Introduction¶
The Generalised Linear Model (GLM) is a flexible way to combine different predictor variables to predict the value of a response variable. Many common statistical procedures are types of GLMs - t-tests, multiple regression, ANOVA, ANCOVA, etc. Furthermore, many more complicated statistical procedures are extensions of GLMs. An understanding of how GLMs work is therefore very useful.
In the next few lectures we want to provide an overview of how GLMs work, with some simple examples. In particular, we want you to understand
- how to build design matrices
- the meaning of coefficients in a design matrix
- what a link function is
General form of the model¶
The general form of a GLM can be written as
$y = f(\mathbf{X}\beta) + \epsilon$
$\mathbf{X}$: the design matrix, where columns are predictors, rows are observations
$y$ is the response variable we're trying to predict (a column vector)
$\beta$ is a vector of weights, with the same length as the number of columns in the design matrix
$\mathbf{X} \beta$ is called the linear predictor
$f()$ is some transformation of the weighted design matrix (the link function). Examples include logistic, logarithmic, Poisson...
$\epsilon$ is random noise of some type.

(note that in the picture above, the vector of weights $\beta$ should really appear tall rather than wide because it's a column vector; I've drawn it this way to emphasise that it has the same length as the number of columns in $\mathbf{X}$)
Another way to write $\mathbf{X}\beta$ is in vector notation:
$x_1\beta_1 + x_2\beta_2 + ... + x_n\beta_n$
- $x_n$ are column vectors
- $\beta_n$ is a scalar
So the term $\mathbf{X}\beta$ is a linear weighted sum of predictors (i.e., it's a linear model). Let's look at a concrete example of this in action.
Example: simple linear regression¶
Let's go back to our "Sleep in Mammals" dataset from the dataframe / plotting lecture.
First, we import the data, drop missing cases, and create our log bodyweight variable.
# import the dataset csv file into a Pandas dataframe object:
sleep = pd.read_csv('../Datasets/sleep.txt', sep='\t')
sleep = sleep.dropna(axis=0) # drop on axis 0 (i.e. rows)
sleep['log_BodyWt'] = np.log(sleep['BodyWt'])
sleep.info()
<class 'pandas.core.frame.DataFrame'> Int64Index: 42 entries, 1 to 60 Data columns (total 12 columns): Species 42 non-null object BodyWt 42 non-null float64 BrainWt 42 non-null float64 NonDreaming 42 non-null float64 Dreaming 42 non-null float64 TotalSleep 42 non-null float64 LifeSpan 42 non-null float64 Gestation 42 non-null float64 Predation 42 non-null int64 Exposure 42 non-null int64 Danger 42 non-null int64 log_BodyWt 42 non-null float64 dtypes: float64(8), int64(3), object(1) memory usage: 4.3+ KB
g = sns.jointplot("log_BodyWt", "TotalSleep", sleep)
We can see that the log of bodyweight seems to be related to TotalSleep approximately linearly (although the variance is higher for low bodyweights).
How many hours does a species of animal sleep on average, given its bodyweight?
To answer this question we can use a linear regression. We have a single predictor variable (the log of bodyweight) to predict our response variable of interest (Total hours of sleep).
As you know from high school maths, the equation for a line can be written as $y = mx + c$. That is, we would like to estimate two parameters: the a slope as a function of $x$, and an intercept (the value of $y$ when $x$ is zero).
Design matrices¶
The design matrix $\mathbf X$ has predictor variables in its columns. Remember that we will estimate one parameter (the $\beta$ weight) for each column of the design matrix.
The design matrix is not necessarily just a copy of the columns in our data frame. Sometimes variables need to be transformed to have a certain meaning within the model. We can use a module called Patsy to create design matrices automatically from our specification of the model.
Patsy and S-style formulae¶
The Patsy package allows us to specify the model we want to fit in a high-level formula language, based on the one used in the ancient S statistical language (and now more recently in R).
Patsy allows us to specify
- our model
- the data frame containing the variables we want to use
and returns the design matrix $\mathbf X$ (and optionally also $y$).
Formulae have the general form y ~ x.
- everthing to the left of the
~character are the things we want to predict (response variable) - everything to the right are the predictors
- we can use the names of variables in our data frame to make the code more readable
To make this more concrete, let's look at an example for our linear regression above.
