#Imports for running this presentation live
from IPython.html.widgets import interact, interactive
from IPython.display import clear_output, display, HTML
import numpy as np
from scipy import integrate
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib.colors import cnames
from matplotlib import animation
%matplotlib inline
The Problem¶
Even though computers are often considered deterministic, computational software is a rapidly evolving and changing landscape. Libraries are constantly adding new features and fixing issues.
Even libraries with the strictest backwards-compatibility policies can change in significant ways.

A reproducible computational environment is sufficiently consistent for the computational task at hand.
For example, this can consist of
- a similar CPU instruction set
- libraries and executables available with a specific version and configuration options
- a specific version of a given compiler
- a specific version of a libc implementation
- a specific version of the C++ standard library
Close But Not Good Enough¶

Package managers and distributions¶
- There is not, and there will not be, a consensus on the package manager
- Packages become unsupported over time
- What to do if a required library is not packaged?
Enter Docker¶

Docker is new type of tool to easily build, ship, and run reproducible, binary applications.
It is "good enough" for a reproducible computational environment.
In this talk, we will introduce Docker from the perspective a scientific research software engineer. We will
- Generate an understanding of what Docker is by comparing it to existing technologies.
- Give an introduction to basic Docker concepts.
- Go through examples that demonstrate when and how it is beneficial to take advantage of this tool.
Docker is an open-source engine that automates the deployment of any application as a lightweight, portable, self-sufficient container that will run virtually anywhere.
!docker run --rm busybox sh -c 'echo "Hello Docker World!"'
Docker is a combination of a:¶
- Sandboxed chroot
- Copy on write filesystem
- Distributed VCS for binaries
- Easy to use interface
Sandboxed chroot¶
Docker works with images that consume minimal disk space, versioned, archiveable, and shareable. Executing applications in these images does not require dedicated resources and is high performance.
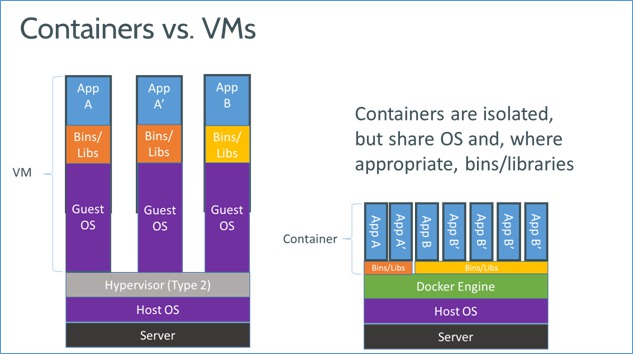
It works with containers as opposed to virtual machines (VM's).
%time !docker run --rm busybox sh -c 'echo "Hello Docker World!"'
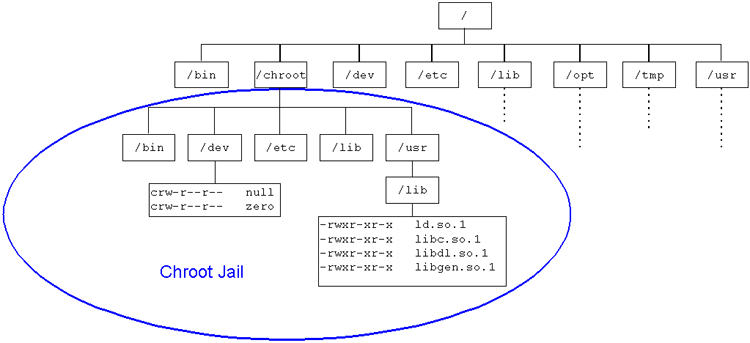
A Docker container is similar to a running an application in a chroot, but it sandboxes processes and the network stack with Linux kernel:
- Namespaces: isolated processes, networking, messaging, file systems, hostname's
- CGroups: groups together cpu, memory, and IO resources
Copy on Write Filesystem¶
Union file systems, or UnionFS, are file systems that operate by creating layers, making them very lightweight and fast. Docker uses union file systems to provide the building blocks for containers. Docker can make use of several union file system variants including:
- AUFS
- btrfs
- vfs
- DeviceMapper

