cytoscape.js¶
In [ ]:
from IPython.html.widgets import interact
from IPython.html.widgets import *
from IPython.html import widgets
# Package to render networks in Cytoscape.js
from py2cytoscape import cytoscapejs as cyjs
layouts = cyjs.get_layouts()
networks = {}
res1 = requests.get(BASE_URL + 'styles/KEGG Style.json')
vs_collection = json.loads(res1.content)
styles = {}
for style in vs_collection:
style_settings = style['style']
title = style['title']
styles[title] = style_settings
# Load local network files
url2 = BASE_URL + 'networks/' + str(pathway_suids[2]['networkSUID'][0])
for i in range(len(pathway_suids)):
net1 = requests.get(BASE_URL + 'networks/' + str(pathway_suids[i]['networkSUID'][0]) + '/views/first')
pathway = json.loads(net1.content)
title = pathway['data']['name']
networks[title] = pathway
In [ ]:
def render_graph(Network, Style, Layout):
cyjs.render(Network, Style, Layout)
interact(render_graph,
Network = DropdownWidget(values=networks),
Style = DropdownWidget(values=styles),
Layout = DropdownWidget(values=layouts, value=layouts['Preset'])
)

kono@ucsd.edu