# this command returns both X and y (hence "dmatrices" plural). See "dmatrix" if you just want X.
y, X = patsy.dmatrices('TotalSleep ~ log_BodyWt', sleep)
y
DesignMatrix with shape (42, 1)
TotalSleep
8.3
3.9
9.8
19.7
6.2
14.5
9.7
12.5
3.9
8.4
8.6
10.7
10.7
3.8
14.4
6.2
13.8
8.2
2.9
9.1
19.9
8.0
13.2
12.8
19.4
17.4
17.0
10.9
13.7
8.4
[12 rows omitted]
Terms:
'TotalSleep' (column 0)
(to view full data, use np.asarray(this_obj))
y in this case is just the column vector of TotalSleep. What about X?
X
DesignMatrix with shape (42, 2)
Intercept log_BodyWt
1 0.00000
1 7.84267
1 2.35613
1 -3.77226
1 5.07517
1 1.19392
1 3.95432
1 -0.85567
1 6.14204
1 -2.59027
1 1.09861
1 -0.24207
1 -1.60944
1 3.31999
1 -2.12026
1 4.44265
1 -2.29263
1 0.03922
1 6.25575
1 -5.29832
1 -4.60517
1 4.12713
1 -3.77226
1 -3.03655
1 0.53063
1 1.25276
1 -0.73397
1 2.30259
1 0.48243
1 5.25750
[12 rows omitted]
Terms:
'Intercept' (column 0)
'log_BodyWt' (column 1)
(to view full data, use np.asarray(this_obj))
Note how the second column of X is just the log bodyweight values from our data frame. However, Patsy has added an additional column: the intercept, where each value is 1. What does this mean?
When we do the regression, remember that we will estimate a $\beta$ weight for each column, and that the value of the linear predictor for each row is $\beta_1 x_1 + \beta_2 x_2 + ... + \beta_n x_n$. Thus, adding a column of ones to the design matrix means we are adding the value of $\beta_1$ to each observation in our data set. This value is the intercept term ($c$) from our equation for a line, $y = mc + c$.
So in our regression, our prediction for the total sleep of each observation will be
$\hat y = \beta_1 + \beta_2 \mathrm{logBodyWt}$
$\beta_1$ and $\beta_2$ are our model coefficents that will be estimated from the data. For now though, let's just think about them as two numbers that we're going to estimate by eye from the scatterplot above.
When log bodyweight is zero, TotalSleep is about 12 hours. So let's set $\beta_1 = 12$. Slope is a bit harder to estimate: I'm going to guess -4. Let's apply these weights to our design matrix to generate some predictions:
betas = np.array([12., -4.]) # the . after the number makes it a floating-point number rather than an integer.
yhat = np.inner(X, betas)
Note that taking the inner product (with the command np.inner) of a matrix and a row vector is the same as multiplying each column by its corresponding beta weight and then summing (i.e. $\beta_1 x_1 + \beta_2 x_2$).
How similar are our predictions against the true values?
# this plot uses straight matplotlib commands, not seaborn:
plt.scatter(yhat, y)
plt.plot((-30, 40), (-30, 40), c=".2", ls="--");
plt.xlabel('yhat (predicted)')
plt.ylabel('y (real)')
plt.show()
From the scatterplot (with dashed identity line) you can see that I got the slope reasonably wrong. Let's compute the errors:
errors = y - yhat # absolute errors for each data point
squared_errors = errors ** 2 # squared errors
sse = squared_errors.sum() # sum of squared errors
sse
array(303857.9545972171)
Statsmodels¶
Now we can use the Statsmodels package to estimate the best coefficients from the data. We will fit it using statsmodels' GLM function.
fit = sm.GLM(y, X).fit() # fit the design matrices we specified above using Patsy...
# let's look at the fit summary:
print(fit.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: TotalSleep No. Observations: 42
Model: GLM Df Residuals: 40
Model Family: Gaussian Df Model: 1
Link Function: identity Scale: 14.0654402509
Method: IRLS Log-Likelihood: -114.09
Date: Tue, 24 Feb 2015 Deviance: 562.62
Time: 17:12:43 Pearson chi2: 563.
No. Iterations: 4
==============================================================================
coef std err z P>|z| [95.0% Conf. Int.]