Kernel and root filesystem¶
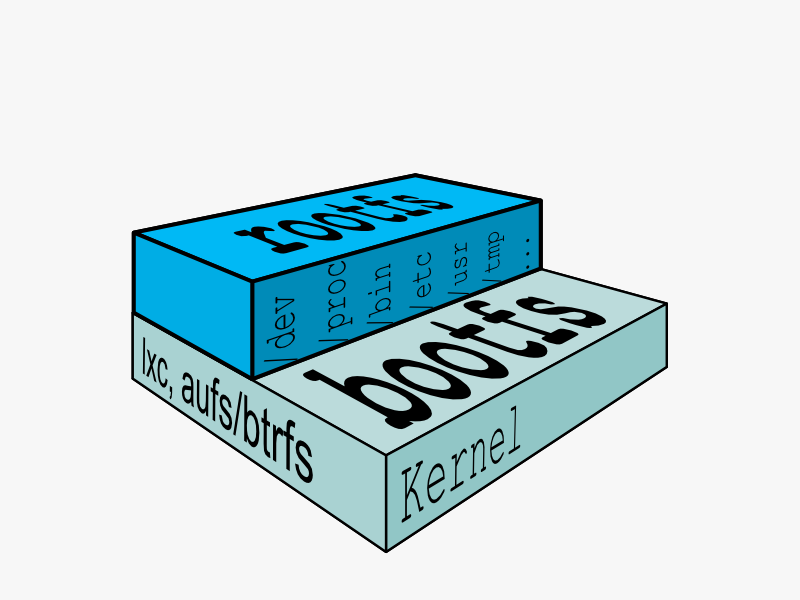

Re-usable, shared filesystem layers¶
Docker is like Git, since¶
- Docker images are identified with hex string or tags
- Interface is
docker <subcommand> docker push,docker pull,docker tagdocker exportwill create a archiveable tarball of an image's filesystem.
Easy to use interface¶
!docker --help
!docker cp --help
Installing¶
Here's what you need:
- Linux kernel with control groups and namespaces
- Support for Another Union File System (AUFS)
- Docker Daemon / Server (written in Go)

Linux¶
- Ubuntu 14.04 or
- See Docker installation instructions for distributions with Kernel 3.8 + later or
- Kernel configuration instructions
Windows and Mac¶
easy install of
- Git Bash
- VirtualBox
- Lightweight Linux distribution
- Docker
Works, but adds layer of complexity
Mac native interface improving
Comes with busybox shell -> Write your Docker build.sh and run.sh in Bourne shell
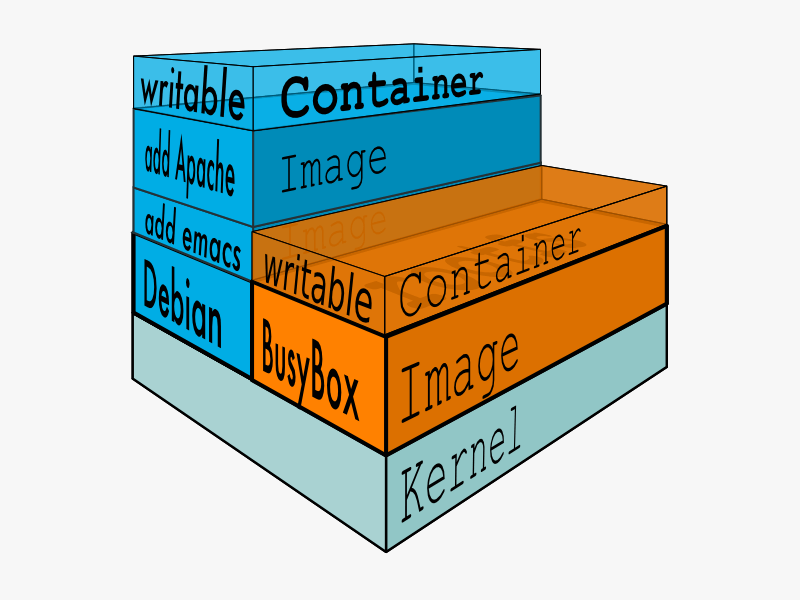
Docker Concepts¶

!docker images
!docker pull odise/busybox-python
!docker images

!docker ps
!docker run -d busybox sh -c 'sleep 3'
!docker ps
!docker ps -a
Volume¶
A mounted directory that is not tracked as a filesystem layer¶
- Data volumes are initialized when a container is created
- Volumes can be shared and reused between containers
- Changes to a data volume are made directly
- Changes to a data volume will not be included when you update an image
- Volume persist until no containers use them
- Host directories can also be mounted as data volumes
!ls $PWD/images/
!docker run --rm --volume $PWD/images:/images busybox \
sh -c 'ls /images'
%%writefile docker-ls-images/Dockerfile
# Best practice for Dockerfile's:
# specify exact versions whenever possible.
FROM busybox:4986bf8c1536
MAINTAINER Matt McCormick <matt.mccormick@kitware.com>
RUN mkdir -p /images
VOLUME /images
CMD ["/bin/sh", "-c", "ls /images"]
Overwriting docker-ls-images/Dockerfile
!docker build -t ls-images ./docker-ls-images
!docker run --rm -v $PWD/images:/images ls-images
Example Applications¶
Creating a Reproducible Computational Environment¶
Reproducing an article on PET image series kinetic analysis
The Article's Build Instructions Part 1¶

The Article's Build Instructions Part 2¶
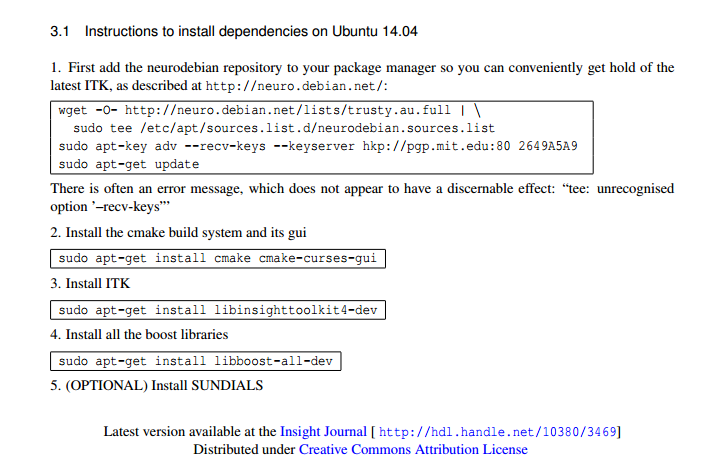
The Article's Build Instructions Part 3¶