------------------------------------------------------------------------------
Intercept 11.4046 0.599 19.049 0.000 10.231 12.578
log_BodyWt -0.9474 0.191 -4.965 0.000 -1.321 -0.573
==============================================================================
The above table shows the results of fitting the simple linear regression. There's a bunch of information here. Let's go through some of it.
Note that we've fit a "Gaussian" model with the "identity" link function. That means we're doing ordinary linear regression, which as we said above is one case of the Generalised Linear Model. The identity link means that $f()$ (see above) is just the identity (i.e. it doesn't modify the linear predictor $\mathbf X \beta$ in any way).
We had 42 observations (i.e. all the complete cases in our sleep set). The residual degrees of freedom equals the number of observations minus 2, because we estimated 2 parameters from the data (intercept and slope).
The table at the bottom shows us information about the individual parameters we estimated. The coefficient value of the intercept is 11.4, which is pretty close to my guess of 12. The slope of log bodyweight is -0.95, so my slope estimate was pretty rubbish.
The
zscore is the coefficient divided by its standard error, and the P value is testing whether that standardised coefficient score is different from zero. Both P values are highly significant here, so in this simple case we can infer that both parameters are contributing something useful to predicting the total amount of sleep. The final two columns give us the bounds of the confidence intervals on the coefficient estimates.
Let's compare the predictions of this model to those of my guesses.
yhat_2 = fit.predict()
# note that internally, the fit.predict() function does something similar
# to our inner product between X and beta, above.
plt.scatter(yhat_2, y)
plt.plot((-30, 40), (-30, 40), c=".2", ls="--");
plt.xlabel('yhat_2 (predicted)')
plt.ylabel('y (real)')
plt.show()
errors = y - yhat_2 # absolute errors for each data point
squared_errors = errors ** 2 # squared errors
sse2 = squared_errors.sum() # sum of squared errors
sse2
array(52753.18037853191)
You can see that the fitted values (SSE = 52753) are far closer to the data than my guesses (SSE = 303857)... which is good.
Statsmodels formula inputs¶
Note that we could also do the same thing by passing our Patsy formula to Statsmodels directly, using the statsmodels formula input:
fit_2 = smf.glm(formula='TotalSleep ~ log_BodyWt', data=sleep).fit()
print(fit_2.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: TotalSleep No. Observations: 42
Model: GLM Df Residuals: 40
Model Family: Gaussian Df Model: 1
Link Function: identity Scale: 14.0654402509
Method: IRLS Log-Likelihood: -114.09
Date: Tue, 24 Feb 2015 Deviance: 562.62
Time: 17:12:44 Pearson chi2: 563.
No. Iterations: 4
==============================================================================
coef std err z P>|z| [95.0% Conf. Int.]
------------------------------------------------------------------------------
Intercept 11.4046 0.599 19.049 0.000 10.231 12.578
log_BodyWt -0.9474 0.191 -4.965 0.000 -1.321 -0.573
==============================================================================
Graphically exploring model fit¶
When fitting models, it's a good idea to look at graphical representations of model fit and the residuals (the distance between your predictions and your observations) to diagnose problems. Many plots of model fit are available in Statsmodels, at least for linear models (see here).
We looked at one representation of the relationship between our model fits and the data above. Here are a few common plots.
# first we add to the sleep data frame, make a data frame of the stuff we want to plot:
plots_df = sleep[['TotalSleep', 'log_BodyWt']].copy()
plots_df['yhat'] = fit.predict()
plots_df['resids'] = plots_df['TotalSleep'] - plots_df['yhat']
# note the fit object also contains various residuals.
plots_df.head()
| TotalSleep | log_BodyWt | yhat | resids | |
|---|---|---|---|---|
| 1 | 8.3 | 0.000000 | 11.404567 | -3.104567 |
| 4 | 3.9 | 7.842671 | 3.974367 | -0.074367 |
| 5 | 9.8 | 2.356126 | 9.172358 | 0.627642 |
| 6 | 19.7 | -3.772261 | 14.978433 | 4.721567 |
| 7 | 6.2 | 5.075174 | 6.596313 | -0.396313 |
# plot predictions against real data:
plt.scatter(sleep['log_BodyWt'], sleep['TotalSleep'], c='k')
plt.plot(sleep['log_BodyWt'], fit.predict(), c='r')
plt.xlabel('Log Bodyweight')
plt.ylabel('Total Sleep')
plt.legend(['Prediction', 'Data'])
plt.show()
Residual plots¶
Looking at the residuals as a function of our predictors can show some problems with the modelling assumptions. Recall that our model expects the observations to be independent and the errors normally distributed. We can check this graphically with a few plots.