The Article's Build Instructions Part 4¶
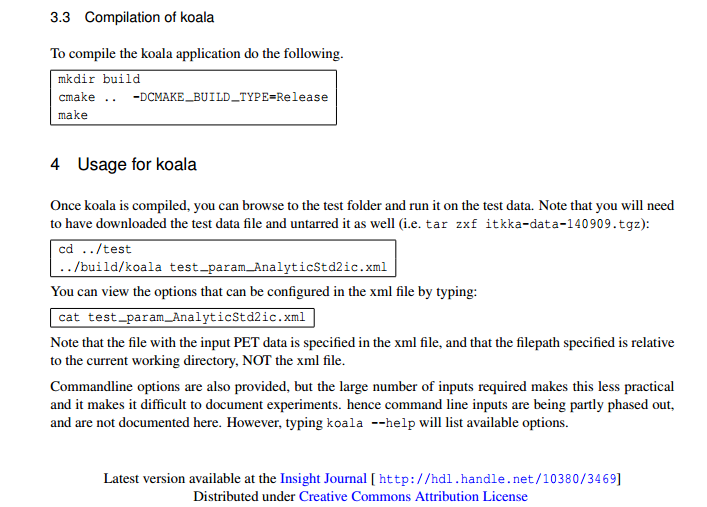
Building and Running with Docker¶
!git clone https://github.com/thewtex/docker-itkka
!cd docker-itkka
!./build.sh
!./run.sh
The requirements for reproducing this computational analysis,
- the Ubuntu 14.04 operating system,
- the GCC 4.8.2-19ubuntu1 compiler,
- and particular versions of the dependencies
- the Insight Toolkit (ITK),
- libxml,
- sundials,
- and Boost,
are now runnable on any x86_64 operating system, archiveable, and shareable.
!docker pull thewtex/itkka
!docker run -it thewtex/itkka bash
Live Presentations, workshops and demos¶
Docker is a good alternative to garlic to stave off the curse of the live demo.
For example, see the course on Modern Scientific Computing with Python from IEEE NSS / MIC or this presentation :-).
from IPython.display import YouTubeVideo
YouTubeVideo("P38cLZJ0w2s")
We can demonstrate interatively how to explore the Lorenz system of differential equations:
$$ \begin{aligned} \dot{x} & = \sigma(y-x) \\ \dot{y} & = \rho x - y - xz \\ \dot{z} & = -\beta z + xy \end{aligned} $$This is one of the classic systems in non-linear differential equations. It exhibits a range of different behaviors as the parameters ($\sigma$, $\beta$, $\rho$) are varied.
We define a function that can integrate the differential equations numerically and then plot the solutions. This function has arguments that control the parameters of the differential equation ($\sigma$, $\beta$, $\rho$), the numerical integration (N, max_time) and the visualization (angle).
def solve_lorenz(N=10, angle=0.0, max_time=4.0, sigma=10.0, beta=8./3, rho=28.0):
fig = plt.figure()
ax = fig.add_axes([0, 0, 1, 1], projection='3d')
ax.axis('off')
# Prepare the axes limits
ax.set_xlim((-25, 25))
ax.set_ylim((-35, 35))