# plot residuals against bodyweight:
resids = sleep['TotalSleep'] - fit.predict()
plt.scatter(plots_df['log_BodyWt'], resids, c='k')
plt.plot((-8, 10), (0, 0), 'r--')
plt.xlabel('Log Bodyweight')
plt.ylabel('Residuals')
plt.tight_layout()
plt.show()
Basically, we want the spread of the residuals around the zero line to look random. There shouldn't be anything systematic there -- if so, our model is missing non-random (i.e. predictable) relationships in the data.
For example (from here):
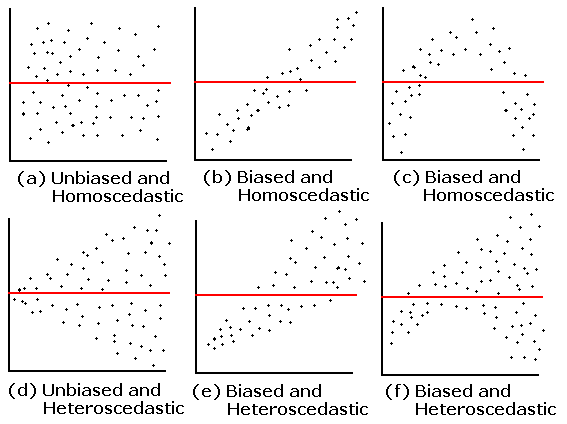
If your residuals plots look something like (b)–(f), think about expanding your model to capture the additional relationships in the data.
The distribution of residuals should also be approximately normal:
plt.hist(resids.values);
In this case, it looks a little right-skewed. That is, we might be violating the assumption of normality in residuals, but probably not drastically.
Seaborn regression plots¶
Handily, Seaborn allows you to do some of these plots directly, without even doing the statsmodels fitting first. This is great for quickly exploring relationships in data, and happily the plots are also much prettier than the default Matplotlib figures. See here for examples and code.
sns.lmplot('log_BodyWt', 'TotalSleep', data=sleep)
sns.despine(offset=10, trim=True);
sns.residplot('log_BodyWt', 'TotalSleep', data=sleep)
sns.despine(offset=10, trim=True);
Internally these functions are fitting the same regression model as we wrote out above, using statsmodels, and wrapping it into pretty exploratory plots for us.
Categorical variables and dummy coding¶
Some variables of interest will be categorical rather than metric / continuous. For example:
- gender is generally thought of as dichotomous (Male, Female)
- a person's religion might be one of a number of categories (Christian, Muslim, Buddhist, nonbeliever, ...)
- "Species" is an example from our dataset above
How are categorical variables handled in the design matrix?
Categorical variables are transformed into one or more design matrix columns with a specific meaning regarding the levels of the variable. The most common method, and the one we explore here, is called Treatment or dummy coding. You can see some examples of other coding schemes built into Patsy here.
For the purposes of this demonstration, let's create a categorical variable "danger_cat" where we consider animals with either high or low danger of predation / exposure.
sleep['danger_cat'] = 'low' # set as a string, "low" for all.
sleep.loc[sleep['Danger'] > 3, 'danger_cat'] = 'high' # set Danger=4 or 5 to "high"
sleep['danger_cat']
1 low 4 high 5 high 6 low 7 high 8 low 9 low 10 high 11 high 14 low 15 low 16 low 17 low 21 high 22 low 24 low 26 low 27 high 28 high 31 high 32 low 33 low 36 low 37 low 38 low 39 low 41 low 42 high 43 low 44 high 45 high 47 low 48 low 49 low 50 low 51 low 53 high 56 low 57 low 58 low 59 high 60 low Name: danger_cat, dtype: object
# how does total sleep appear to change according to our new categories?
sns.set_context('notebook')
g = sns.boxplot(sleep['TotalSleep'], groupby=sleep['danger_cat'])
sns.despine(ax=g, offset=10);
# Note how "violinplot" can be a bit misleading here, due to few cases and the kernel density smooth:
# g = sns.violinplot(sleep['TotalSleep'], groupby=sleep['danger_cat'])
# sns.despine(ax=g, offset=10);
plt.savefig('blah.pdf')
The low danger group seem to sleep for longer on average than the high danger group.
Now imagine we want to use a GLM to test this impression. Is the total amount of sleep in the high danger group less than the low danger group?
y, X = patsy.dmatrices('TotalSleep ~ danger_cat', sleep)
X
DesignMatrix with shape (42, 2)
Intercept danger_cat[T.low]
1 1
1 0
1 0
1 1
1 0
1 1
1 1
1 0
1 0
1 1
1 1
1 1
1 1
1 0
1 1
1 1
1 1
1 0
1 0
1 0
1 1
1 1
1 1
1 1
1 1
1 1
1 1
1 0
1 1
1 0
[12 rows omitted]
Terms:
'Intercept' (column 0)
'danger_cat' (column 1)
(to view full data, use np.asarray(this_obj))
Now see that the design matrix column for danger_cat takes on two values: 0 (for high danger) and 1 (for low danger). What does this mean for the predictions? Let's think through what happens when we multiply these results by $\beta$.
When danger_cat = 1,
$\hat y = \beta_1 * 1 + \beta_2 * 1$
when danger_cat = 0,
$\hat y = \beta_1 * 1 + \beta_2 * 0$
That is, the intercept coefficient $\beta_1$ represents the mean for the high danger group (danger_cat = 0), and the $\beta_2$ coefficient represents the difference between the means of the high group and the low group.
Let's see what those coefficients are:
fit = smf.glm(formula='TotalSleep ~ danger_cat', data=sleep).fit()
print(fit.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: TotalSleep No. Observations: 42
Model: GLM Df Residuals: 40
Model Family: Gaussian Df Model: 1
Link Function: identity Scale: 16.8412589286
Method: IRLS Log-Likelihood: -117.87
Date: Tue, 24 Feb 2015 Deviance: 673.65
Time: 17:12:46 Pearson chi2: 674.
No. Iterations: 4
=====================================================================================
coef std err z P>|z| [95.0% Conf. Int.]
-------------------------------------------------------------------------------------
Intercept 7.2929 1.097 6.649 0.000 5.143 9.443
danger_cat[T.low] 5.0250 1.343 3.741 0.000 2.392 7.658
=====================================================================================
The intercept coefficient is 7.29 hours of sleep. You can see from the box plot above that this corresponds to the mean of the high danger group. The difference between the high and low group (the coefficient for danger_cat) is 5.02 hours. That is, the mean of the low danger group is 7.29 + 5.02 = 12.31 hours (again, check that this looks right according to the box plot).
We can see from the z scores that given the variance in the samples (std err), the intercept is significantly different to zero (i.e. the high danger group sleep significantly more than zero hours) and that the difference between the high and low danger groups is also significant (i.e. the low danger group sleeps for significantly longer than the high danger group).
We've just tested two hypotheses. What do these tests remind you of?
t-tests¶
Testing if the coefficient of the danger_cat variable is different to zero is (almost) the same as an independent samples t-test! It's not exactly the same because glm uses a z-test -- if we fitted the model with ordinary least squares [smf.ols] it would be exactly the same in this case.
To prove to ourselves that these are equivalent, let's do the appropriate tests:
high_array = sleep['TotalSleep'][sleep['danger_cat']=='high']
low_array = sleep['TotalSleep'][sleep['danger_cat']=='low']
# how does this compare to an independent groups z-test?
t, p = sm_stats.weightstats.ztest(high_array, low_array)
'Independent samples z-test: z = {t}, p = {p}'.format(t=np.round(t, decimals=3), p=p)
'Independent samples z-test: z = -3.741, p = 0.00018341854744738338'
'Model values: z = {t}, p = {p}'.format(
t=np.round(fit.tvalues[1], decimals=3),
p=fit.pvalues[1])
'Model values: z = 3.741, p = 0.00018341854744738338'
So we can see that the values from the model are exactly the same as a z-test. What about t?
fit_ols = smf.ols(formula='TotalSleep ~ danger_cat', data=sleep).fit()
print(fit_ols.summary())
OLS Regression Results
==============================================================================
Dep. Variable: TotalSleep R-squared: 0.259
Model: OLS Adj. R-squared: 0.241
Method: Least Squares F-statistic: 13.99
Date: Tue, 24 Feb 2015 Prob (F-statistic): 0.000575
Time: 17:12:46 Log-Likelihood: -117.87
No. Observations: 42 AIC: 239.7
Df Residuals: 40 BIC: 243.2
Df Model: 1
Covariance Type: nonrobust
=====================================================================================
coef std err t P>|t| [95.0% Conf. Int.]