ax.set_zlim((5, 55))
def lorenz_deriv((x, y, z), t0, sigma=sigma, beta=beta, rho=rho):
"""Compute the time-derivative of a Lorentz system."""
return [sigma * (y - x), x * (rho - z) - y, x * y - beta * z]
# Choose random starting points, uniformly distributed from -15 to 15
np.random.seed(1)
x0 = -15 + 30 * np.random.random((N, 3))
# Solve for the trajectory
t = np.linspace(0, max_time, int(250*max_time))
x_t = np.asarray([integrate.odeint(lorenz_deriv, x0i, t)
for x0i in x0])
# Choose a different color for each trajectory
colors = plt.cm.jet(np.linspace(0, 1, N))
for i in range(N):
x, y, z = x_t[i,:,:].T
lines = ax.plot(x, y, z, '-', c=colors[i])
plt.setp(lines, linewidth=2)
ax.view_init(30, angle)
plt.show()
return t, x_t
# A static view provides limited insights
t, x_t = solve_lorenz(angle=0, N=10)
# An interative visualization provides much more insight
# into the parameters
w = interactive(solve_lorenz, angle=(0.,360.),
N=(0,50), sigma=(0.0,50.0), rho=(0.0,50.0))
display(w)
YouTubeVideo('QqfjiuqVrV4')
Graphical Applications and Docker¶
A portable Docker image will only assume standard CPU/memory/disk/network resources are available. The use of local USB devices and video card devices, for example, will not make the images runnable anywhere.
- Use IPython Notebooks
- Use a VNC Server / Client
- Install node-mirror for a browser viewer/terminal/browser/editor
- No OpenGL
Run Exotic or Binary-only Applications¶
MakerBot Makerware¶
The Makerware software to use our MakerBot 3D printer are binaries that only work with certain versions of Ubuntu and Fedora.
With the Makerware image, it is possible to run the latest Makerware with any host Linux distribution.
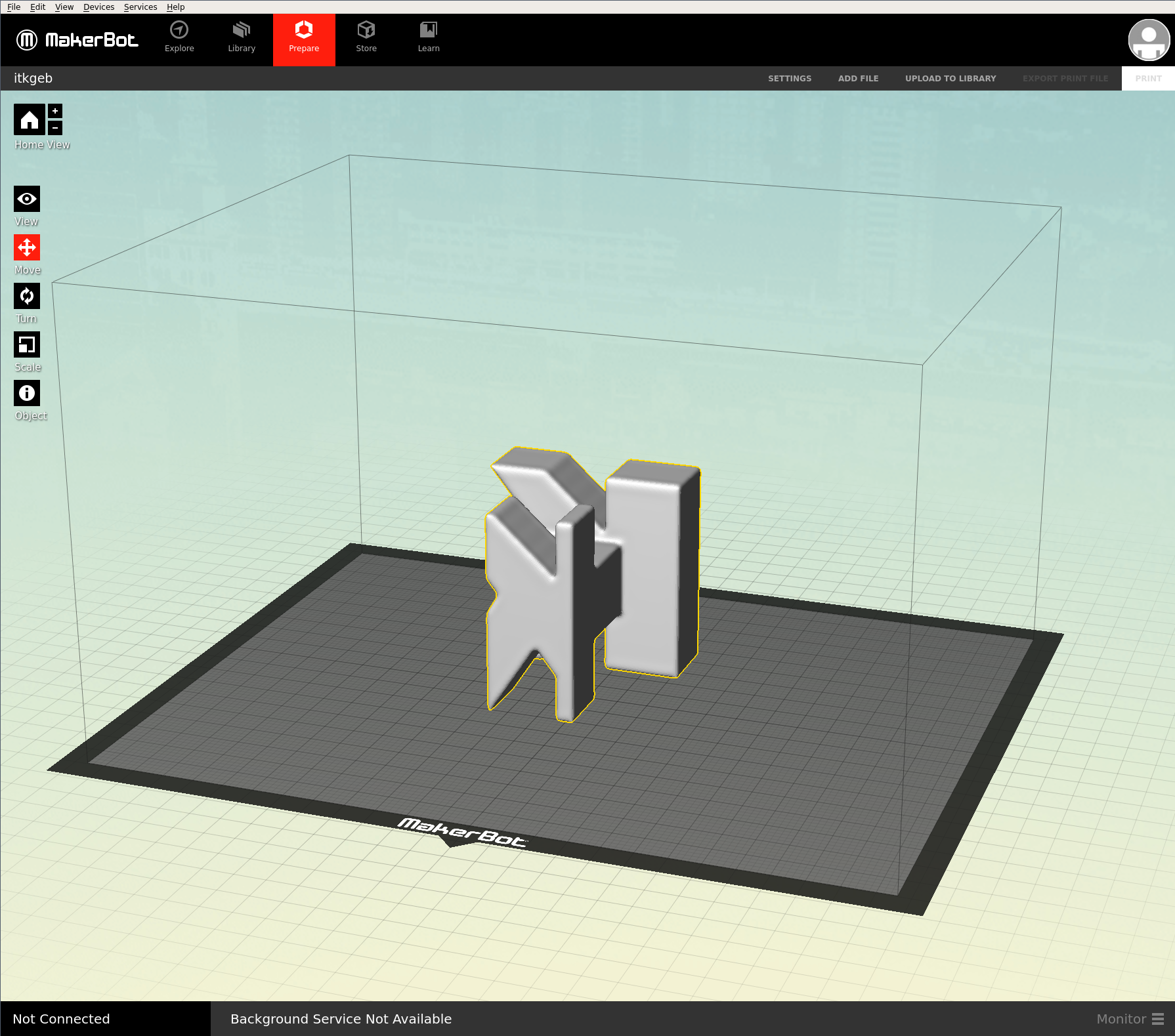
VTK Rocks!¶
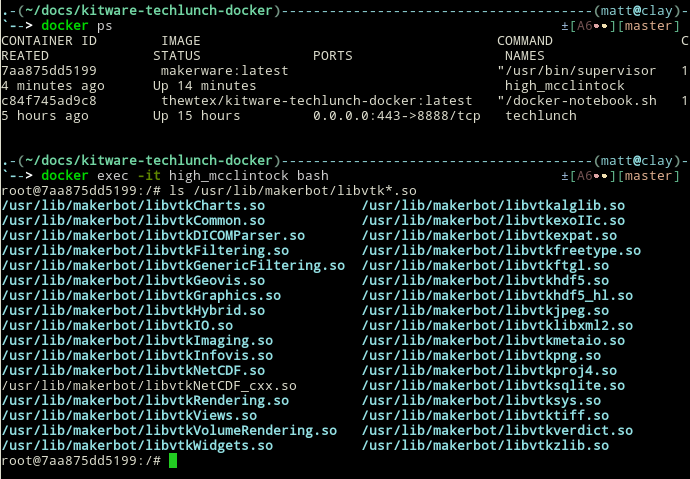
Isn't there supposed to be no OpenGL?