-------------------------------------------------------------------------------------
Intercept 7.2929 1.097 6.649 0.000 5.076 9.510
danger_cat[T.low] 5.0250 1.343 3.741 0.001 2.310 7.740
==============================================================================
Omnibus: 2.740 Durbin-Watson: 1.526
Prob(Omnibus): 0.254 Jarque-Bera (JB): 1.557
Skew: 0.165 Prob(JB): 0.459
Kurtosis: 2.117 Cond. No. 3.23
==============================================================================
Warnings:
[1] Standard Errors assume that the covariance matrix of the errors is correctly specified.
t, p, df = sm_stats.weightstats.ttest_ind(low_array, high_array)
'Independent samples t-test: t({df}) = {t}, p = {p}'.format(
t=np.round(t, decimals=3), df=int(df), p=p)
'Independent samples t-test: t(40) = 3.741, p = 0.0005752964234035196'
'Model values: t = {t}, p = {p}'.format(
t=np.round(fit_ols.tvalues[1], decimals=3),
p=fit_ols.pvalues[1])
'Model values: t = 3.741, p = 0.0005752964234035218'
They are identical to high precision!
One-sample test (mean of high different to zero)¶
The test of the regression coefficient for the intercept is conceptually the same as a one-sample t-test: we ask whether the mean of the "high" group is different to zero. In practice however, because the standard errors in the regression are calculated using the entire design matrix, the test statistics and confidence intervals for the intercept are not the same as a one-sample t-test. Loosely, the standard errors of the intercept depend on the total variance in the dataset (i.e. all variables included in the regression), whereas in the separate one-sample test they do not.
# using statsmodels, must first compute "descrstats":
d1 = sm_stats.weightstats.DescrStatsW(high_array)
t, p = d1.ztest_mean(0)
'Independent z-test: z = {t}, p = {p}'.format(
t=np.round(t, decimals=3), p=p)
'Independent z-test: z = 8.632, p = 6.026877856701133e-18'
Is this the same as from the linear model fit?
'Model values: z = {t}, p = {p}'.format(
t=np.round(fit.tvalues[0], decimals=3),
p=fit.pvalues[0])
'Model values: z = 6.649, p = 2.9453554513479515e-11'
So we can see that the test statistics for the intercept are not the same: 8.632 in the separate test compared to 6.64 in the regression model. As mentioned above, the total variance in the design matrix contributes to the standard errors.
We can check that it's equivalent to fitting an intercept-only model to just the high dataset:
# cell to check "high" alone
high_only = sleep.copy()
high_only = high_only[high_only['danger_cat']=='high']
fit_high = smf.glm(formula='TotalSleep ~ 1', data=high_only).fit()
print(fit_high.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: TotalSleep No. Observations: 14
Model: GLM Df Residuals: 13
Model Family: Gaussian Df Model: 0
Link Function: identity Scale: 9.99302197802
Method: IRLS Log-Likelihood: -35.460
Date: Tue, 24 Feb 2015 Deviance: 129.91
Time: 17:12:46 Pearson chi2: 130.
No. Iterations: 4
==============================================================================
coef std err z P>|z| [95.0% Conf. Int.]
------------------------------------------------------------------------------
Intercept 7.2929 0.845 8.632 0.000 5.637 8.949
==============================================================================
So in this case, where the low cases are excluded and we only fit an intercept, the GLM is the same as the separate one-sample test.
Categorical variable with more than one level¶
We learned above that "dummy coding" is when you transform a categorical variable into 0s and 1s in the design matrix. How does this work for categorical variables with more than two levels?
Let's try by treating the "Lifespan" variable (continuous, lifespan in years) as a categorical rather than continuous variable. I will create a new variable, grouping lifespan into three categories: short, medium, long.
To do this we will use Pandas' qcut function, which cuts a variable into groups based on quantiles. This creates a new Pandas Series whose type is categorical. While categories are going to be really useful in the future, right now they don't play nice with other packages (like statsmodels). For this reason, we will convert the category data type to a plain old object. See the docs for categories here.
sleep['life_cat'] = pd.qcut(sleep['LifeSpan'], [0, .33, .66, 1])
sleep['life_cat']
# category labels are a bit ugly, reflecting range of cuts:
1 [2, 6.765] 4 (20.428, 100] 5 (20.428, 100] 6 (6.765, 20.428] 7 (20.428, 100] 8 (20.428, 100] 9 (20.428, 100] 10 (6.765, 20.428] 11 (20.428, 100] 14 [2, 6.765] 15 (20.428, 100] 16 [2, 6.765] 17 (6.765, 20.428] 21 (6.765, 20.428] 22 [2, 6.765] 24 (20.428, 100] 26 (6.765, 20.428] 27 (6.765, 20.428] 28 (20.428, 100] 31 [2, 6.765] 32 (20.428, 100] 33 (20.428, 100] 36 [2, 6.765] 37 [2, 6.765] 38 [2, 6.765] 39 [2, 6.765] 41 (6.765, 20.428] 42 (6.765, 20.428] 43 (6.765, 20.428] 44 (20.428, 100] 45 (6.765, 20.428] 47 [2, 6.765] 48 (6.765, 20.428] 49 (20.428, 100] 50 (6.765, 20.428] 51 [2, 6.765] 53 (6.765, 20.428] 56 [2, 6.765] 57 (6.765, 20.428] 58 [2, 6.765] 59 (20.428, 100] 60 [2, 6.765] Name: life_cat, dtype: category Categories (3, object): [[2, 6.765] < (6.765, 20.428] < (20.428, 100]]
# change category labels, then convert to object:
sleep['life_cat'] = sleep['life_cat'].cat.rename_categories(['short', 'med', 'long']) # rename categories
sleep['life_cat'] = sleep['life_cat'].astype('object') # make it an object, rather than a category
sleep['life_cat']
1 short 4 long 5 long 6 med 7 long 8 long 9 long 10 med 11 long 14 short 15 long 16 short 17 med 21 med 22 short 24 long 26 med 27 med 28 long 31 short 32 long 33 long 36 short 37 short 38 short 39 short 41 med 42 med 43 med 44 long 45 med 47 short 48 med 49 long 50 med 51 short 53 med 56 short 57 med 58 short 59 long 60 short Name: life_cat, dtype: object
So now we have a new categorical variable, life_cat, with three levels. In this case it would be appropriate to analyse the categories as ordered (see here for how to do that in Patsy, and here for a discussion of different coding schemes). However, for simplicity here we're going to treat this as unordered, meaning that our model thinks the category order doesn't matter (e.g. with something like religion).
What does Patsy do when we make a design matrix using a categorical predictor with more than 2 levels?
y, X = patsy.dmatrices('TotalSleep ~ life_cat', sleep)
X
DesignMatrix with shape (42, 3)
Intercept life_cat[T.med] life_cat[T.short]
1 0 1
1 0 0
1 0 0
1 1 0
1 0 0
1 0 0
1 0 0
1 1 0
1 0 0
1 0 1
1 0 0
1 0 1
1 1 0
1 1 0
1 0 1
1 0 0
1 1 0
1 1 0
1 0 0
1 0 1
1 0 0
1 0 0
1 0 1
1 0 1
1 0 1
1 0 1
1 1 0
1 1 0
1 1 0
1 0 0
[12 rows omitted]
Terms:
'Intercept' (column 0)
'life_cat' (columns 1:3)
(to view full data, use np.asarray(this_obj))
You can see that we now have the intercept plus two additional columns, whose values can be zero or one. Take a look down the columns and compare them to the values of sleep['life_cat'] above.
Let's take for example the first 5 rows:
sleep['life_cat'].head()
1 short 4 long 5 long 6 med 7 long Name: life_cat, dtype: object
X[0:5, :] # first five rows, and second two cols of X
array([[ 1., 0., 1.],
[ 1., 0., 0.],
[ 1., 0., 0.],
[ 1., 1., 0.],
[ 1., 0., 0.]])