It is possible to run accelerated X11 OpenGL 3D applications, but the Docker images that are built will only work on host systems with the same video driver and compatible video card.
See the docker-opengl-nvidia and the docker-opengl-mesa repositories.
Webapp Development and Deployment¶
Jenkins is outstanding, but it runs on Java.

# Easy execution and sandboxing
!docker pull jenkins
!docker run --name myjenkins -p 8080:8080 -v /var/jenkins_home jenkins
Basic Web App Development and Deployment¶
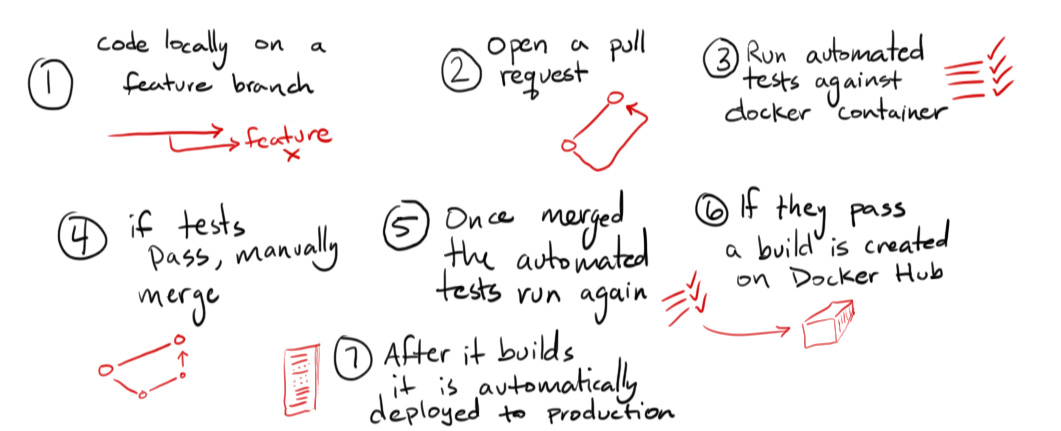
Automated deploy commands:
docker pull mjhea0/flask-docker-workflow
docker stop flask
docker rm flask
docker run --name flask -d -p 80:80 mjhea0/flask-docker-workflow
Many Other Deployment Tools¶
There are many other tools for coordinating many containers that work together, spawning many containers on a cluster, ...
Recap and Next Steps¶
Docker is¶
- Sandboxed chroot +
- Incremental, copy on write filesystem +
- Distributed VCS for binaries +
- Easy to use interface
Concepts¶
- Image: A read-only file system layer
- Container: A writable image with processes running in memory, or an exited container with a modified filesystem
- Volume: A mounted directory that is not tracked as a filesystem layer
- Dockerfile: A sequence of instructions to generate a Docker image
Applications¶
- Reproducible computational environment
- Live presentations, workshops, demos
- Run exotic applications in a sandbox
- Webapp development and deployment
Learn more!¶
Docker vs. LXC¶
- LXC is a set of tools and API to interact with Linux kernel namespaces, cgroups, etc.
- LXC used to be the default execution enviroment for Docker
- Docker provides LXC function, plus:
- Portable deployment across machines
- Application-centric
- Automatic builds
- Versioning
- Component re-use
- Sharing
- Tool echosystem