Let's work through what this means for the Beta weights:
For rows where 'life_cat' is 'long', the value of the two X columns is [0, 0]. Therefore, the prediction for TotalSleep in these rows is:
$\hat y = \beta_1 * 1 + \beta_2 * 0 + \beta_3 * 0$
$\hat y = \beta_1$
The first beta weight (the intercept) is therefore the mean of the 'long' group.
For rows where 'life_cat' is 'short', the value of the two X columns is [1, 0]. Therefore, the prediction for TotalSleep in these rows is:
$\hat y = \beta_1 * 1 + \beta_2 * 1 + \beta_3 * 0$
$\hat y = \beta_1 + \beta_2$
The second beta weight is therefore the average difference between the 'long' group and the 'short' group.
Finally, for rows where 'life_cat' is 'med', the value of the two X columns is [0, 1]. Therefore, the prediction for TotalSleep in these rows is:
$\hat y = \beta_1 * 1 + \beta_2 * 0 + \beta_3 * 1$
$\hat y = \beta_1 + \beta_3$
The third beta weight is therefore the average difference between the 'long' group and the 'med' group.
Let's see what these coefficients turn out to be, by fitting the model:
fit = smf.glm('TotalSleep ~ life_cat', sleep).fit()
print(fit.summary())
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: TotalSleep No. Observations: 42
Model: GLM Df Residuals: 39
Model Family: Gaussian Df Model: 2
Link Function: identity Scale: 20.0783516484
Method: IRLS Log-Likelihood: -121.03
Date: Tue, 24 Feb 2015 Deviance: 783.06
Time: 17:12:46 Pearson chi2: 783.
No. Iterations: 4
=====================================================================================
coef std err z P>|z| [95.0% Conf. Int.]
-------------------------------------------------------------------------------------
Intercept 8.7071 1.198 7.271 0.000 6.360 11.054
life_cat[T.med] 1.6000 1.694 0.945 0.345 -1.719 4.919
life_cat[T.short] 4.2071 1.694 2.484 0.013 0.888 7.527
=====================================================================================
If we add up these coefficients, we get some predictions for the mean amount of sleep in each category (intercept is just for the 'long' group):
long_mean = fit.params[0]
short_mean = fit.params[0] + fit.params[2]
med_mean = fit.params[0] + fit.params[1]
print('the predictions are: \
\n long = {} \
\n med = {} \
\n short = {}'.format(np.round(long_mean, decimals=1),
np.round(med_mean, decimals=1),
np.round(short_mean, decimals=1)))
the predictions are: long = 8.7 med = 10.3 short = 12.9
Let's check that against a boxplot (which plots medians, but will give us a good idea):
g = sns.boxplot(sleep['TotalSleep'], groupby=sleep['life_cat'])
sns.despine(ax=g, offset=10);
So that looks about right. As before, the z-tests of the coefficients compare groups. The 'short' lifespan group gets significantly more hours of sleep than the 'long' lifespan group (P = .013), but the difference between the 'med' and 'long' groups is not significant (P = .345).
Reference level¶
In the scheme of setting up design matrices we use here, a categorical predictor will have $k - 1$ columns in the design matrix, where $k$ is the number of categories. For example, in our two category (Danger = low or high) example from before, there was $2 - 1 = 1$ column of the design matrix for this categorical predictor.
Note how the intercept takes on a meaning in these designs: the prediction for one of the categories. The category that corresponds to the intercept (or, when all the columns for this variable are zero) is called the reference level or reference category. The additional design matrix columns tell us how the non-reference categories differ from the reference level.
In Patsy, we can specify the reference level manually (e.g. if we wanted the intercept to be the mean of the 'short' group rather than the 'long' group). See the Patsy documentation here for how to do this.
Your homework: multiple predictors¶
In each example above, we have fit GLMs with single predictor variables. It is possible to expand our design matrix to include multiple predictors. In Patsy, the + operator is used to add a predictor to the design matrix.
In a new notebook file, you should
- Import the sleep data
- Create the new variables (log_BodyWt, danger_cat, life_cat) by copying the code from this notebook.
- Fit the model "
TotalSleep ~ log_BodyWt + danger_cat + life_cat" - Think about the meaning of the coefficients. Write down what each coefficient means.
- Bonus: work out the predicted total sleep for a species that is
- low danger
- medium lifespan
- weighs 13 kg.
from IPython.core.display import HTML
def css_styling():
styles = open("../custom_style.css", "r").read()
return HTML(styles)
css_styling()